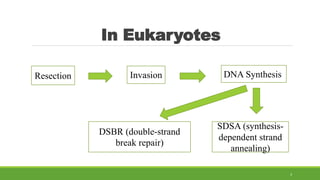
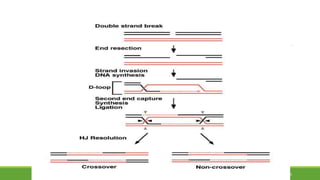
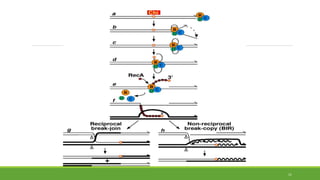
This document summarizes homologous recombination in eukaryotes and bacteria. In eukaryotes, homologous recombination repairs double-strand DNA breaks through either the double-strand break repair (DSBR) pathway or synthesis-dependent strand annealing (SDSA) pathway. The DSBR pathway forms double Holliday junctions that are resolved to result in crossover or non-crossover products. In bacteria, the RecBCD pathway repairs double-strand breaks and the RecF pathway repairs single-strand gaps. Both pathways involve strand invasion and branch migration to facilitate homologous recombination.