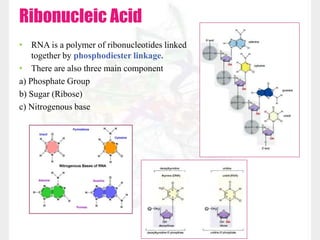
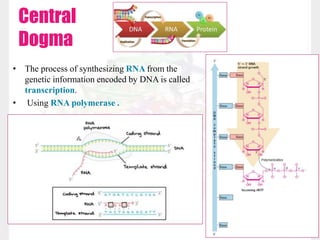
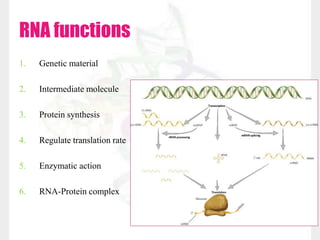
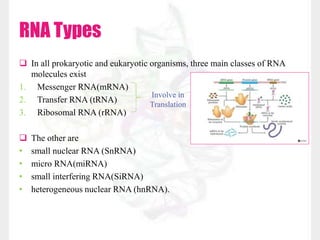
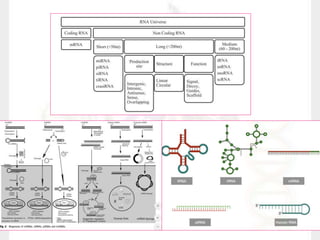
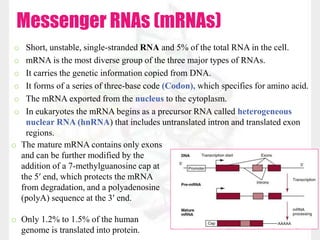
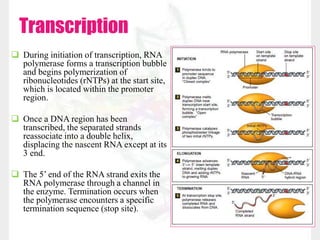
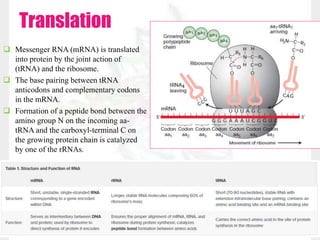
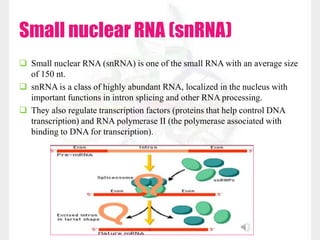
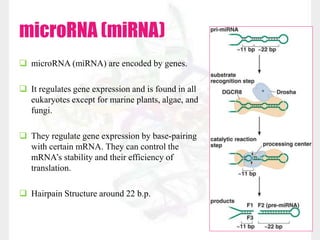
This document discusses RNA structure and types. It begins by describing the basic components and functions of RNA, including its role in transcription and as an intermediate molecule in protein synthesis. It then discusses the different forms and structures of RNA, including primary, secondary and tertiary structures. The main types of RNA - mRNA, tRNA, rRNA and others like miRNA and siRNA - are then summarized in terms of their roles and characteristics. Applications of RNA interference are also briefly outlined.