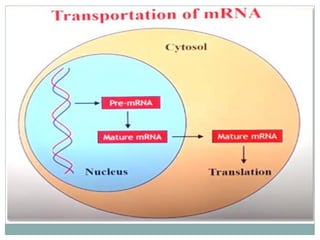
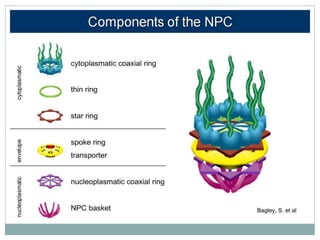
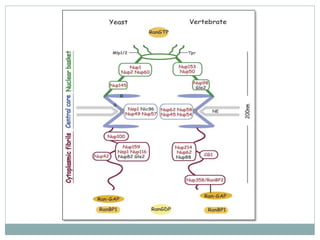
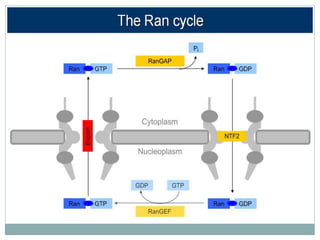
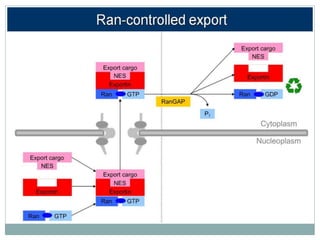
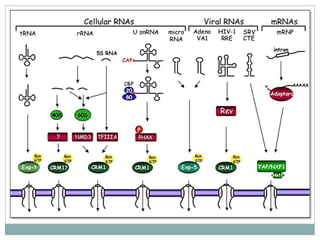
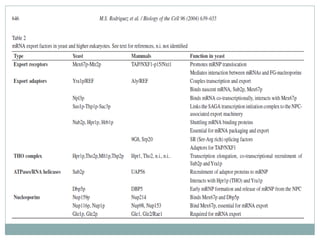
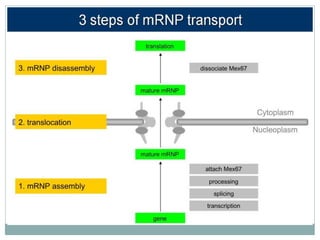
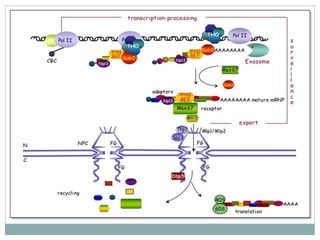
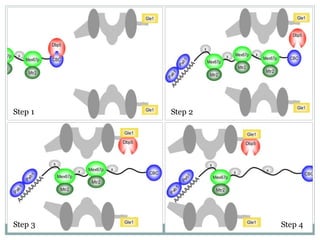
This document summarizes the process of nuclear export of messenger RNA (mRNA). It begins with an introduction describing how mRNA must be exported from the nucleus to the cytoplasm to be translated into protein. It then discusses the importance of nuclear export and describes the nuclear pore complex that facilitates transport. The document outlines the roles of Ran GTPase and transport receptors in nuclear export. It provides details on the adaptor-receptor system and multistep process of mRNA export, including recruitment of export factors, translocation through the nuclear pore, and release into the cytoplasm. The summary concludes with sections on regulation and quality control of mRNA export.