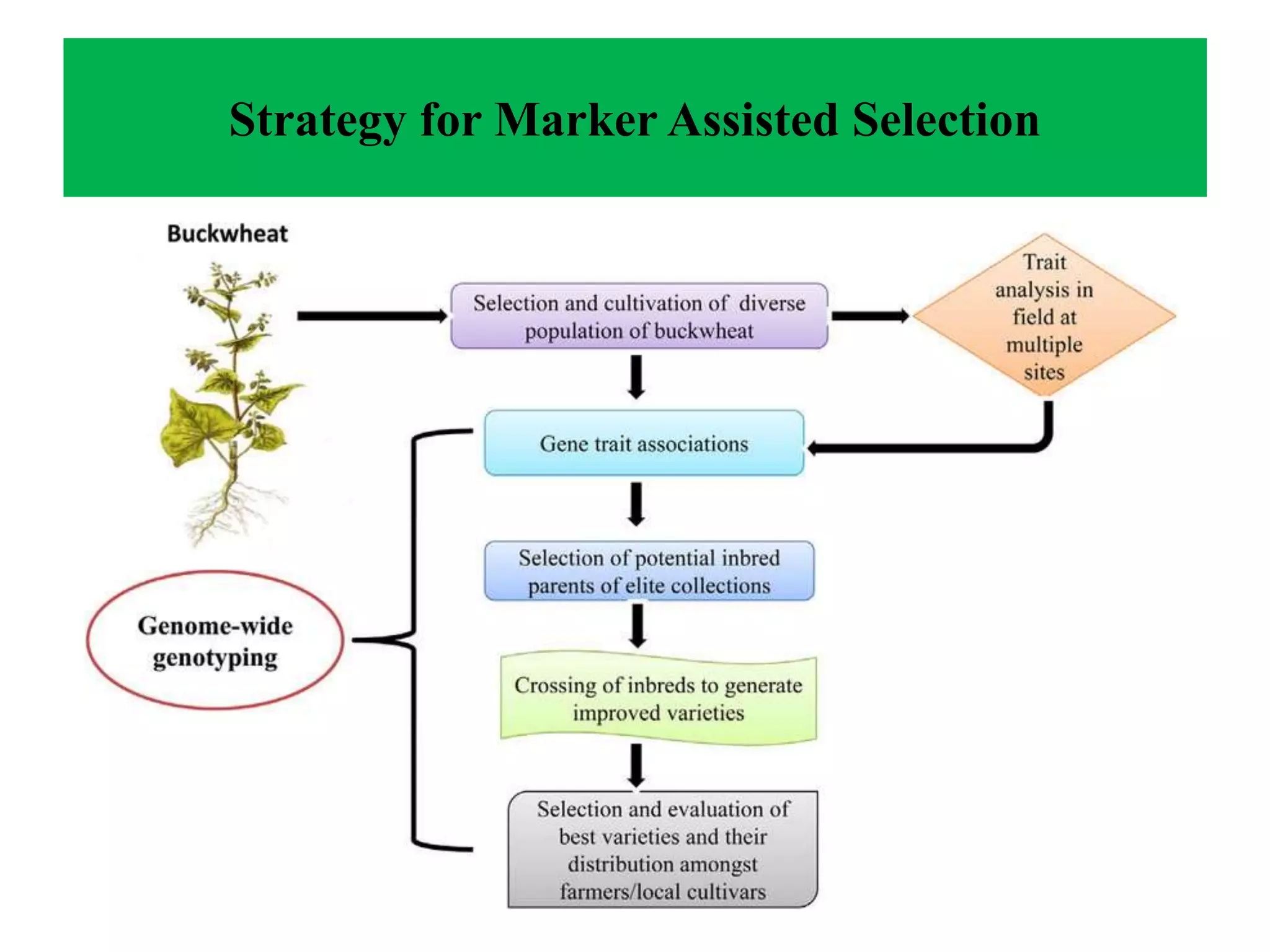
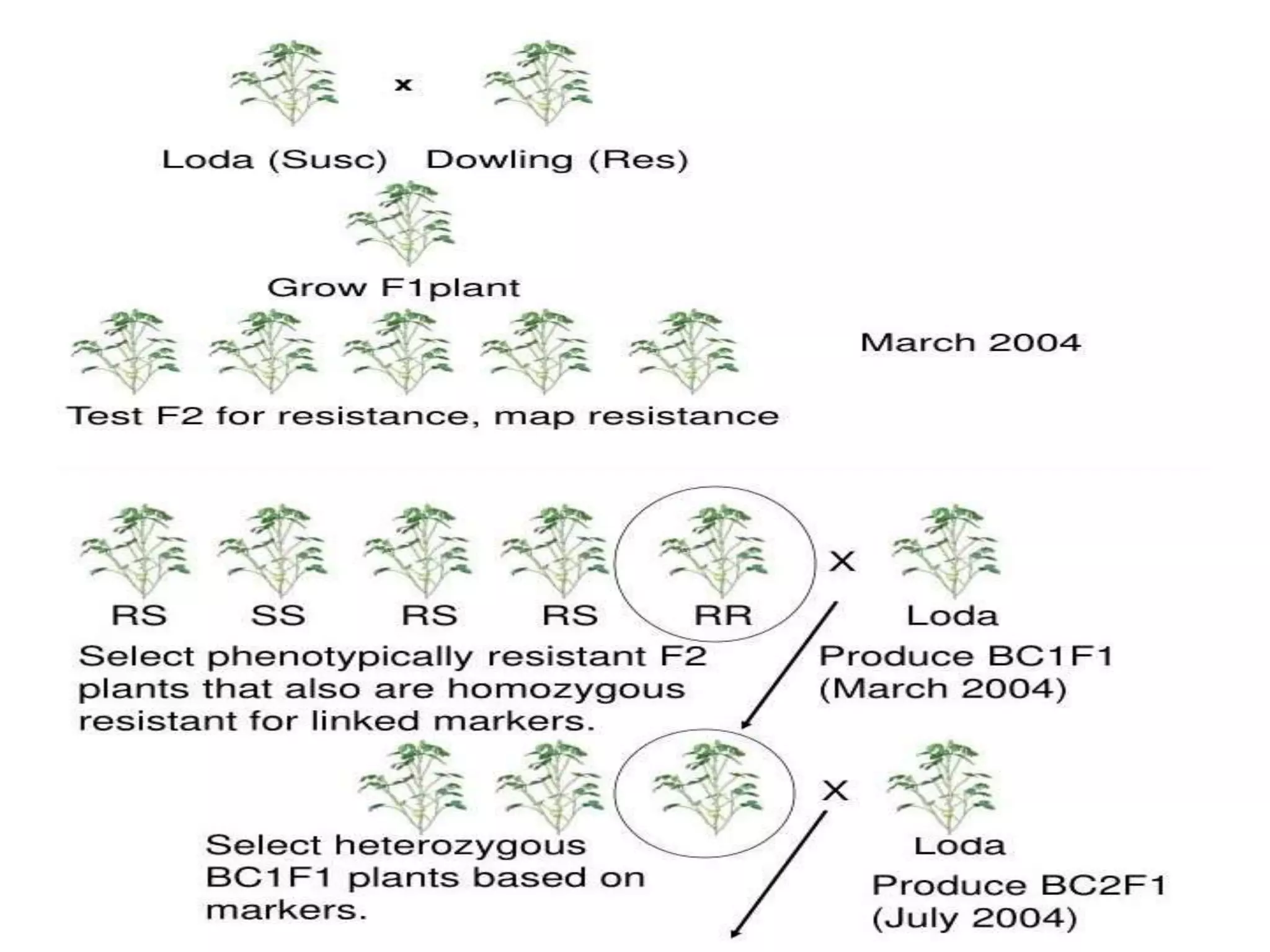
Marker-assisted selection (MAS) integrates genomic and phenotypic data to enhance plant breeding efficiency, enabling more accurate selection of traits like disease resistance and quality improvement. It involves various molecular markers and techniques that facilitate the identification of desirable traits while overcoming limitations of traditional breeding methods. However, MAS is associated with challenges such as higher costs and potential marker-polymorphism issues, necessitating continuous advancements in the field.