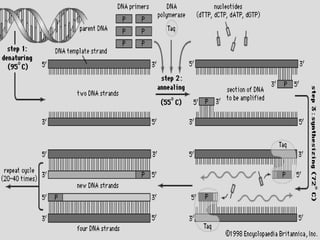
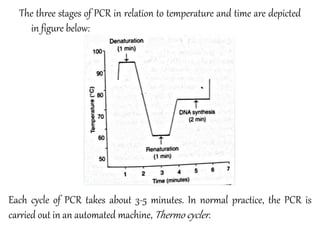
Polymerase chain reaction (PCR) is a technique developed by Kerry Mullis in 1984 that uses thermal cycling to amplify a specific DNA across several orders of magnitude, generating millions of copies of the target DNA segment. It involves repeated cycles of separating DNA strands through heating, annealing primers to the strands through cooling, and extending the primers with a thermostable DNA polymerase through heating. This allows for rapid and efficient amplification of targeted DNA regions.