This document discusses various strategies for gene cloning, including:
1. Extracting target DNA from organisms and purifying it for cloning.
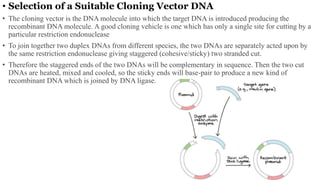
2. Selecting a suitable cloning vector with specific properties.
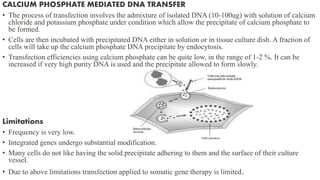
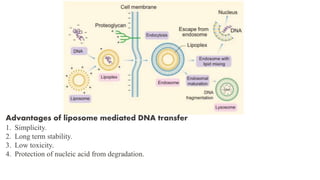
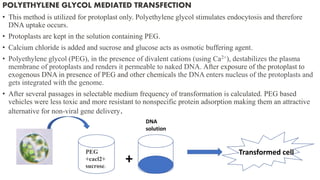
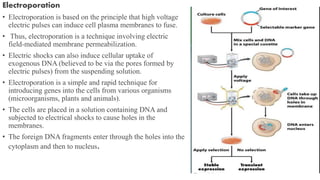
3. Introducing the target DNA into host cells using transformation methods like calcium phosphate transfection, liposome transfection, electroporation, etc.
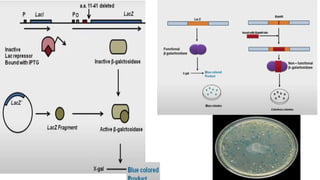
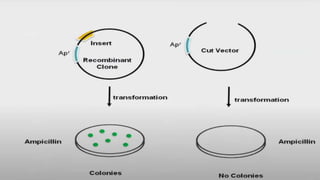
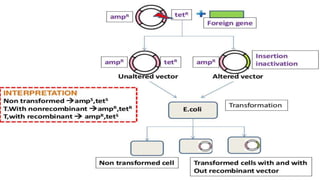
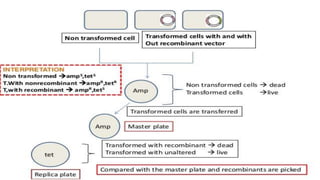
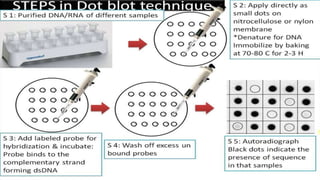
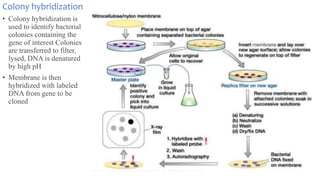
4. Screening and identifying recombinant clones using techniques like antibiotic resistance, colorimetric assays, and nucleic acid hybridization.