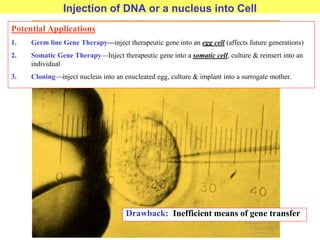
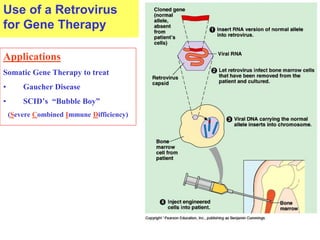
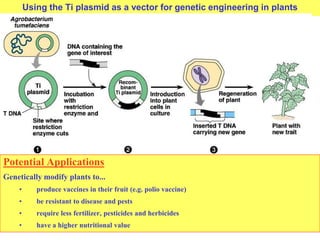
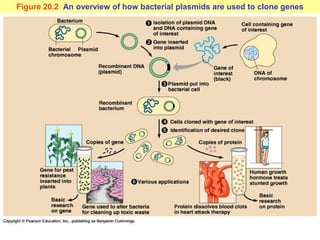
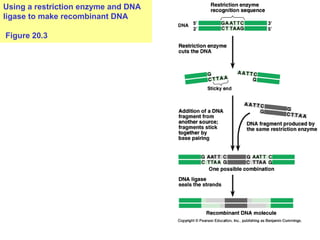
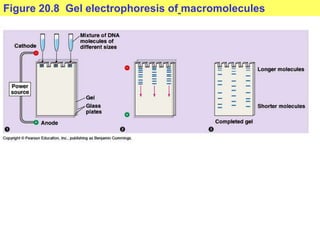
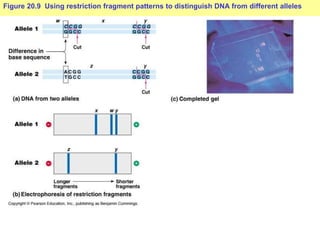
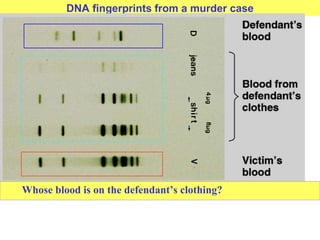
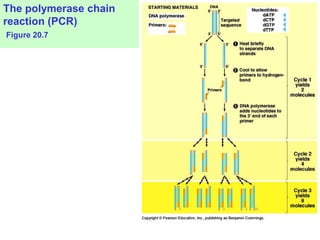
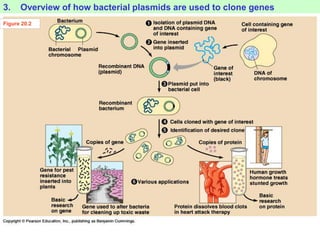
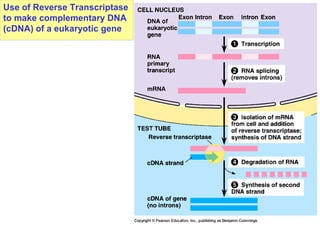
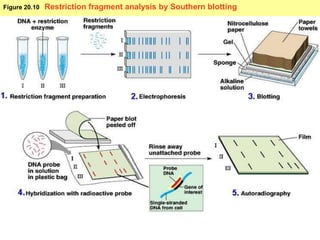
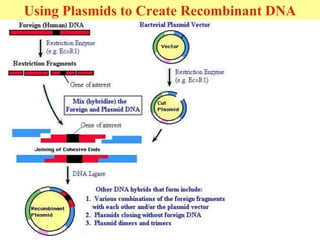
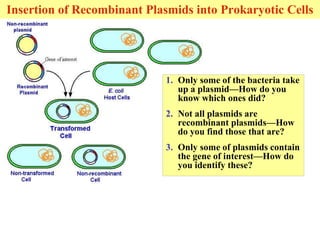
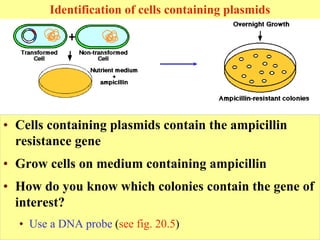
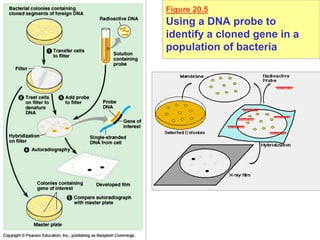
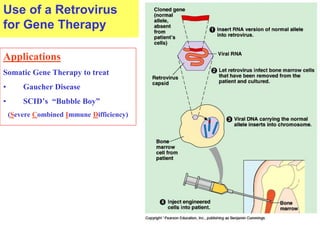
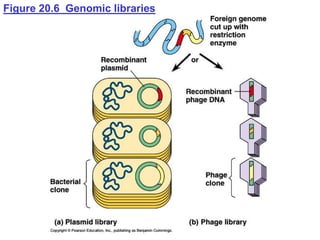
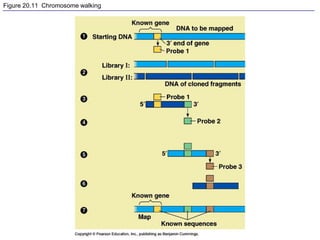
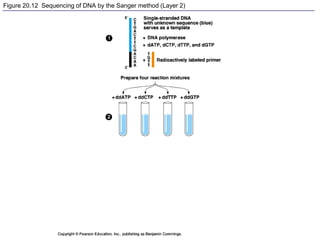
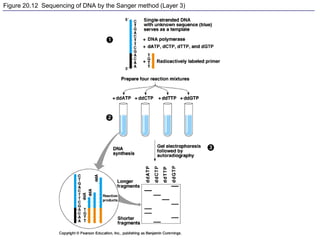
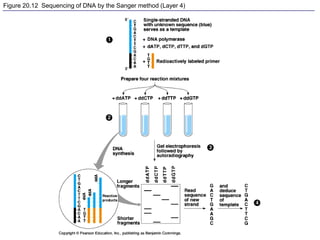
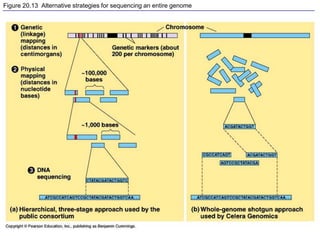
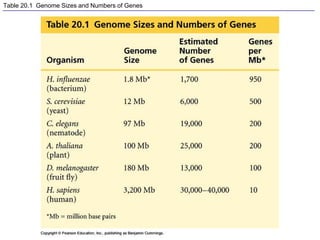
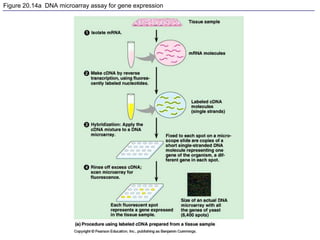
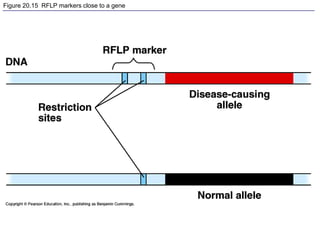
This document provides an overview of advanced molecular biology techniques including recombinant DNA technologies, gene therapy, genetic modification of plants and animals, and tools used in genetic engineering. Key techniques discussed include using restriction enzymes and DNA ligase to create recombinant DNA, gel electrophoresis to separate DNA fragments, PCR to make copies of DNA, and bacterial plasmids to clone genes. Strategies for genetically engineering bacteria and applications of gene therapy like treating immune deficiencies are also summarized.
![PROFESOR DR HJH FARIDA
ZURAINA MOHD YUSOF
Dean of FSG
ADVANCED MOLECULAR BIOLOGY
TECHNIQUES [OVERVIEW & TOOLS]](https://image.slidesharecdn.com/advancedbiotechnology-230203082258-00b4c853/75/Advanced-Biotechnology-ppt-1-2048.jpg)