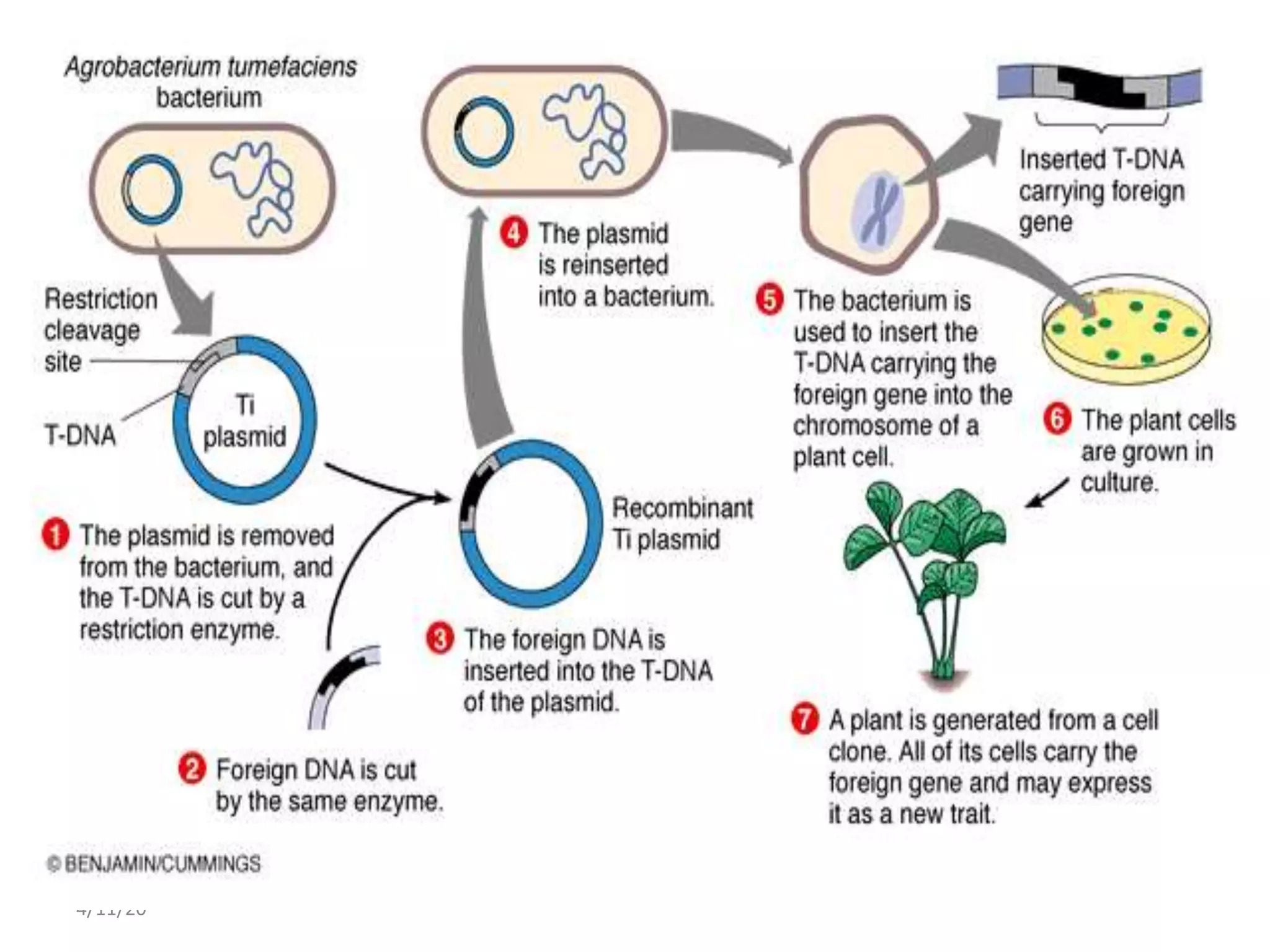
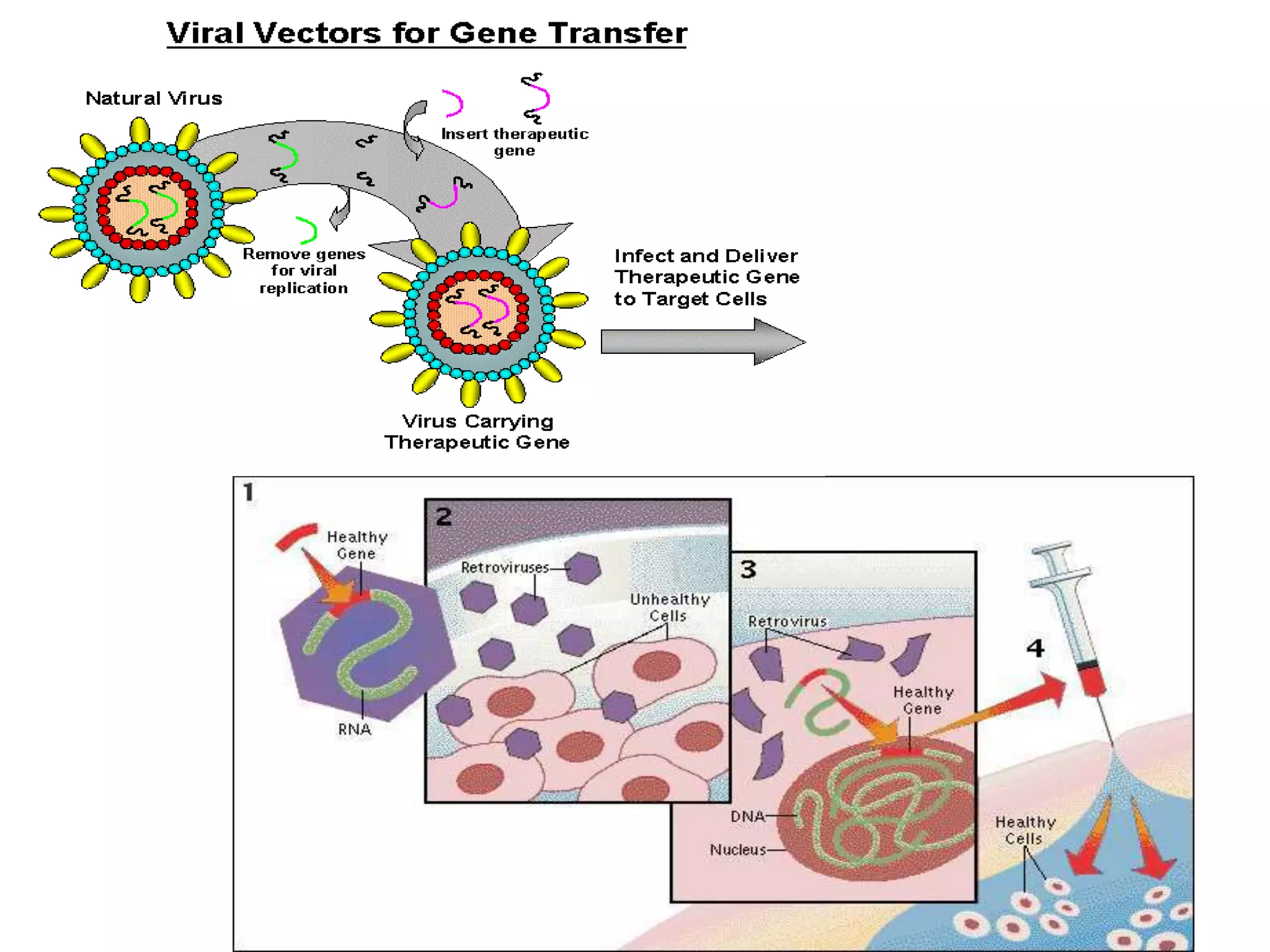
The document discusses gene transfer technology, defined as the stable introduction of foreign genes into target cells' genomes, resulting in the formation of transgenic organisms. It outlines the history and various methods of gene transfer including natural (conjugation, transformation, transduction) and artificial approaches (electroporation, microinjection, biolistics, and liposome-mediated transfer). Each method's advantages and disadvantages are highlighted, showcasing efficiencies and applications in transforming a variety of organisms.