Bacteriophage vectors
•Download as PPTX, PDF•
155 likes•104,517 views
Bacteriophage vectors Bacteriophage WHY BACTERIOPHAGE AS A VECTOR? M13 phage Genome of m13 phage Life cycle and dna replication of m13 CONSTRUCTION M13 AS PHAGE VECTOR M13 MP 2 vector M13MP7 VECTOR Selection of recombinants Lambda replacement vectors LAMBDA EMBL 4 VECTOR P1 PHAGE GENOME OF P1 PHAGE P1 PHAGE AS VECTOR P1 phage vector system
Report
Share
Report
Share
Recommended
Linker, Adaptor, Homopolymeric Tailing & Terminal Transferase

A brief idea about the problems and solutions in cloning regarding Linker, Adaptor, Homopolymeric Tailing & Terminal Transferase enzyme.
bacterial artificial chromosome & yeast artificial chromosome

BAC & YAC are artificially prepared chromosomes to clone DNA sequences.yeast artificial chromosome is capable of carrying upto 1000 kbp of inserted DNA sequence
cDNA Library

This is a brief overview of the process of making a complementary DNA (cDNA) library from mRNA (messenger RNA).
Screening and selection of recombinants 

Screening and selection of recombinants- Direct selection and indirect selection of recombinants
Recommended
Linker, Adaptor, Homopolymeric Tailing & Terminal Transferase

A brief idea about the problems and solutions in cloning regarding Linker, Adaptor, Homopolymeric Tailing & Terminal Transferase enzyme.
bacterial artificial chromosome & yeast artificial chromosome

BAC & YAC are artificially prepared chromosomes to clone DNA sequences.yeast artificial chromosome is capable of carrying upto 1000 kbp of inserted DNA sequence
cDNA Library

This is a brief overview of the process of making a complementary DNA (cDNA) library from mRNA (messenger RNA).
Screening and selection of recombinants 

Screening and selection of recombinants- Direct selection and indirect selection of recombinants
Artificial chromosomes - YAC and BAC

Artificial chromosome - Yeast Artificial Chromosome and Bacterial Artificial Chromosome.
Shuttle vector - a plasmid vector used in rDNA technology. 

shuttle vector used to in rDNA technology to create recombinants
Selection & Screening of Recombinant cells & expression of recombinant (2) (1)

selection ,screening & expression
Selection and screening of recombinant clones 

methods for screening and selection of recombinant clones
Molecular cloning vectors

The molecular cloning vectors used in genetic engineering-plasmids,phage vectors,Hbrid vectors and artificial chromosomes
More Related Content
What's hot
Artificial chromosomes - YAC and BAC

Artificial chromosome - Yeast Artificial Chromosome and Bacterial Artificial Chromosome.
Shuttle vector - a plasmid vector used in rDNA technology. 

shuttle vector used to in rDNA technology to create recombinants
Selection & Screening of Recombinant cells & expression of recombinant (2) (1)

selection ,screening & expression
Selection and screening of recombinant clones 

methods for screening and selection of recombinant clones
What's hot (20)
Shuttle vector - a plasmid vector used in rDNA technology. 

Shuttle vector - a plasmid vector used in rDNA technology.
Selection & Screening of Recombinant cells & expression of recombinant (2) (1)

Selection & Screening of Recombinant cells & expression of recombinant (2) (1)
Viewers also liked
Molecular cloning vectors

The molecular cloning vectors used in genetic engineering-plasmids,phage vectors,Hbrid vectors and artificial chromosomes
Nucleic Acid Purification 

Nucleic Acids: Composition,Purpose of Purification,Modes of Purification & Latest Research Trends.
Topoisomerases

topoisomerases simplified
Subscribe to my youtube channel for such great contents
www.youtube.com/c/AwesomeBiochemistry
Ribotyping

Ribotyping is a molecular technique for bacterial identification that uses information from rRNA-based phylogenetic analyses. It is rapid and specific method widely used in clinical diagnostics and analysis of microbial communities in food, water, and beverages.
Molecular Cloning - Vectors: Types & Characteristics

Molecular Cloning - Vectors: Types & Characteristics
Mitochondria and chloroplast structure and genome organisation

the ppt gives the information about mitochondria and chloroplast structure and genome organisation
Viewers also liked (20)
Molecular detection of food borne pathogens-presentation

Molecular detection of food borne pathogens-presentation
Molecular Cloning - Vectors: Types & Characteristics

Molecular Cloning - Vectors: Types & Characteristics
Mitochondria and chloroplast structure and genome organisation

Mitochondria and chloroplast structure and genome organisation
Similar to Bacteriophage vectors
Phi x 174 phage.

This presentation is about the Phi X 174 phage which include detail information of morphology, life cycle and genome organization of phi X 174 phage.
Bacteriophage vector

my preparation on bacteriophage vector for a brief 10-minute class presentation.
PRESENTATION ON T7 BACTERIOPHAGE.pptx

This ppt is about the T7 phage, it's structure, reproduction, and its genome.
cloning vectors and their role in genetic engineering

cloning vectors and their role in genetic engineering
Major features of e.coli

It is a circular DNA molecule 4.6 million base pairs in length, containing 4288 annotated protein-coding genes (organized into 2584 operons), seven ribosomal RNA (rRNA) operons, and 86 transfer RNA (tRNA) genes.
LAMBDA PHAGE AND THEIR REPRODUCTION 

Introduction, Structure, Lambda DNA, Life cycle of Lambda phage, Genetic switch in Lambda phage
PAC

P1 derived Artificial Chromosomes (PAC) is a genome derived from Phage P1. P1 phage utilises PAC sites for efficient breaking and packaging of the genome and its efficient delivery in transfection stage.
Types of vectors, Prokaryotic and Eukaryotic

vectors types
prokaryotic and eukaryotic
prokaryotic vectors= 1. ti-plasmid 2. pUC18 3. pBR322 4. lambda phage
Eukaryotic vectors= 1.BAC 2. YAC
by
Malaka Madhusudhana Reddy
Research Scholar
Dept.of :: Zoology
Yogivemana University, kadapa.
Andhrapradesh, India.
Similar to Bacteriophage vectors (20)
Vectors part 1 | molecular biology | biotechnology

Vectors part 1 | molecular biology | biotechnology
cloning vectors and their role in genetic engineering

cloning vectors and their role in genetic engineering
12 DNA recomb Tech for the new gene and new info(1).ppt

12 DNA recomb Tech for the new gene and new info(1).ppt
More from priyanka raviraj
CROP PROSPECTS and FOOD SITUATION, 2020

CROP PROSPECTS and FOOD SITUATION, 2020
FAO of UN
world cereal production
Guide TO FINDING YOUR NATURAL TALENTS AND STRENGTHS

THE ULTIMATE GUIDE
"TO FINDING YOUR NATURAL TALENTS AND STRENGTHS"
CLINICAL LABORATORY TECHNIQUES

Protocol for all basic laboratory tests:
Biochemistry tests, microbiology tests, serology tests, histopathology tests
Laboratory Blood/ Specimen collection

Blood Specimen Collection and Processing
VENIPUNCTURE BUTTERFLY NEEDLE METHOD
Sites to draw blood
Order of Draw
Labelling the sample
Areas to Avoid When Choosing a Site for Blood Draw
Techniques to Prevent Hemolysis (which can interfere with many tests)
SAMPLE REJECTION
Blood Sample Handling and Processing
RBC ZINC TEST
HIV 1&2 WESTERN BLOT
Metabolomic Profiling of Spent Biomass Of Marine Microalgae, Chlorella vulgaris

OBJECTIVE:
To evaluate the presence of any high value added compounds in the spent biomass of C. vulgaris
To identify the biological activity of the extracted compounds
To evaluate the structure and nature of the compounds using Nuclear Magnetic Resonance Spectroscopy and other analytical techniques.
Development of economically viable methodologies for the simultaneous extraction of by-products from a single set of biomass.
biological activities performed -Total antioxidant capacity, Anti bacterial activity, Anti-tuberculosis activity, Anti proliferative assay
Photochemistry Mediated Synthesis and Characterization of Thyroxine Capped Si...

Objective:
Silver nanoparticles (AgNPs) are one of the noble metal nanoparticles studied due to their amenability of synthesis, functionalization and ease of detection. Synthesis of silver nanoparticles using thyroxine as a reducing and capping agent through the one step photochemical method
Characterization of synthesized silver nanoparticles (Thy-AgNPs)
1. UV-Spectroscopy Analysis
2. Fourier Transforms-Infra Red Spectroscopy (FT-IR)
3. High Resolution Transmission Electron Microscopy(HR-TEM)
4. Field Emission Scanning Electron Microscopy(FE-SEM)
5. Dynamic Light Scattering (DLS)
6. Zeta potential
Uses:
*AgNPs have unique optical, electrical, and thermal properties
*Exhibit high plasmon efficiency
*More sensitive towards localized surface plasmon resonance
*Less time consuming, economic and more ecofriendly
*It is used in electronics, food industry, cosmetics, photochemical, biomedicine and chemistry.
Rna polymerase

RNA Polymerase
Introduction
Purification
History
PRODUCTS OF RNAP
Messenger RNA
Non-coding RNA or "RNA genes
Transfer RNA
Ribosomal RNA
Micro RNA
Catalytic RNA (Ribozyme)
prokaryotic and eukaryotic
Transcription by RNA Polymerase
TYPES OF RNA POLYMERASE
Type I
Type II
Type III
Prokaryotic Transcription Unit
EXPRESSION OF A PROKARYOTIC GENE
Prokaryotic Polycistronic Message Codes for Several Different Proteins
Eukaryotic Transcription Unit
ENHANCERS AND SILENCERS
RESULT OF THE TRANSCRIPTION CYCLE
RNAP III TRANSCRIBES HUMAN MICRORNAS
RNAP I–specific subunits promote
polymerase clustering to enhance the rRNA gene
transcription cycle
RNAP II–TFIIB STRUCTURE AND
MECHANISM OF TRANSCRIPTION INITIATION
FIVE CHECKPOINTS MAINTAINING THE FIDELITY OF
TRANSCRIPTION BY RNAP IN STRUCTURAL AND
ENERGETIC DETAILS
Dna

DNA
history
structure
X-Ray diffraction image of DNA
base pairing principle
base pairs
bonding patterns of DNA
base stacking different conformations of DNA
different forms of DNA
function of DNA
replication
encoding information
mutation/recombination
gene expression
Application of DNA
Bacterial plasmids

plasmids
history
properties
classification
ColE1 plasmid
characteristics
replication
regulation
application
Sv 40 plasmid
history
properties
applications
PMB 9 Plasmid
construction
application
Third Generation Sequencing 

whole genome analysis
history
needs
steps involved
human genome data
NGS
pyrosequencing
illumina
SOLiD
Ion torrent
PacBio
applications
problems
benefits
Sericulture

Introduction
Sericulture, or silk farming, is the rearing of silkworms for the production of silk.
Species of silkworm
Mulberry silkworm
Tasar silkworm
Muga silkworm
Eri silkworm
Oak silkworm
Giant silkworm
History
Types of silk
Tasar
Eri
Mulberry
Muga
Life cycle
Advantages
Uses
Diseases
Pebrene
Grasserie
Flacherie
Muscardine
Production of silk India
Research Institutes
Artificial production
In vitro culture of embryo
Tissue culture media- Grace’s medium
Cell line production
Nutrition production
More from priyanka raviraj (12)
Guide TO FINDING YOUR NATURAL TALENTS AND STRENGTHS

Guide TO FINDING YOUR NATURAL TALENTS AND STRENGTHS
Metabolomic Profiling of Spent Biomass Of Marine Microalgae, Chlorella vulgaris

Metabolomic Profiling of Spent Biomass Of Marine Microalgae, Chlorella vulgaris
Photochemistry Mediated Synthesis and Characterization of Thyroxine Capped Si...

Photochemistry Mediated Synthesis and Characterization of Thyroxine Capped Si...
Recently uploaded
FAIR & AI Ready KGs for Explainable Predictions

The increased availability of biomedical data, particularly in the public domain, offers the opportunity to better understand human health and to develop effective therapeutics for a wide range of unmet medical needs. However, data scientists remain stymied by the fact that data remain hard to find and to productively reuse because data and their metadata i) are wholly inaccessible, ii) are in non-standard or incompatible representations, iii) do not conform to community standards, and iv) have unclear or highly restricted terms and conditions that preclude legitimate reuse. These limitations require a rethink on data can be made machine and AI-ready - the key motivation behind the FAIR Guiding Principles. Concurrently, while recent efforts have explored the use of deep learning to fuse disparate data into predictive models for a wide range of biomedical applications, these models often fail even when the correct answer is already known, and fail to explain individual predictions in terms that data scientists can appreciate. These limitations suggest that new methods to produce practical artificial intelligence are still needed.
In this talk, I will discuss our work in (1) building an integrative knowledge infrastructure to prepare FAIR and "AI-ready" data and services along with (2) neurosymbolic AI methods to improve the quality of predictions and to generate plausible explanations. Attention is given to standards, platforms, and methods to wrangle knowledge into simple, but effective semantic and latent representations, and to make these available into standards-compliant and discoverable interfaces that can be used in model building, validation, and explanation. Our work, and those of others in the field, creates a baseline for building trustworthy and easy to deploy AI models in biomedicine.
Bio
Dr. Michel Dumontier is the Distinguished Professor of Data Science at Maastricht University, founder and executive director of the Institute of Data Science, and co-founder of the FAIR (Findable, Accessible, Interoperable and Reusable) data principles. His research explores socio-technological approaches for responsible discovery science, which includes collaborative multi-modal knowledge graphs, privacy-preserving distributed data mining, and AI methods for drug discovery and personalized medicine. His work is supported through the Dutch National Research Agenda, the Netherlands Organisation for Scientific Research, Horizon Europe, the European Open Science Cloud, the US National Institutes of Health, and a Marie-Curie Innovative Training Network. He is the editor-in-chief for the journal Data Science and is internationally recognized for his contributions in bioinformatics, biomedical informatics, and semantic technologies including ontologies and linked data.
insect taxonomy importance systematics and classification

documents provide information about insect classification and taxonomy of insect
Structures and textures of metamorphic rocks

It is useful for the Under Graduating students for easy understanding and it's useful for the exam preparations.
Seminar of U.V. Spectroscopy by SAMIR PANDA

Spectroscopy is a branch of science dealing the study of interaction of electromagnetic radiation with matter.
Ultraviolet-visible spectroscopy refers to absorption spectroscopy or reflect spectroscopy in the UV-VIS spectral region.
Ultraviolet-visible spectroscopy is an analytical method that can measure the amount of light received by the analyte.
extra-chromosomal-inheritance[1].pptx.pdfpdf![extra-chromosomal-inheritance[1].pptx.pdfpdf](data:image/gif;base64,R0lGODlhAQABAIAAAAAAAP///yH5BAEAAAAALAAAAAABAAEAAAIBRAA7)
![extra-chromosomal-inheritance[1].pptx.pdfpdf](data:image/gif;base64,R0lGODlhAQABAIAAAAAAAP///yH5BAEAAAAALAAAAAABAAEAAAIBRAA7)
Slide 1: Title Slide
Extrachromosomal Inheritance
Slide 2: Introduction to Extrachromosomal Inheritance
Definition: Extrachromosomal inheritance refers to the transmission of genetic material that is not found within the nucleus.
Key Components: Involves genes located in mitochondria, chloroplasts, and plasmids.
Slide 3: Mitochondrial Inheritance
Mitochondria: Organelles responsible for energy production.
Mitochondrial DNA (mtDNA): Circular DNA molecule found in mitochondria.
Inheritance Pattern: Maternally inherited, meaning it is passed from mothers to all their offspring.
Diseases: Examples include Leber’s hereditary optic neuropathy (LHON) and mitochondrial myopathy.
Slide 4: Chloroplast Inheritance
Chloroplasts: Organelles responsible for photosynthesis in plants.
Chloroplast DNA (cpDNA): Circular DNA molecule found in chloroplasts.
Inheritance Pattern: Often maternally inherited in most plants, but can vary in some species.
Examples: Variegation in plants, where leaf color patterns are determined by chloroplast DNA.
Slide 5: Plasmid Inheritance
Plasmids: Small, circular DNA molecules found in bacteria and some eukaryotes.
Features: Can carry antibiotic resistance genes and can be transferred between cells through processes like conjugation.
Significance: Important in biotechnology for gene cloning and genetic engineering.
Slide 6: Mechanisms of Extrachromosomal Inheritance
Non-Mendelian Patterns: Do not follow Mendel’s laws of inheritance.
Cytoplasmic Segregation: During cell division, organelles like mitochondria and chloroplasts are randomly distributed to daughter cells.
Heteroplasmy: Presence of more than one type of organellar genome within a cell, leading to variation in expression.
Slide 7: Examples of Extrachromosomal Inheritance
Four O’clock Plant (Mirabilis jalapa): Shows variegated leaves due to different cpDNA in leaf cells.
Petite Mutants in Yeast: Result from mutations in mitochondrial DNA affecting respiration.
Slide 8: Importance of Extrachromosomal Inheritance
Evolution: Provides insight into the evolution of eukaryotic cells.
Medicine: Understanding mitochondrial inheritance helps in diagnosing and treating mitochondrial diseases.
Agriculture: Chloroplast inheritance can be used in plant breeding and genetic modification.
Slide 9: Recent Research and Advances
Gene Editing: Techniques like CRISPR-Cas9 are being used to edit mitochondrial and chloroplast DNA.
Therapies: Development of mitochondrial replacement therapy (MRT) for preventing mitochondrial diseases.
Slide 10: Conclusion
Summary: Extrachromosomal inheritance involves the transmission of genetic material outside the nucleus and plays a crucial role in genetics, medicine, and biotechnology.
Future Directions: Continued research and technological advancements hold promise for new treatments and applications.
Slide 11: Questions and Discussion
Invite Audience: Open the floor for any questions or further discussion on the topic.
Orion Air Quality Monitoring Systems - CWS

Professional air quality monitoring systems provide immediate, on-site data for analysis, compliance, and decision-making.
Monitor common gases, weather parameters, particulates.
Nutraceutical market, scope and growth: Herbal drug technology

As consumer awareness of health and wellness rises, the nutraceutical market—which includes goods like functional meals, drinks, and dietary supplements that provide health advantages beyond basic nutrition—is growing significantly. As healthcare expenses rise, the population ages, and people want natural and preventative health solutions more and more, this industry is increasing quickly. Further driving market expansion are product formulation innovations and the use of cutting-edge technology for customized nutrition. With its worldwide reach, the nutraceutical industry is expected to keep growing and provide significant chances for research and investment in a number of categories, including vitamins, minerals, probiotics, and herbal supplements.
Richard's entangled aventures in wonderland

Since the loophole-free Bell experiments of 2020 and the Nobel prizes in physics of 2022, critics of Bell's work have retreated to the fortress of super-determinism. Now, super-determinism is a derogatory word - it just means "determinism". Palmer, Hance and Hossenfelder argue that quantum mechanics and determinism are not incompatible, using a sophisticated mathematical construction based on a subtle thinning of allowed states and measurements in quantum mechanics, such that what is left appears to make Bell's argument fail, without altering the empirical predictions of quantum mechanics. I think however that it is a smoke screen, and the slogan "lost in math" comes to my mind. I will discuss some other recent disproofs of Bell's theorem using the language of causality based on causal graphs. Causal thinking is also central to law and justice. I will mention surprising connections to my work on serial killer nurse cases, in particular the Dutch case of Lucia de Berk and the current UK case of Lucy Letby.
insect morphology and physiology of insect

the documents provide complete information of insect morphology and their structure
SCHIZOPHRENIA Disorder/ Brain Disorder.pdf

This pdf is about the Schizophrenia.
For more details visit on YouTube; @SELF-EXPLANATORY;
https://www.youtube.com/channel/UCAiarMZDNhe1A3Rnpr_WkzA/videos
Thanks...!
Recently uploaded (20)
insect taxonomy importance systematics and classification

insect taxonomy importance systematics and classification
platelets- lifespan -Clot retraction-disorders.pptx

platelets- lifespan -Clot retraction-disorders.pptx
Nutraceutical market, scope and growth: Herbal drug technology

Nutraceutical market, scope and growth: Herbal drug technology
Lateral Ventricles.pdf very easy good diagrams comprehensive

Lateral Ventricles.pdf very easy good diagrams comprehensive
Body fluids_tonicity_dehydration_hypovolemia_hypervolemia.pptx

Body fluids_tonicity_dehydration_hypovolemia_hypervolemia.pptx
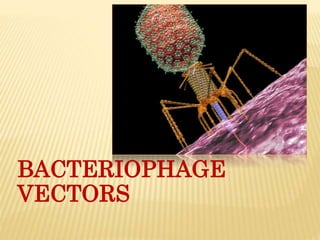
Bacteriophage vectors
- 2. BACTERIOPHAGE Virus that infect bacteria is known as bacteriophage. It was discovered by Frederick.W.Twort in Great Britian (1915) and Felix d’ Herelle in France(1917). D’ Herelle coined the term bacteriophage meaning ‘bacterial eater’ to describe the agent’s bacteriocidal activity.
- 3. Phages are very simple in structure, consisting merely of a DNA (or occasionally ribonucleic acid (RNA)) molecule carrying a number of genes,surrounded by a protective coat or capsid made up of protein molecules. They can undergo two life cycle Lytic cycle Lysogenic cycle
- 4. WHY BACTERIOPHAGE AS A VECTOR? It can accept very large pieces of foreign DNA. Genetic engineers have constructed numerous derivatives of phage vectors that contain only one or two sites for a variety of restriction enzymes. Phage that have a stuffer fragment are called substitution vectors because they are designed to have a piece removed and substituted with something else. Examples are Lambda phage, M13 phage, T4,T7 phage, P1 phage etc.
- 5. M13 PHAGE Bacteriophage M13 was first isolated from wastewater in Munich (Hofschneider, 1963). Hence named as M13 phage. It is a filamentous phage which has 6407 nucleotides. It possess single stranded circular DNA. It was sequenced by Sanger in 1982.
- 6. GENOME OF M13 PHAGE Genes (III, VI, VII, VIII and IX encode structural proteins Genes (II, V and X) for phage DNA replication Genes (I, IV andXI) encode products for assembly and secretion
- 7. LIFE CYCLE AND DNA REPLICATION OF M13
- 8. CONSTRUCTION M13 AS PHAGE VECTOR 1. The first step in construction of an M13 cloning vector was to introduce the lacZ′ gene into the intergenic sequence. 2. This gave rise to M13mp1, which forms blue plaques on X-gal agar. 3. M13mp1 does not possess any unique restriction sites in the lacZ′ gene. ori Lac Z IV II M13MP1 VECTOR
- 9. M13 MP 2 VECTOR It contains the hexanucleotide GGATTC near the start of the gene. A single nucleotide change would make this GAATTC, which is an EcoRI site. This alteration was carried out using in vitro mutagenesis, resulting in M13mp2. ori Lac Z IV II EcoRI
- 10. M13MP7 VECTOR A polylinker, which consists of a series of restriction sites and has EcoRI sticky ends. This polylinker was inserted into the EcoRI site of M13mp2, to give M13mp7 a more complex vector with four possible cloning sites (EcoRI, BamHI, SalI, and PstI). The polylinker is designed so that it does not totally disrupt the lacZ′ gene. Although it is altered, b-galactosidase enzyme is still produced.
- 11. SELECTION OF RECOMBINANTS The vector is then inserted into a competent host cell viable for transformation, which are then grown in the presence of X-gal. Cells transformed with vectors containing recombinants will produce white colonies; non-recombinant plasmids (i.e. only the vector) grow into blue colonies.
- 12. LAMBDA REPLACEMENT VECTORS Replacement vectors A replacement vector has two recognition sites for the restriction endonuclease used for cloning. These sites flank a segment of DNA that is replaced by the DNA to be cloned,. Often the replaceable fragment (or “stuffer fragment” in cloning jargon) carries additional restriction sites that can be used to cut it up into small pieces, so that its own re-insertion during a cloning experiment is very unlikely. Replacement vectors are generally designed to carry larger pieces of DNA than insertion vectors can handle. An example of a replacement vectors is: LAMBDA EMBL4 can carry up to 20 kb of inserted DNA by replacing a segment flanked by pairs of EcoRI, BamHI, and SalI sites. Any of these three restriction endonucleases can be used to remove the stuffer fragment, so DNA fragments with a variety of sticky ends can be cloned. Recombinant selection with eEMBL4 can be on the basis of size, or can utilize the Spi phenotype.
- 13. LAMBDA EMBL 4 VECTOR
- 14. P1 PHAGE P1 is a temperate bacteriophage (phage) that infects Escherichia coli and a some other bacteria. When undergoing a lysogenic cycle the phage genome exists as a plasmid in the bacterium. Unlike other phages it integrate into the host DNA. P1 has an icosahedral "head" containing the DNA attached to a contractile tail with six tail fibers.
- 15. GENOME OF P1 PHAGE The genome of the P1 phage is moderately large, around 93Kbp in length. In the viral particle it is in the form of a linear double stranded DNA molecule. Once inserted into the host it circularizes and replicates as a plasmid. The genome is especially rich in Chi sequences recognized by the bacterial recombinase RecBCD. The genome contains two origins of replication, oriR which replicates it during the lysogenic cycle and oriL which replicates it during the lytic stage.
- 16. P1 PHAGE AS VECTOR The development of a bacteriophage P1 cloning system capable of accepting DNA fragments as large as 100 kilobase pairs (kbp). Phage particles has two P1 loxP recombination sites to cyclize the packaged DNA once it has been injected into a strain of Escherichia coli containing the P1 Cre recombinase, a kanamycin resistant gene to select bacterial clones containing the cyclized DNA. P1 plasmid replicon to stably maintain that DNA in E. coli at one copy per cell chromosome, and a lac promoter-regulated P1 lytic replicon to amplify the DNA before it is reisolated.
- 17. P1 PHAGE VECTOR SYSTEM
