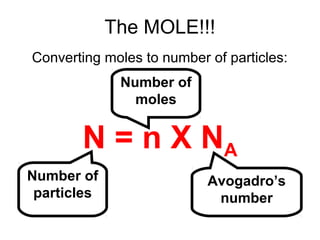
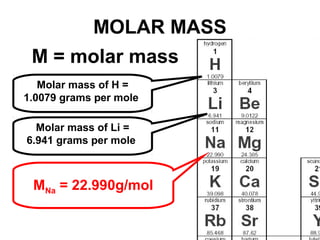
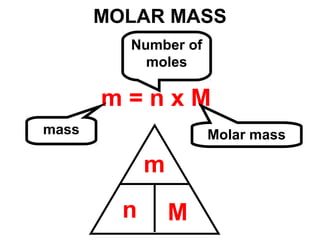
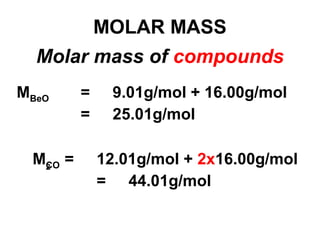The document discusses the mole concept in chemistry. It defines the mole as 6.022x10^23 particles, which is known as Avogadro's number. The mole can refer to individual atoms, molecules, ions or other particles. The document provides examples of how to calculate the number of particles or moles of a substance using the formula N=n×NA. It also discusses molar mass and how to calculate the mass of a substance using the formula m=n×M.