Mutation, repair, recombination
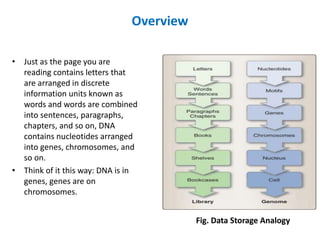
- 1. Overview • Just as the page you are reading contains letters that are arranged in discrete information units known as words and words are combined into sentences, paragraphs, chapters, and so on, DNA contains nucleotides arranged into genes, chromosomes, and so on. • Think of it this way: DNA is in genes, genes are on chromosomes. Fig. Data Storage Analogy
- 2. Physical Organization • The human genome is contained within two distinct compartments : the nucleus and the mitochondria. • The bulk of the genome, containing about 20,000 to 25,000 genes encoded by DNA, is contained within a set of linear chromosomes within the cell nucleus and contains genetic material for both maternal and paternal origin. • In contrast, mitochondrial DNA contains 37 genes that are essential for normal mitochondrial function and is exclusively of maternal origin. Fig. Genes are found in the nucleus and mitochondria
- 3. Human genome = nuclear genome + mitochondrial genome NUCLEAR GENOME 24 distinct chromosomes (22 autosomal + X + Y) 3,200 Mbp 25,000 genes Mitochondrial genome 16,569 bp 37 genes
- 4. DNA building blocks • DNA contains the structural blueprint for all genetic instructions. • The genetic code contained within the DNA is composed of four letters or bases. • Two of the bases are heterocyclic compounds or purines – adenine (A) and guanine (G) – and two are six-member rings known as pyrimidines – cytosine (C) and thymidine (T). • The famous double-helix structure of DNA derives from its phosphate-deoxyribose backbone. • The backbone comprises five-carbon sugar (pentose) molecules bound to a nucleoside (A,G,C, or T). • The pentose molecules are also asymmetrically joined to phosphate groups by phosphodiester bonds. • Hydrogen bonds between complementary (G:C or A:T) nucleotides (a nucleoside linked to a sugar and one or more phosphate groups) interact to stabilize and form the double-helix structure. Fig. Nuclear structure of eukaryotic DNA
- 5. Histones • Chromatin consists of very long double-stranded DNA molecules, nearly an equal mass of rather small basic proteins termed histones, as well as smaller amounts of nonhistones proteins, and a small quantity of ribonucleic acid (RNA). • Histones are heterogenous group of closely related arginine- and lysine-rich basic proteins, which together make up one fourth of amino acid residues. • These positively charged amino acids help histones to bind tightly to the negatively charged sugar phosphate backbone of DNA. • Functionally, histones provide for the compaction of chromatin. Fig. Nuclear genomes packed with proteins to form chromatin
- 6. Human chromosomes 23 pairs 46 chromosomes 22 pairs autosomes 1 pair sex chromosomes 46,XY Normal male 3 billion base pairs in the haploid genome
- 8. DNA packing • The nucleus of a human cell is typically 6 μm in diameter but contains DNA which at its maximum stages of condensation is only about 1/50,000th of its linear length. • Total uncoiled DNA within a single human cell would stretch to more than a meter. • At least four levels of packaging of DNA take place in order that DNA in individual chromosomes fits into the 1.4 μm chromosome seen at metaphase ( a stage in mitosis in the cell cycle where the DNA is most condensed). • Nucleosome are the fundamental organization upon which the higher order packing of chromatin is built. • Each nucleosome core consists of a complex of 8 histone proteins (two molecules each of histone H2A, H2B, H3, and H4) with double stranded DNA wound around it. • 146 base pairs (bp) of DNA are associated with the nucleosome particle and a 50 to 70bp span of linker DNA bound by a linker histone H1 separates each nucleosome. Fig. Structure of a nucleosome
- 9. Chromosome structure • Chromosome structure varies with the cell cycle, from the loose thread like appearance in G1 phase to the tightly compacted state observed during M phase. • Chromosome require three sequence elements for their propagation and maintenance as individual units. • Telomeres are hexameric DNA repeats [(TTAGGG)n] found at the ends of chromosomes that serve to protect the chromosome from degradation. • Sequence elements known as centromeres serve “handles”, which allow mitotic spindles to attach to the chromosome during cell division. • As the cell progresses through the mitotic or M phase of the cell cycle, the nuclear envelope breaks down, and chromosome segregate into the opposite poles of the cells (to form daughter cells), while kinetochore forms consisting of the centromere and mitotic spindles.
- 10. Information organization • Ploidy refers to the number of chromosome copies within a cell. • Most of the somatic cells within the body are diploid, meaning that each nucleus has two copies of each chromosome, one deriving from the mother and other from the father. • Germ cells are the exception to this rule, which contain a single copy of each chromosome and are known haploid. • The haploid genome of each human cell consists of 3.0 x 109 bp of DNA, divided into 23 (22 somatic and 1 sex) chromosomes. • The entire haploid genome contains sufficient DNA to code for nearly 1.5 million pairs of genes. • The human genome project has shown humans to have only about 20,000 to 25,000 genes. • Our genome has approximately the same number of genes as a fruit fly, yet considerably more complex that that organism. • The human proteome, or total number of protein species, is five to ten times larger than that of the fruit fly. Fig. Organization of the genome
- 11. Unique DNA sequences • Single copy DNA or genes generally encode information for specific protein products. • The 20,000 to 25,000 genes within the human genome can be divided into four general categories. • Approximately 5,000 genes are involved with genome maintenance; nearly 5,000 with signal transduction; and 4,000 with general biochemical functions; the largest portion, 9,000 genes, is involved in other activities. Fig. Unique (nonrepetitive) DNA distribution with the genome
- 12. Repeat sequences • Repeat sequences do not encode proteins, but they make up at least 50% of the human genome. • These sequences fall into two main classes: satellite DNA and LINES and SINES. 1. Satellite DNA • These highly repetitive sequences tend to be clustered and repeated many times in tandem ( a head-to-toe arrangement). • They are generally not transcribed and are present in 1 to 10 million copies per haploid genome. • These sequences are also associated with the centromeres and telomeres of chromosomes. • Satellite DNA sequences are categorized according to the number of base pairs within the repeat sequences: • Alpha satellite – 171 bp sequence that extends several million base pairs or more in length. • Minisatellite - 20 to 70 bp in length and a total legth of a few thousand base pairs. • Microsatellite – repeat units only 2, 3, or 4 bp in length and a total length of a few hundred.
- 13. 2. LINES and SINES • these clustered sequences are found interspersed with unique sequences. • These are also present at less than 106 copies per haploid genome. • They are transcribed into RNA and can be according to their size. • LINES (long interspersed elements) 7,000 bp (20 to 50,000 copies) • SINES (short interspersed elements) 90 to 500 bp (about 100,000 copies)
- 14. Functional organization • Functions within the cell are usually encoded by genes. • Genes are present on the chromosome and in the mitochondria. • Not all genes are active in all tissues and specific alterations to these genes are necessary for tissue-specific gene expression. A. Genes • A gene is the complete sequence region necessary for generating a functional product. • This encompasses promoters and control regions necessary for the transcription, processing, and, if applicable, translation of gene. • About 2% of the genome encodes instructions for the synthesis of proteins. B. Alteration to the genome • Aside from mutation, all cells in an individual have identical DNA content and sequence. • However, different tissues and cells require a specific set of genes to carry out their functions. Fig. chromatin structure is tissue specific
- 15. Genome • The genome is all the DNA in a cell. – All the DNA on all the chromosomes – Includes genes, intergenic sequences, repeats • Specifically, it is all the DNA in an organelle. • Eukaryotes can have 2-3 genomes – Nuclear genome – Mitochondrial genome – Plastid genome • If not specified, “genome” usually refers to the nuclear genome.
- 16. Genomics • Genomics is the study of genomes, including large chromosomal segments containing many genes. • The initial phase of genomics aims to map and sequence an initial set of entire genomes. • Functional genomics aims to deduce information about the function of DNA sequences. – Should continue long after the initial genome sequences have been completed.
- 17. Genomes and Genomics • The word “genome,” coined by German botanist Hans Winkler in 1920, was derived simply by combining gene and the final syllable of chromosome. • If not specified, “genome” usually refers to the nuclear genome! • An organism’s genome is defined as the complete haploid genetic complement of a typical cell. • The genetic content of the organelles in the cell, is not considered part of the nuclear genome. • In diploid organisms, sequence variations exist between the two copies of each chromosome present in a cell. • The genome is the ultimate source of information
- 18. "Genes" are units of genetic information present on the DNA in the chromosomes and chromatin. " Genome" is all the DNA contained in an organism or a cell, which includes the chromosomes plus the DNA in mitochondria (and DNA in the chloroplasts of plant cells).
- 19. • The number of genomes sequenced in their entirety is now in the thousands and includes organisms ranging from bacteria to mammals. • The first complete genome to be sequenced was that of the bacterium Haemophilus influenzae, in 1995. • The first eukaryotic genome sequence, that of the yeast Saccharomyces cerevisiae, followed in 1996. • The genome sequence for the bacterium Escherichia coli became available in 1997 . • The much larger effort directed at the human genome was also accelerating.
- 20. Prokaryotes and Eukaryotes genome Prokaryotes Eukaryotes Single cell Single or multi cell No nucleus nucleus One piece of circular DNA Chromosomes No mRNA post transcriptional modification Exons/Introns splicing
- 21. Karyotype o The study of chromosomes, their structure and their inheritance is known as Cytogenetics. o Each species has a characteristic number of chromosomes and this is known as karyotype. • Bacteria 1 • Fruit fly 8 • Garden Pea 14 • Yeast 16 • Frog 26 • Cat 38 • Fox 34 • Mouse 40 • Rat 42 • Rabbit 44 • Human 46 • Chicken 78
- 22. o Prior to 1950's it was believed that humans had 48 chromosomes but in 1956 it was confirmed that each human cell has 46 chromosomes (Tjio and Levan, 1956). o On the chromosomes the genes are situated in a linear order. o Each gene has a precise position or locus. o The size of bacterial chromosomes ranges from 0.6 -10 Mbp, and the size of Archael range from 0.5 - 5.8 Mbp, whereas Eukaryotic chromosomes range from 2.9 - 4,000 Mbp.
- 23. Human Genome: General Information • Genetic material in humans is stored in two organelles: nucleus (about 3200 Mbp) and mitochondria (16.6 kb). • Human chromosomes are not of equal sizes; the smallest, chromosome 21, and the largest, chromosome 1. • Only a very small amount of human DNA is responsible for the differences among humans, indeed among all organisms.
- 24. Number of genes in the human genome • Number of genes at least 100,000. • HOWEVER, the number of protein‐encoding genes is only ~20,000 to 25,000. • How can we explain this?
- 25. Lecture no. 5
- 26. Genomics Vs. Genetics • Genetics: study of inherited phenotypes Peter Goodfellow (1997, Nature Genetics 16:209-210): "...I would define genetics as the study of inheritance and genomics as the study of genomes. The latter informs the former and includes the sequencing of genomes. The concept of functional genetics is a tautology (the whole point of genetics is to link genes with phenotypes). Functional genomics is the attachment of information about function to knowledge of DNA sequence' paradoxically, genetics is a major tool for functional genomics."
- 27. Genomics, Genetics and Biochemistry • Genetics: study of inherited phenotypes • Genomics: study of genomes • Biochemistry: study of the chemistry of living organisms and/or cells • Revolution lauched by full genome sequencing – Many biological problems now have finite (albeit complex) solutions. – New era will see an even greater interaction among these three disciplines
- 28. Distinct components in complex genomes • Highly repeated DNA – R (repetition frequency) >100,000 – Almost no information, low complexity • Moderately repeated DNA – 10<R<10,000 – Little information, moderate complexity • “Single copy” DNA – R=1 or 2 – Much information, high complexity
- 29. MUTATION DEFINTION • A mutation can be defined as a permanent change in the DNA segment of a gene which may be phenotypically silent or expressed. • The substances (chemicals) which can induce mutations are collectively known as mutagens. CAUSES OF MUTATIONS • Mutations are cause by 1. Error in proofreading during DNA Replication • DNA polymerase corrects the mismatch during replication by proofreading. If the proofreading is not effective, the lesion remains in the DNA molecule. 2. Error in DNA Repair • If the change in base sequence occurs after DNA replication, and if the DNA repair is defective, the lesion remains in the DNA molecule. 3. Error in DNA Recombination • DNA recombination occurs by translocation and rearrangement of genes. If the recombination is defective, it can lead to mutations.
- 30. 4. Chemical Mutagens • Chemical mutagens alter DNA bases or structure of DNA. If these alterations are not repaired, they lead to mutations. • Examples of mutagens include base analogs, alkylating agents, intercalating agents, deaminating agents, hydrazines and aflatoxin. 5. Irradiation • Ultraviolet light or ionizing radiations can alter the structure of DNA. These changes lead to mutations, if they are not repaired. 6. Spontaneous Mutations • Mutations can occur without any underlying cause. TYPES OF MUTATIONS • Mutations are classified into two major categories, i.e.: 1. Point mutation, and 2. Frame-shift mutation.
- 31. 1. POINT MUTATIONS • The replacement of one base pair by another results in point mutation. • They are of two sub- types. A. Transitions: • In this case, a purine (or a pyrimidine) is replaced by another. B. Transversions: • These are characterized by replacement of a purine by a pyrimidine or vice versa.
- 32. Consequences of point mutations : • The change in a single base sequence in point mutation may cause one of the following. 1. Silent mutation • The codon (of mRNA) containing the changed base may code for the same amino acid. • For instance, UCA codes for serine and change in the third base (UCU) still codes for serine. Therefore, there are no detectable effects in silent mutation. 2. Missense mutation • In this case, the changed base may code for a different amino acid. • For example, UCA codes for serine while ACA codes for threonine. • The mistaken (or missense) amino acid may be acceptable, partially acceptable or unacceptable with regard to the function of protein molecule. • Sickle-cell anemia is a classical example of missense mutation. 3. Nonsense mutation • Sometimes, the codon with the altered base may become a termination (or nonsense) codon. • For instance, change in the second base of serine codon (UCA) may result in UAA. • The altered codon acts as a stop signal and causes termination of protein synthesis, at that point.
- 33. Fig. An illustration of point mutations represented by a codon of mRNA
- 34. 2. Frameshift mutations • These occur when one or more base pairs are inserted in or deleted from the DNA, respectively, causing insertion of deletion mutations. Consequences of frameshift mutations: • The insertion or deletion of a base in a gene results in an altered reading of the mRNA (hence the name frameshift). • The machinery of mRNA (containing codons) does not recognize that a base was missing or a new base was added. • Since there are no punctuations in the reading of codons, translation continues. • The result is that the protein synthesized will have several altered amino acids and/or prematurely terminated protein.
- 36. REPAIR OF DNA INTRODUCTION • DNA is the only macromolecule that is repaired rather than degraded. • DNA repair is a high-priority process for the maintenance of integrity of the genetic information and other cellular functions within the cell both, in prokaryotes as well as eukaryotes. • It is a very efficient process with less than 1 out of 1000 accidental changes which may result in a mutation. • The cell possesses an inbuilt system to repair the damaged DNA. • This may be achieved by four distinct mechanisms: 1. Base excision repair 2. Nucleotide excision repair 3. Mismatch repair 4. Double-strand break repair
- 37. 1. Base excision-repair • This type of repair eliminates modified bases like those that have been deaminated, methylated or modified chemically. • The bases cytosine, adenine and guanine can undergo spontaneous, depurination to respectively form uracil, hypoxanthine and xanthine. • These altered bases do not exist in the normal DNA, and therefore need to be removed. • This is carried out by base excision repair. • A defective DNA in which cytosine is deaminated to uracil is acted upon by the enzyme uracil DNA glycosylase. • This results in the removal of the defective base uracil. • An endonuclease cuts the backbone of DNA strand near the defect and removes a few bases. • The gap so created is filled up by the action of repair DNA polymerase and DNA ligase.
- 38. 2. Nucleotide excision repair • Nucleotide excision repair is ideally suited for large- scale defects in DNA. • The process is activated when a bulky lesion has been produced, such as DNA damage due to ultraviolet light, ionizing radiation and other environmental factors, often results in the modification of certain bases, strand breaks, cross-linkages etc. • After the identification of the defective piece of DNA, the DNA double helix is unwound to expose the damaged part. • An excision nuclease (exinuclease) cuts the DNA on either side ( upstream and downstream) of the damaged DNA. This defective piece is degraded. • The gap created by the nucleotide excision is filled up by the DNA polymerase which gets ligated by DNA ligase. • Xeroderma pigmentosum (XP) is a rare autosomal recessive disease. The affected patients are photosensitive and susceptible to skin cancers. It is now recognized that XP is due to a defect in the nucleotide excision repair of the damaged DNA.
- 39. 3. Mismatch repair • Despite high accuracy in replication, defects do occur when the DNA is copied. • For instance, cytosine (instead of thymine) could be incorporated opposite to adenine. • Mismatch repair corrects a single mismatch base pair e.g. C to A, instead of T to A. • The template strand of the DNA exists in a methylated form, while the newly synthesized strand is not methylated. • This difference allows the recognition of the new strands. • The enzyme GATC endonuclease cuts the strand at adjacent methylated GATC sequence. • This is followed by an exonuclease digestion of the defective strand, and thus its removal. • A new strand is now synthesized to replace the damaged one. • Hereditary nonpolyposis colon cancer (HNPCC) is one of the most common inherited cancers. • This cancer is now linked with faulty mismatch repair of defective DNA.
- 40. Fig. A diagrammatic representation of mismatch repair of DNA
- 41. 4. Double-strand break repair • Double-strand breaks (DSBs) in DNA are dangerous. • They result in genetic recombination which may lead to chromosomal translocation, broken chromosomes, and finally cell death. • DSBs can be repaired by homologous recombination or non- homologous end joining. • Homologous recombination occurs in yeasts while in mammals, non-homologous and joining dominates.
- 43. RECOMBINATION INTRODUCTION • Recombination basically involves the exchange of genetic information. • There are mainly two types of recombination. 1. Homologous recombination 2. Non-homologous recombination 1. Homologous recombination • This is also called as general recombination, and occurs between identical or nearly identical chromosomes (DNA sequences). • The best example is the recombination between the paternal and maternal chromosomal pairs. • It is a known fact that the chromosomes are not passed on intact from generation to generation. • Instead, they are inherited from both the parents. • This is possible due to homologous recombination.
- 44. • Three models have been put forth to explain homologous recombination. 1. Holliday model • Holliday model is the simplest among the homologous recombination. • The two homologous chromosomes come closer, get properly aligned, and form single-strand breaks. • This results in two aligned DNA duplexes. • Now, the strands of each duplex partly unwind and invade in the opposite direction to form a two strands cross between the DNA molecules. • There occurs simultaneous unwinding and rewinding of the duplexes in such a way that there is no net change in the amount of base pairing, but the position of crossover moves. This phenomenon referred to as branch migration, results in the formation of heteroduplex DNA. • The enzyme DNA ligase seals the nick. • The two DNA duplexes (4 strands of DNA), joined by a single crossover point can rotate to create a four stranded Holliday junction. • Now the DNA molecules are subjected to symmetrical cuts in either of the two directions, and the cut ends are resealed by ligase.
- 45. • The DNA exchange is determined by the direction of the cuts, which would be horizontal or vertical. • If the cross strands are cut horizontally (cut 1), the flanking genes 9or markers, i.e. Ab/ab) remain intact, and no recombination occurs. • On the other hand, if the parental strands are cut vertically (cut 2), the flanking genes get exchanged (i.e. Ab/aB) due to recombination.
- 47. 2. Non-homologous recombination • The recombination process without any special homologous sequences of DNA is regarded as non-homologous recombination. TRANSPOSITION: • Transposition primarily involves the movement of specific pieces of DNA in the genome. • The mobile segments of DNA are called transposons or transposable elements. • They were first discovered by Barbara McClintock in maize. • Transposons are mobile and can move almost to any place in the target chromosome. • There are two modes of transposition. One that involves an RNA intermediate, and the other which does not involve RNA intermediate. • Retrotransposition: • Transposition involving RNA intermediate represents retrotransposition. • By the normal process of transcription, a copy of RNA is formed from a transposon. • Then by the enzyme reverse transcriptase, DNA is copied from the RNA.
- 48. • The newly formed DNA which is a copy of the transposon gets integrated into the genome. • This integration may occur randomly on the same chromosome or, on a different chromosome. • As a result of the retrotransposition, there are now two copies of the transposon, at different points on the genome. • DNA transposition: • Some transposons are capable of direct transposition of DNA to DNA. • This may occur either by replicative transposition or conservative transposition. • Both the mechanisms require enzymes that are mostly coded by the genes within the transposons. • DNA transposons is less common than retrotransposition in case of eukaryotes. • However, in case of prokaryotes, DNA transposons are more important than RNA transposons.
- 49. • Significance of transposition: • A large fraction of human genome has resulted due to the accumulation of transposons. • Short interspersed elements (SINEs) are repeats of DNA sequences which are present in about 500,000 copies per haploid human genome e.g. Alu sequences. • Long interspersed elements (LINEs) are also repeated DNA sequences and are present in about 50,000 copies in the human genome e.g. L1 elements.
Editor's Notes
- Autosome : Any chromosome other than the sex (X and Y) chromosomes.
- Germ cell : a cell containing half the number of chromosomes of a somatic cell and able to unite with one from the opposite sex to form a new individual; a gamete. An embryonic cell with the potential of developing into a gamete. Somatic cell : any cell of a living organism other than the reproductive cells.
- Phenotypically : The observable physical or biochemical characteristics of an organism, as determined by both genetic makeup and environmental influences. The expression of a specific trait, such as stature or blood type, based on genetic and environmental influences.
