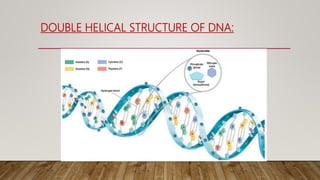
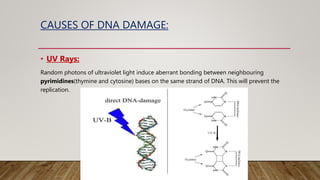
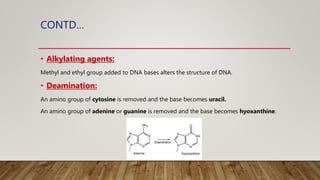
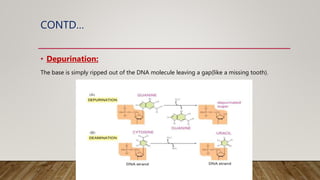
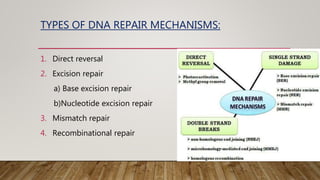
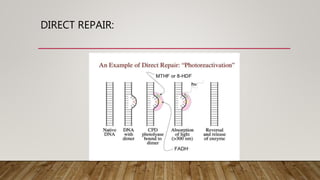
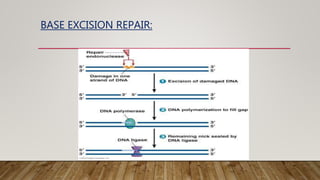
The document discusses DNA repair, emphasizing its critical role in maintaining genetic stability. It elaborates on the types of DNA damage, sources, causes, consequences, and various repair mechanisms, including direct repair, excision repair, mismatch repair, and recombinational repair. Ultimately, these processes are essential to prevent genome instability, cancer risk, and other diseases.