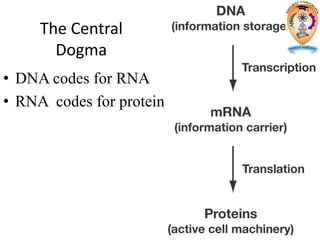
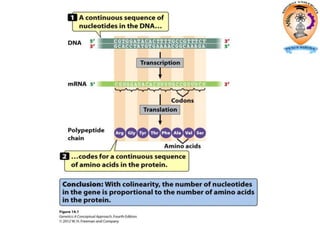
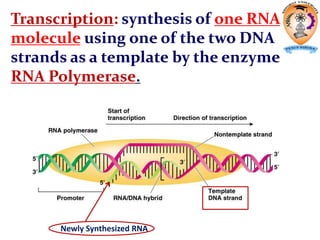
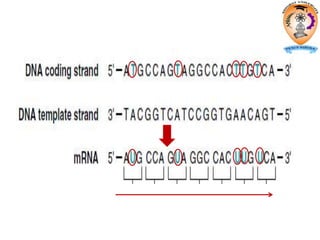
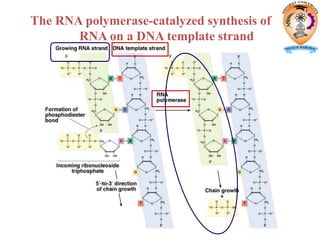
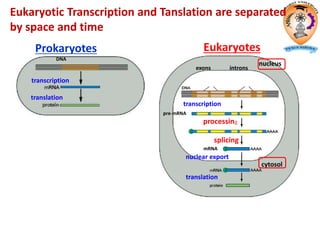
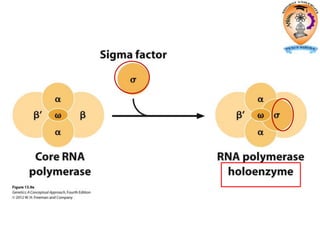
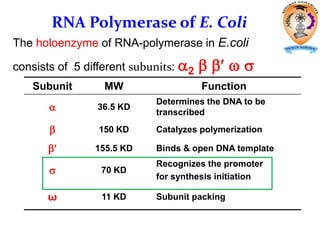
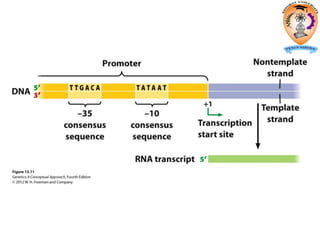
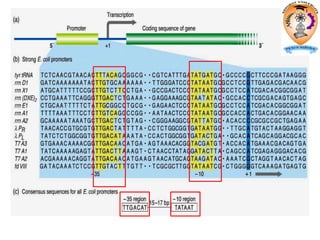
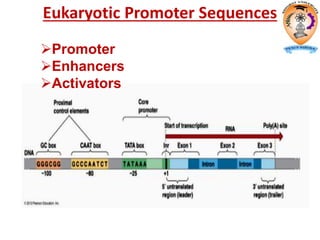
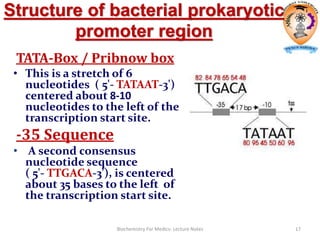
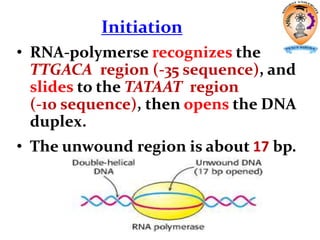
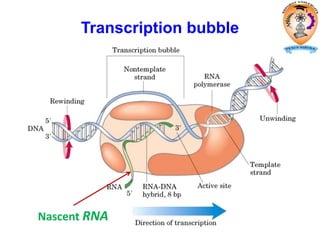
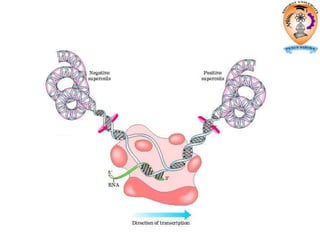
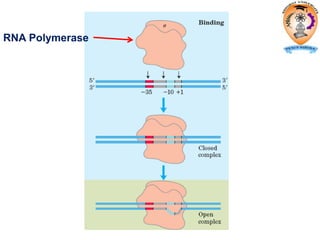
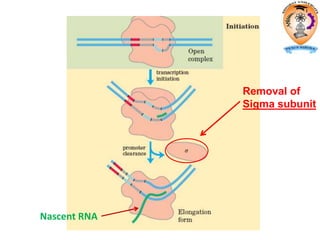
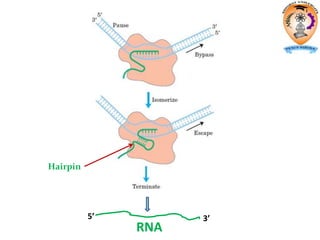
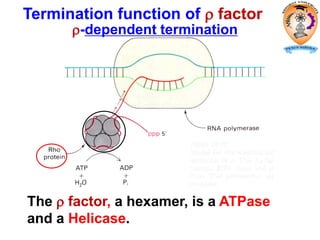
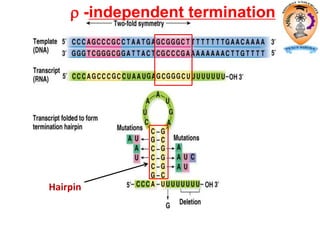
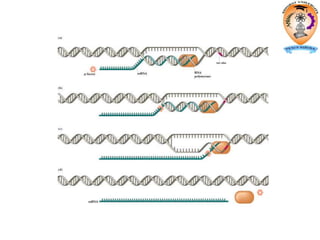
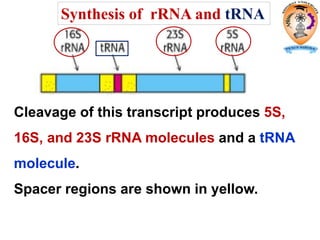
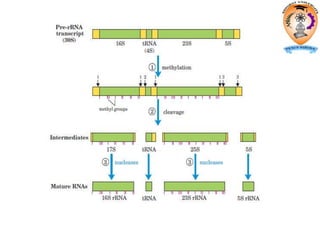
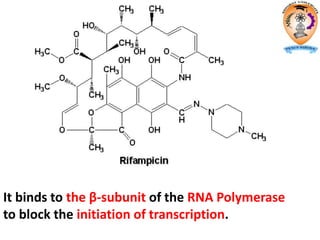
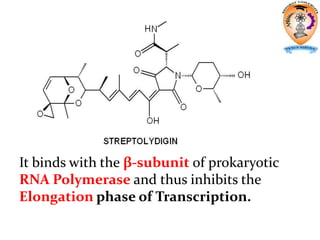
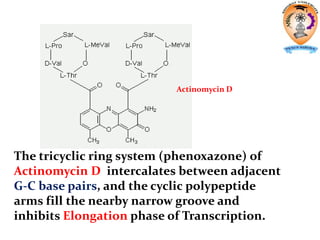
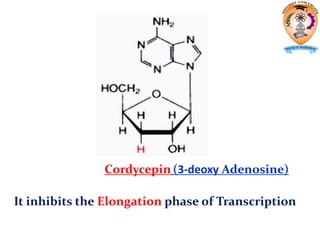
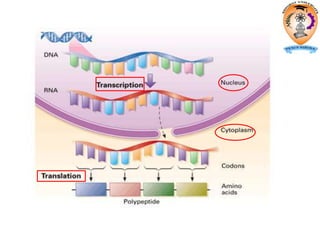
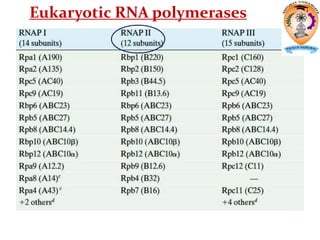
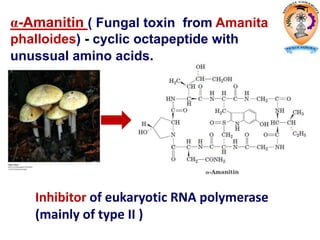
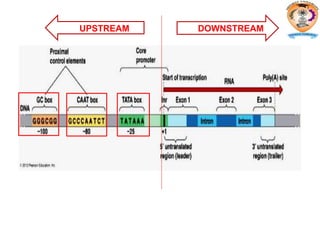
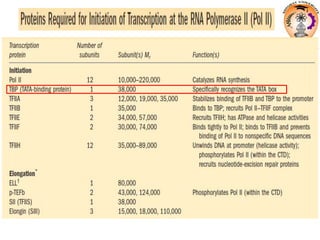
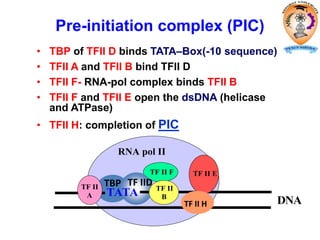
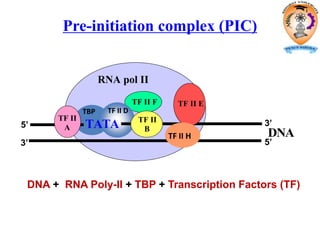
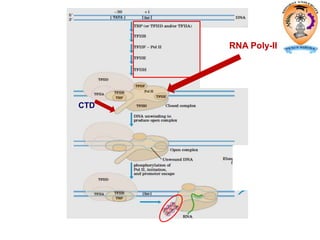
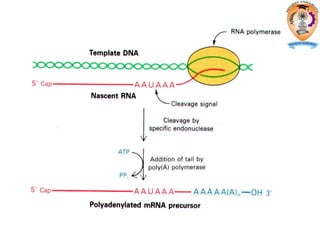
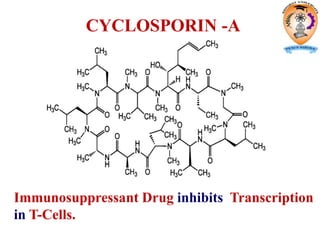
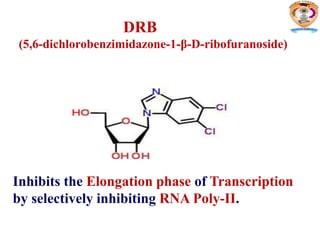
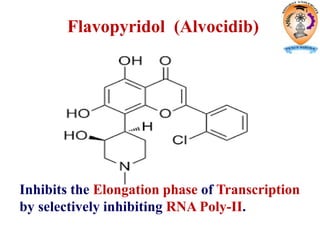
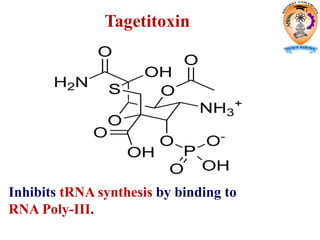
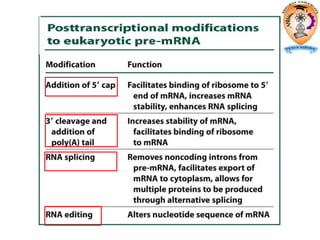
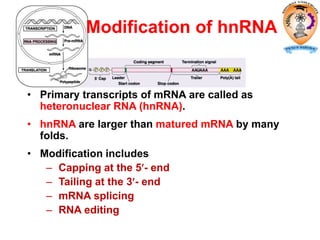
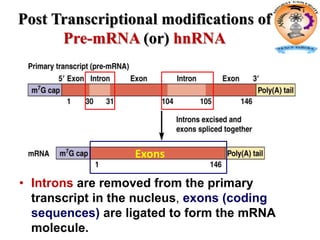
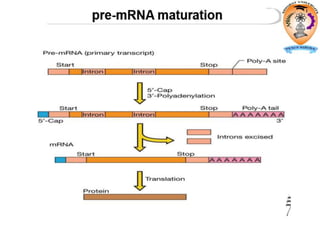
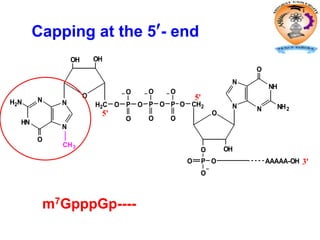
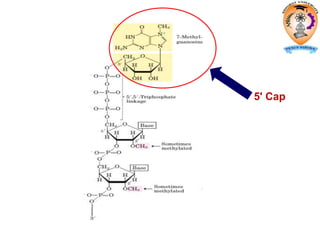
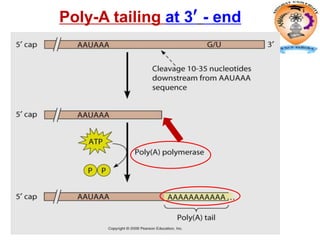
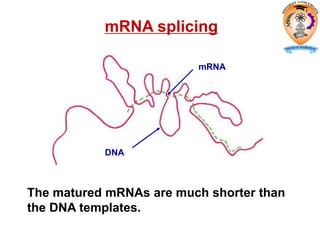
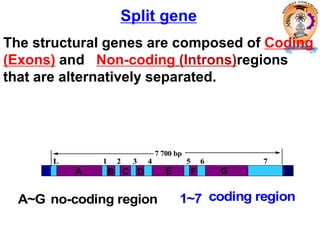
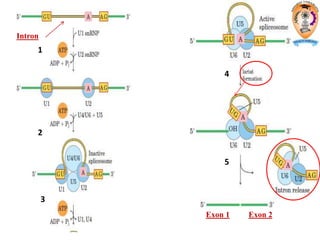
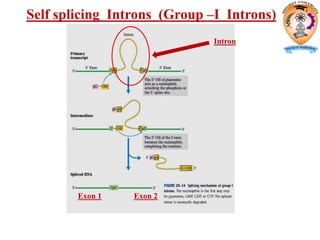
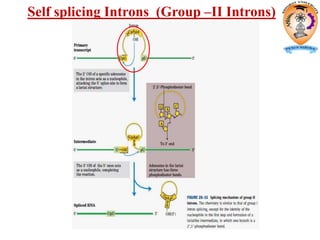
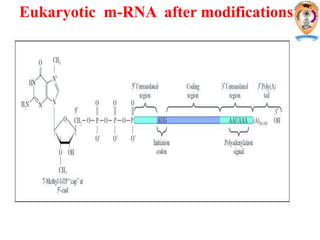
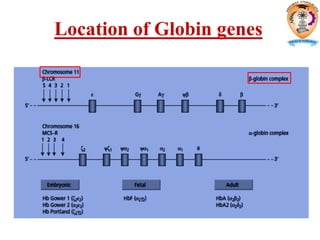
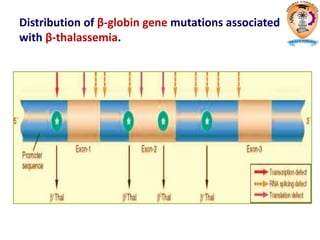
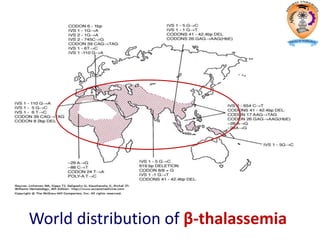
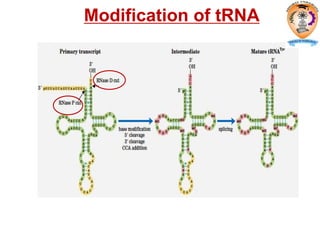
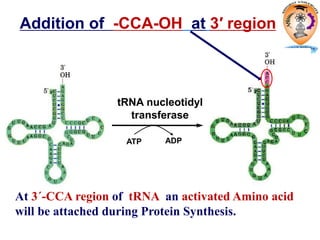
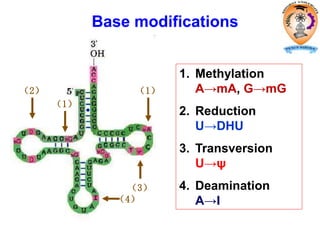
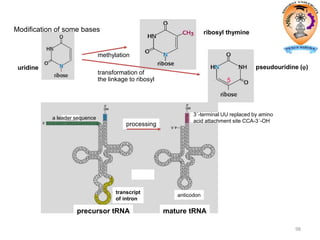
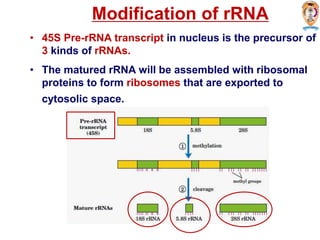
The document discusses transcription in prokaryotes and eukaryotes. In prokaryotes, RNA polymerase binds to promoter sequences and transcribes DNA into RNA through initiation, elongation, and termination. Transcription requires RNA polymerase and proceeds similarly in eukaryotes but involves multiple RNA polymerases and occurs in the nucleus. Eukaryotic transcription is more complex, utilizing regulatory sequences, transcription factors, and RNA processing to modify pre-mRNA into mature mRNA through splicing, capping, polyadenylation, and other modifications. Mutations can affect splicing and cause genetic disorders like beta-thalassemia.