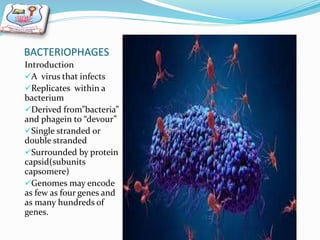
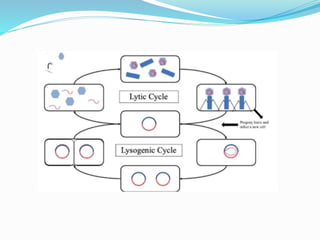
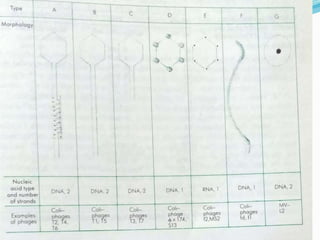
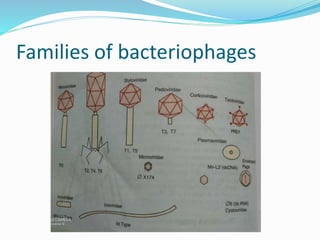
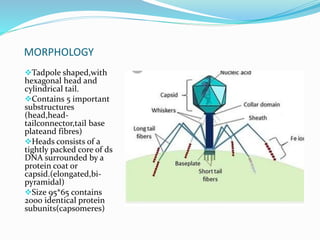
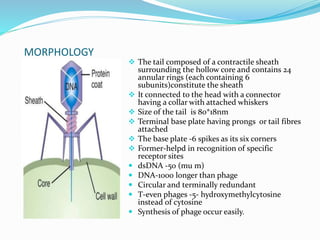
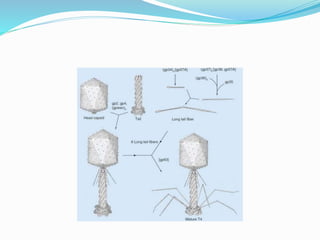
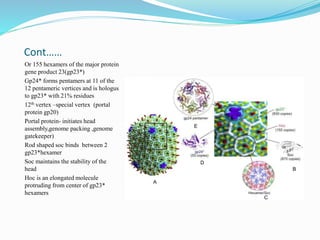
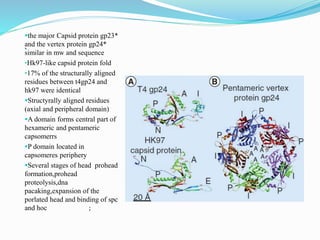
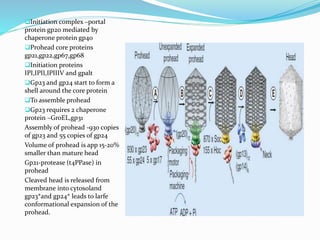
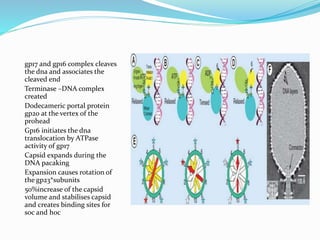
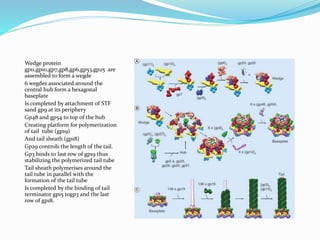
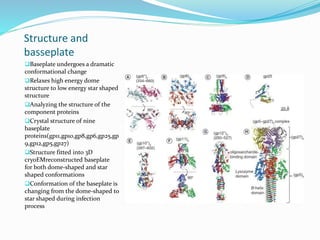
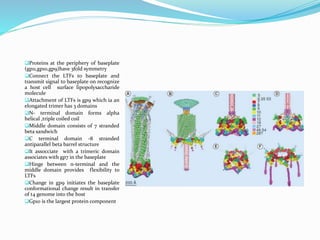
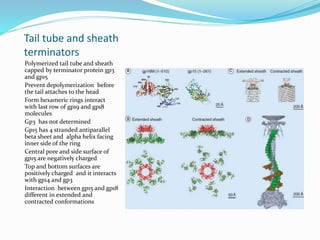
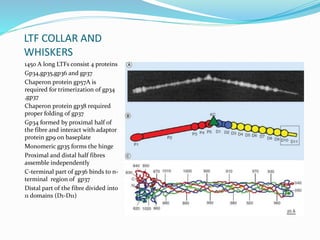
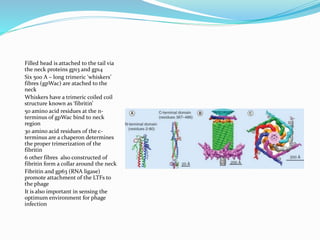
This document provides a history of bacteriophages and discusses their structure and classification. It notes that bacteriophages were discovered in the late 1890s and early 1900s and were named and more fully described between 1915-1917. It describes the basic tadpole shape of bacteriophages, consisting of a protein-encased DNA core surrounded by a hexagonal head and tail fibers. Bacteriophages are classified based on their morphology, nucleic acid composition, and genome size/genes. The document focuses on the T4 bacteriophage, detailing its structure, life cycle, and the proteins involved in head and tail assembly.