Protein structures determine their functions. There are four levels of protein structure:
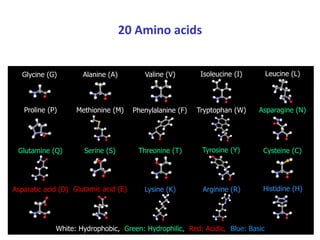
1) Primary structure is the amino acid sequence
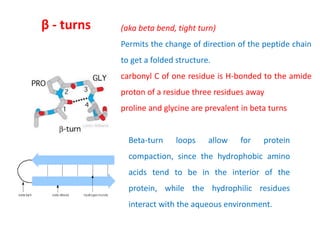
2) Secondary structure involves local patterns like alpha helices and beta sheets
3) Tertiary structure describes the overall 3D shape formed by secondary structures
4) Quaternary structure refers to the arrangement of multiple polypeptide chains
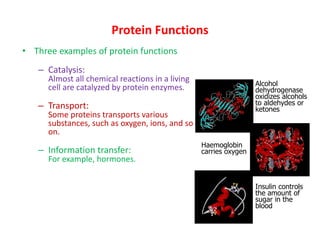
The most common secondary structures are alpha helices, stabilized by hydrogen bonds between amino acids i and i+4, and beta sheets formed by hydrogen bonding between strands. Protein structure enables functions like catalysis, transport, and information transfer.






















































![Fibrous proteins: Fibroin
Fibroin
Fibroins are the silk proteins. They also form the spider webs
Made with a -sheet structures with Gly on one face and
Ala/Ser on the other
Fibroins contain repeats of [Gly-Ala-Gly-Ala-Gly-Ser-Gly-Ala-
Ala-Gly-(Ser-Gly-Ala-Gly-Ala-Gly)8]
The -sheet structures stack on top of each other
Bulky regions with valine and tyrosine interrupt the -sheet
and allow the stretchiness](https://image.slidesharecdn.com/doc-20210601-wa0061-230718071330-a6838b77/85/Protein-Structure-60-320.jpg)







