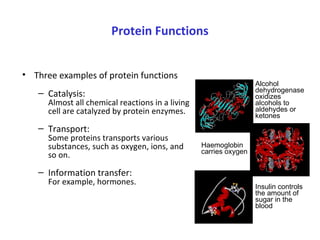
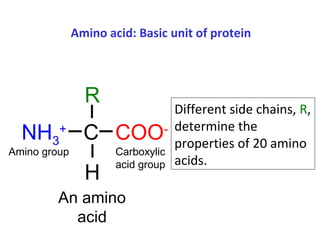
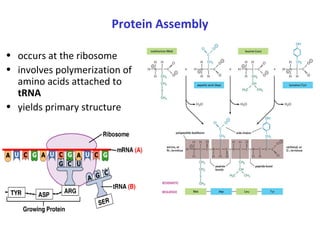
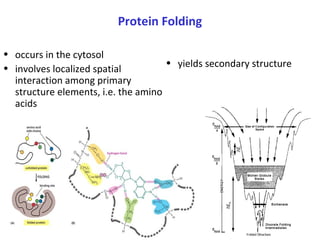
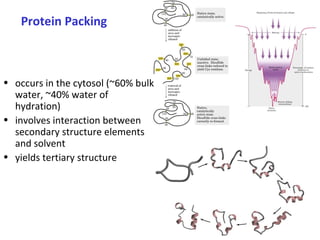
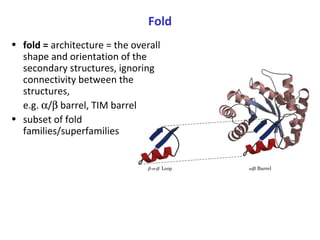
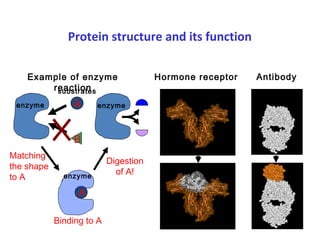
Proteins have three main functions: catalysis, transport, and information transfer. They are made of amino acids that polymerize and fold into complex three-dimensional structures, from primary to quaternary levels, that determine their unique functions. Protein structure and function are closely linked, as the structure allows proteins to bind specifically to other molecules and carry out reactions.