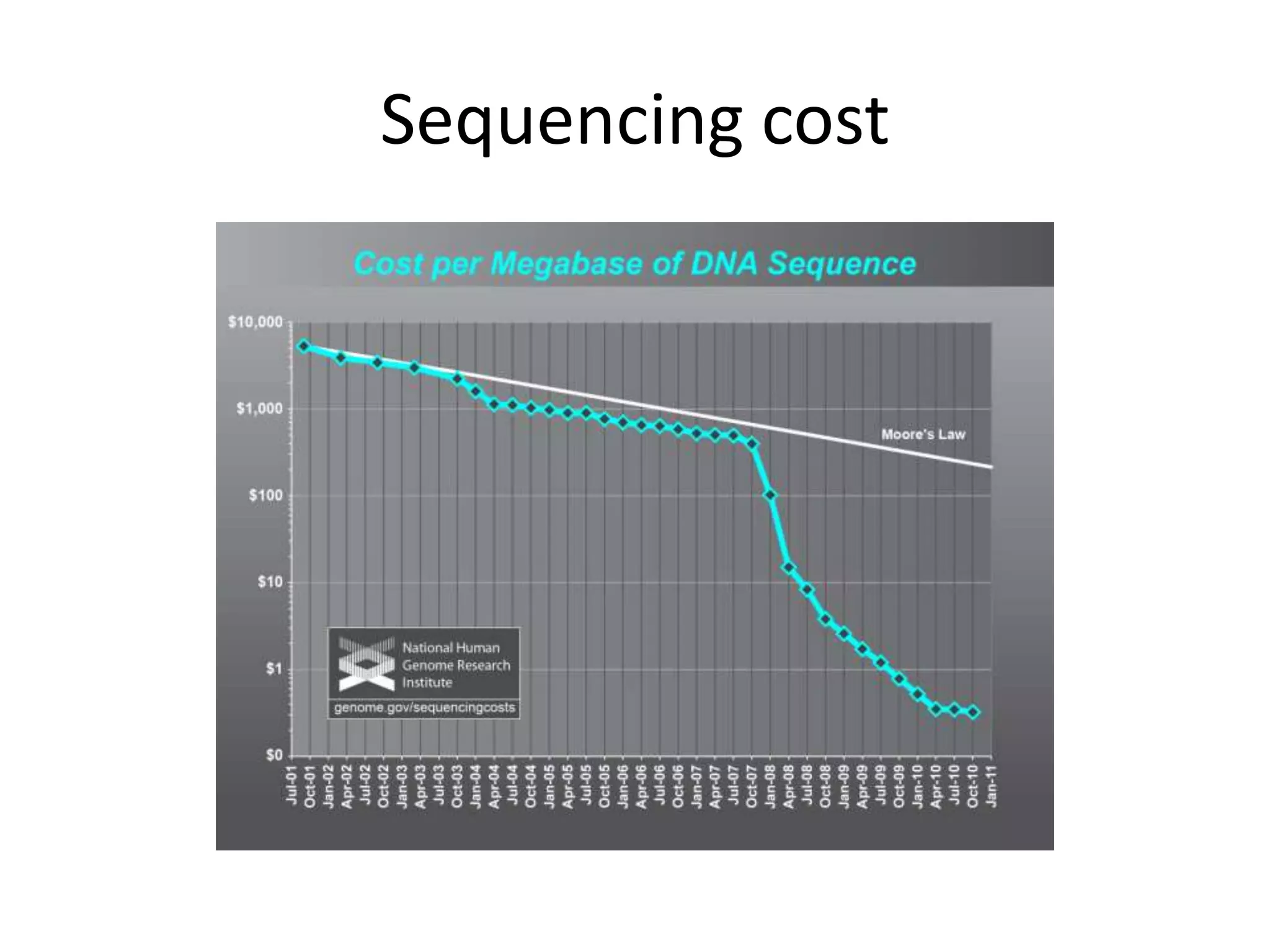
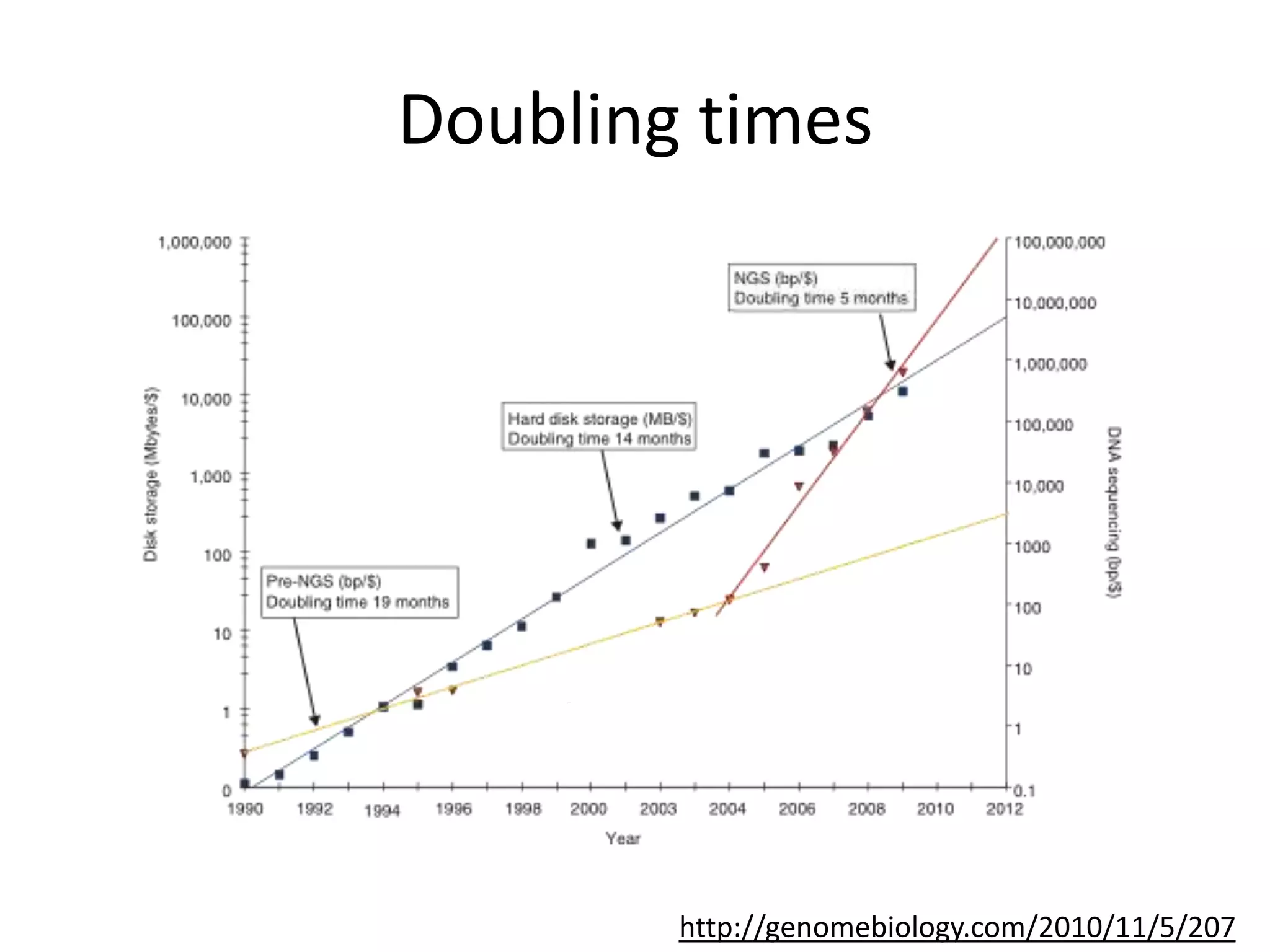
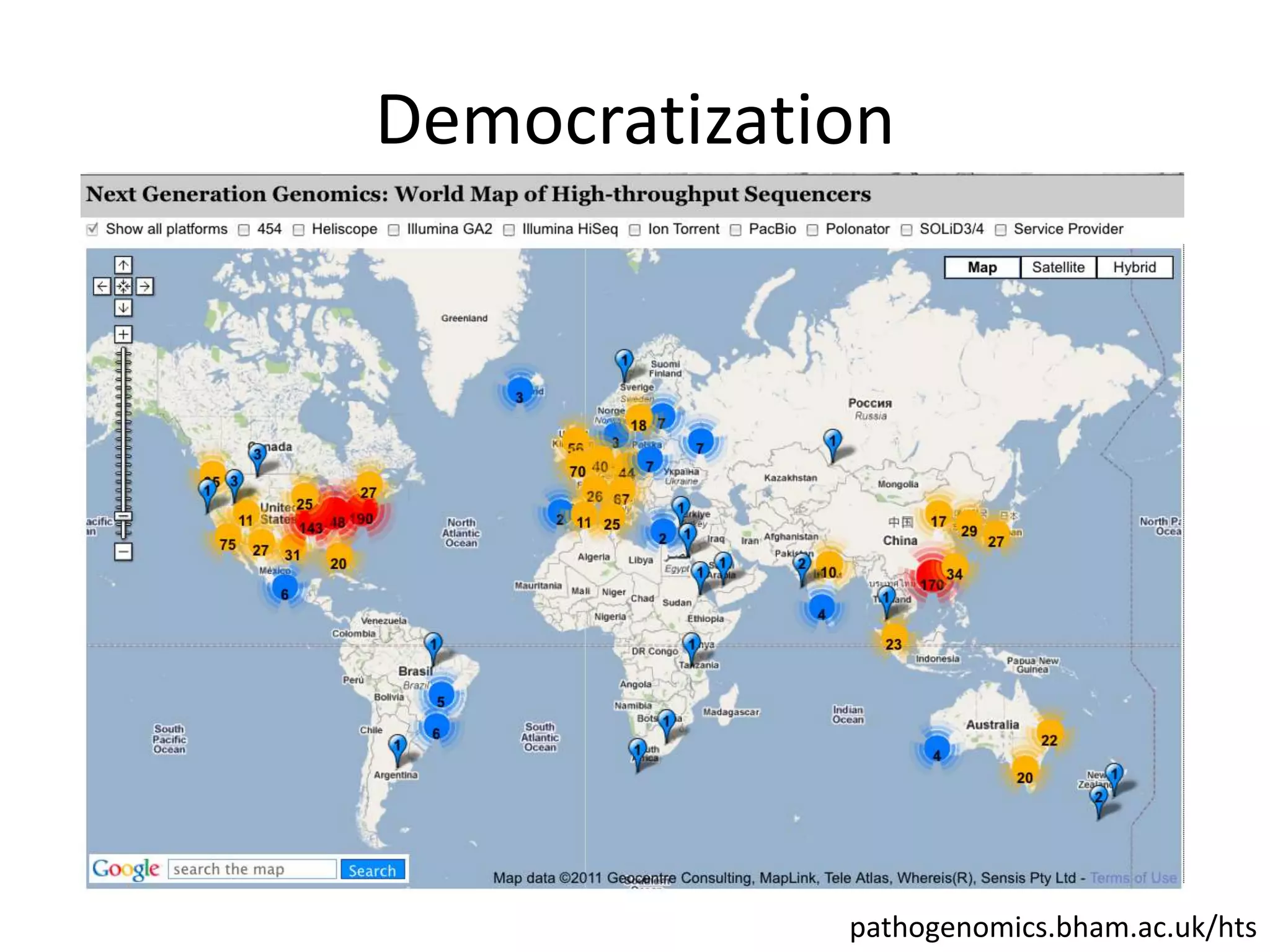
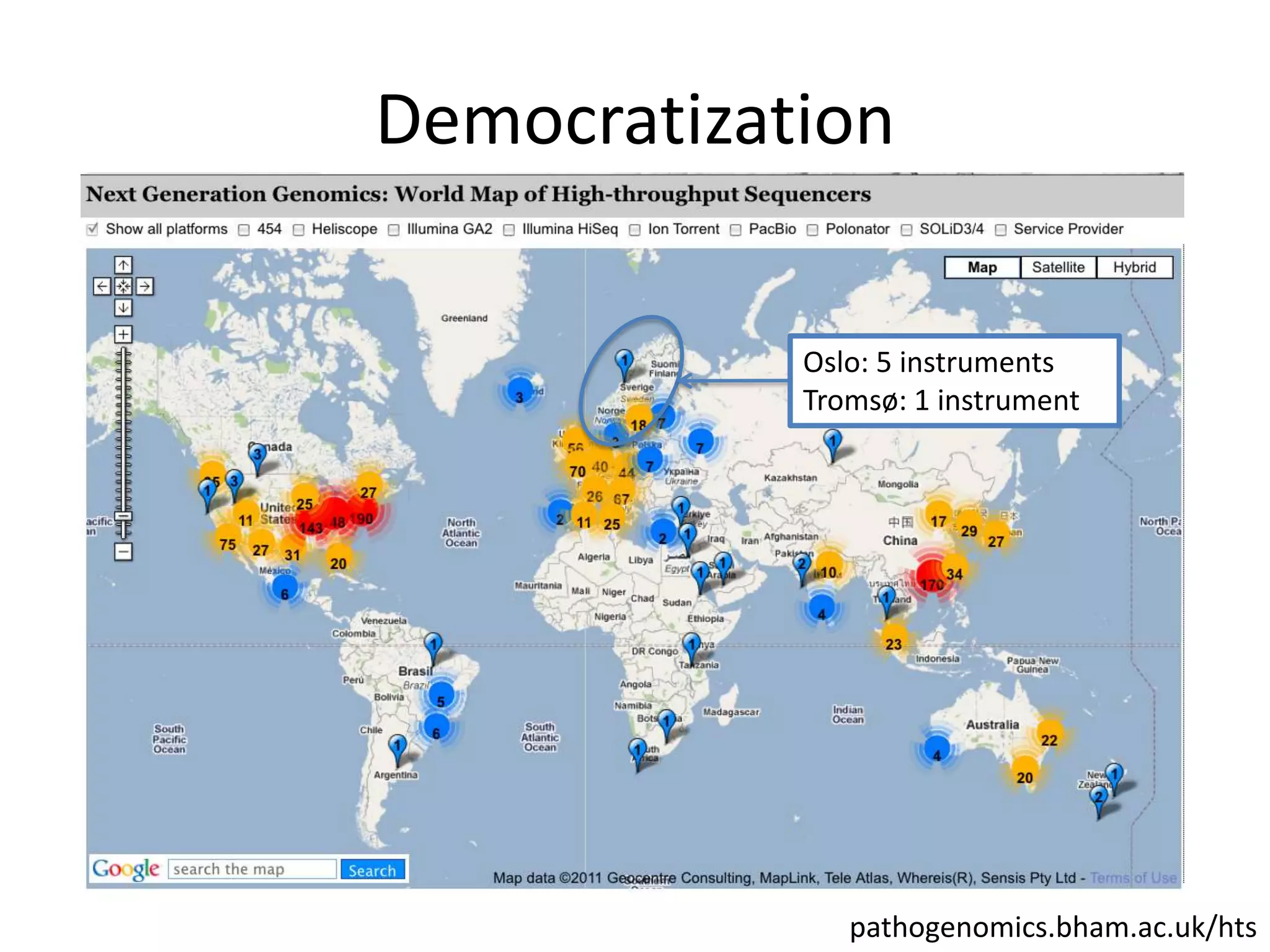
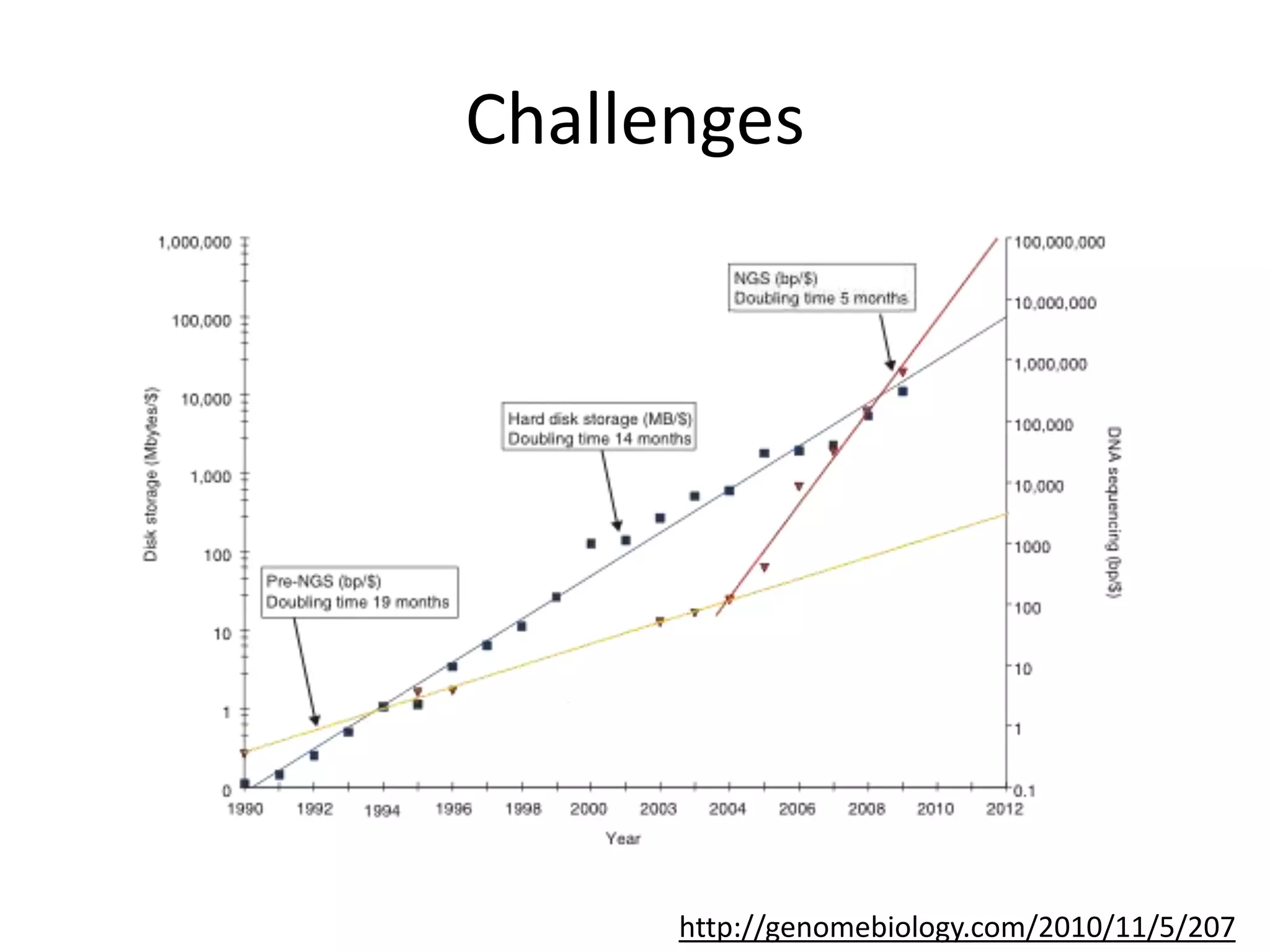
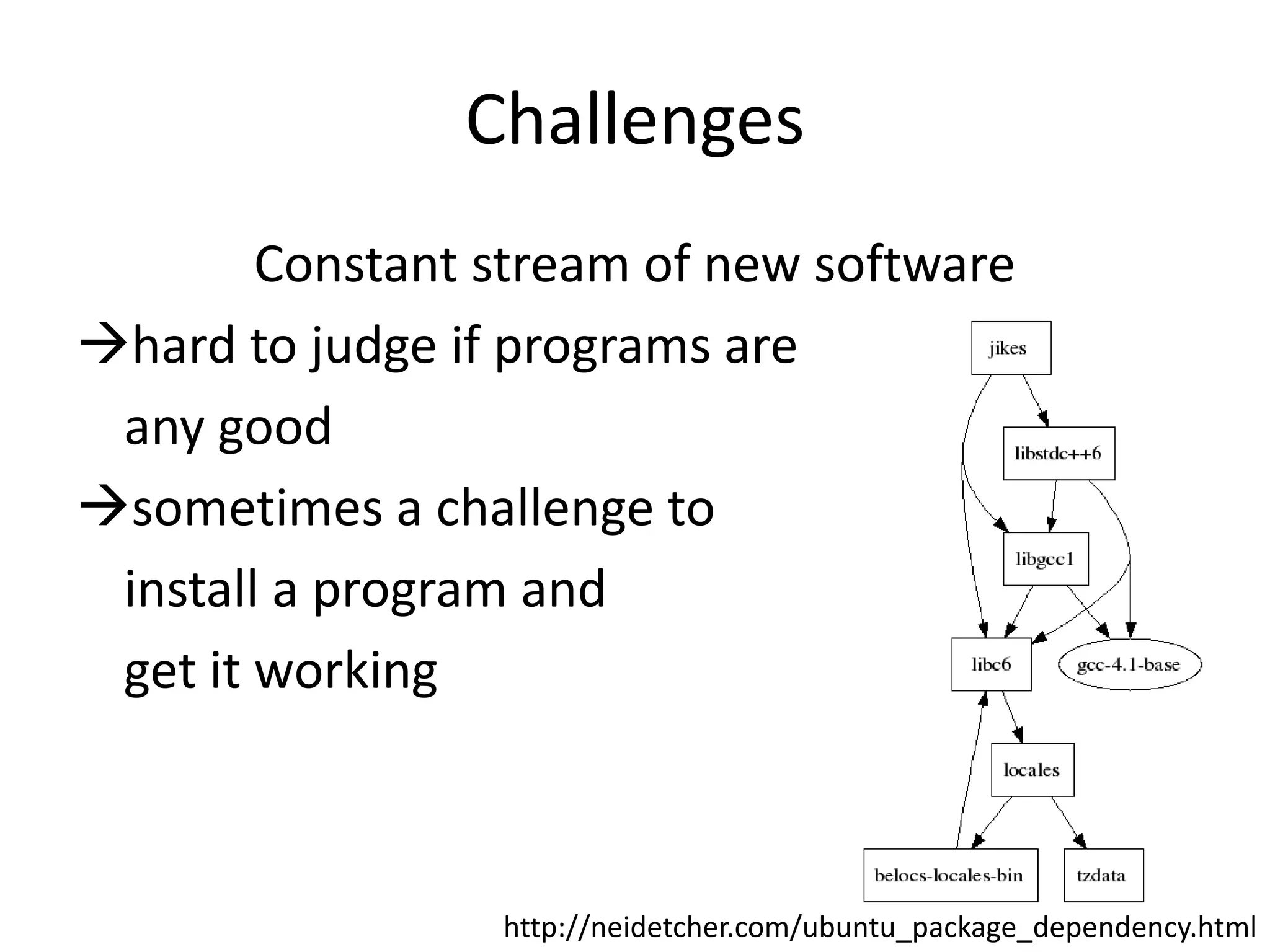
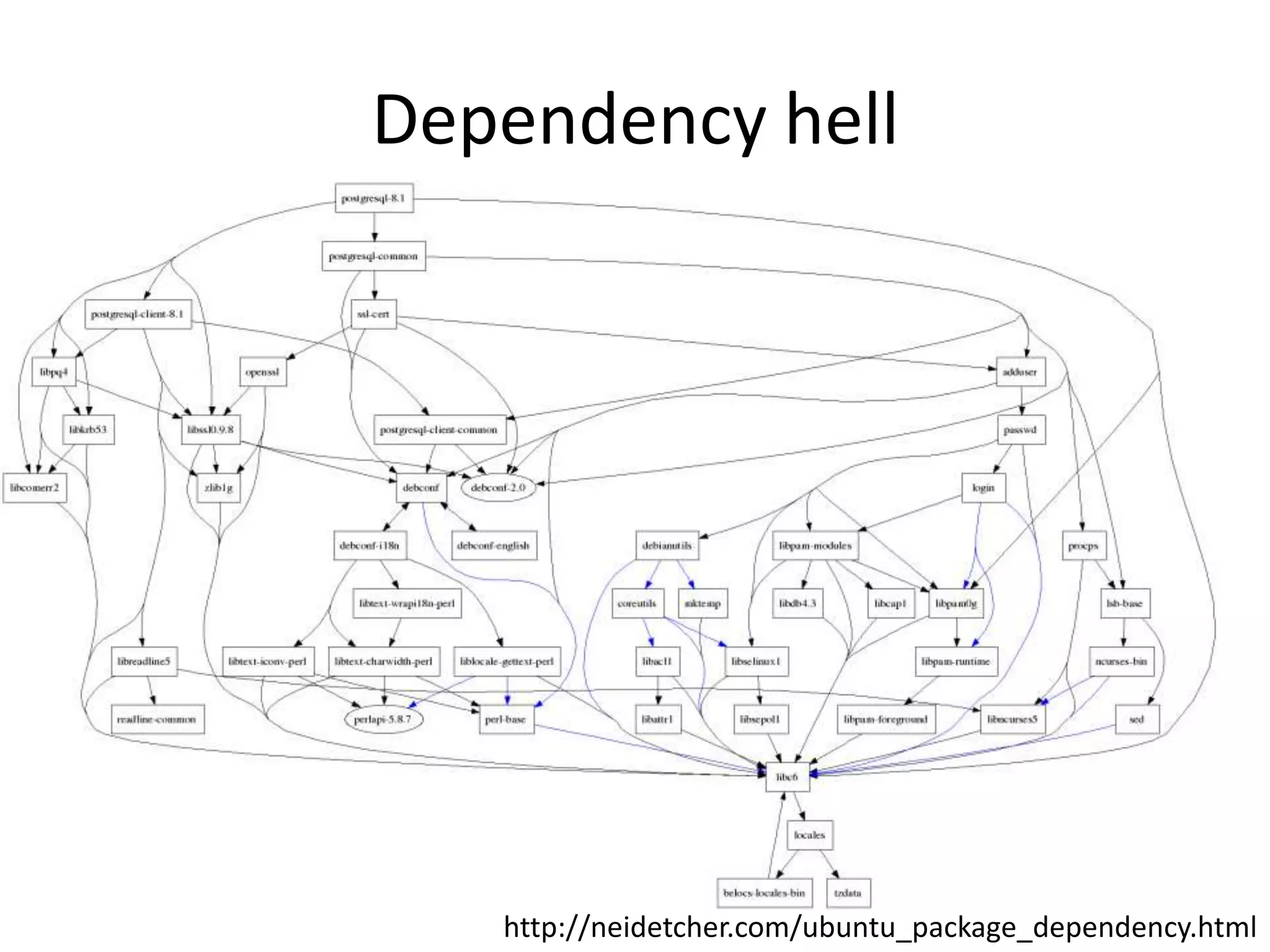
The document discusses the opportunities and challenges in next-generation sequencing (NGS) research, highlighting the need for affordable sequencing and the associated bioinformatics challenges. It emphasizes the importance of computational resources, training, and collaboration to effectively analyze the increasing data generated by NGS technologies. The document also suggests potential solutions like web-based programs and contributions to data standards.