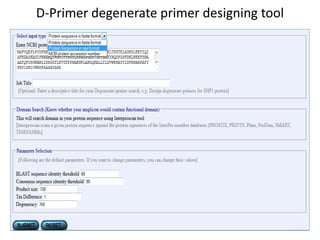
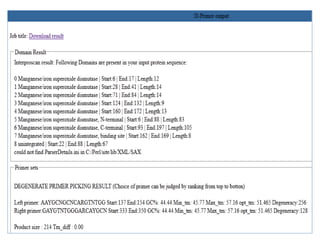
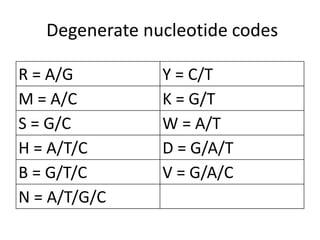
Degenerate primers are designed from conserved amino acid sequences aligned from multiple species. They contain nucleotide degeneracies that allow binding to related gene sequences. Key steps are:
1) Identifying conserved regions over 5+ amino acids long and within 200-600 bp for primers.
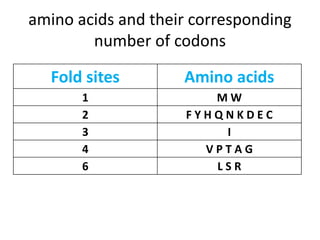
2) Calculating primer degeneracy based on contributing amino acids to minimize values over 64.
3) Optimizing primers by avoiding amino acids with 6-fold degeneracy and adding 5' tails.