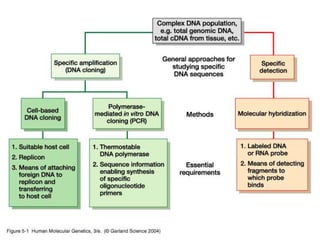
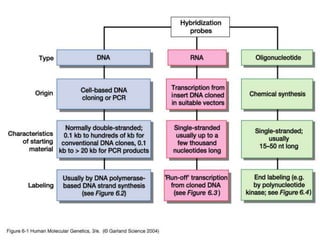
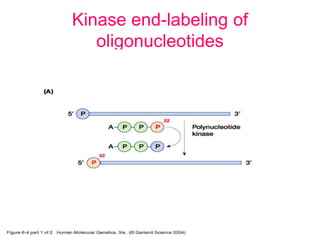
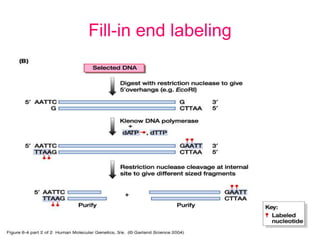
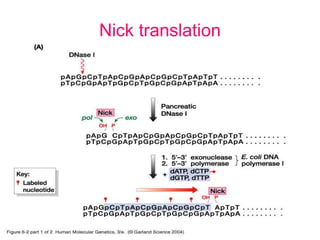
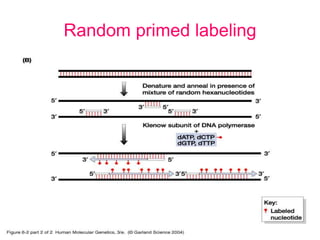
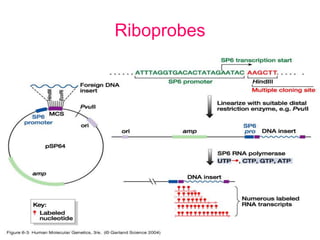
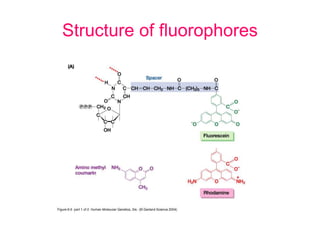
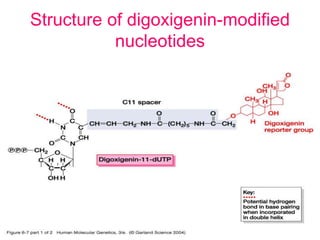
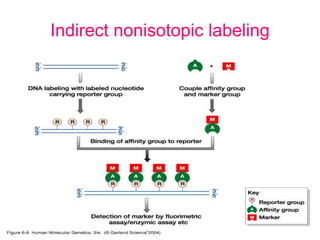
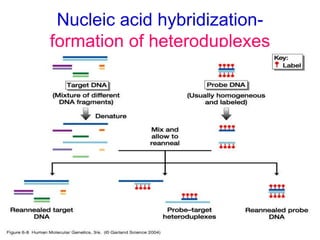
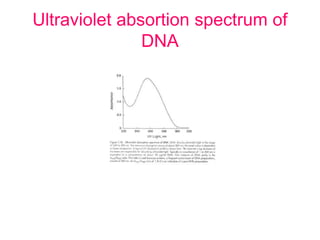
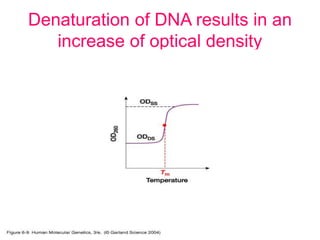
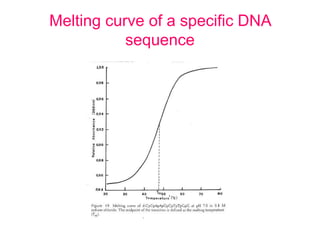
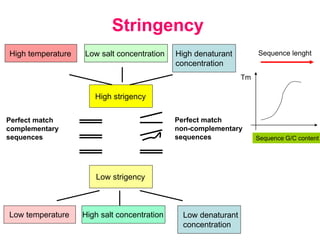
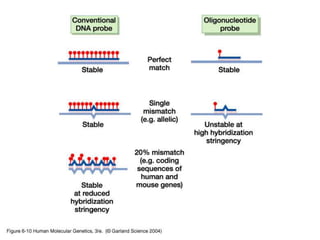
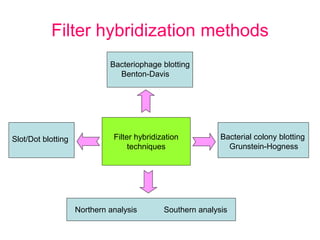
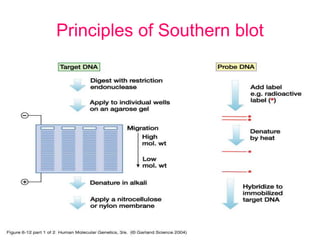
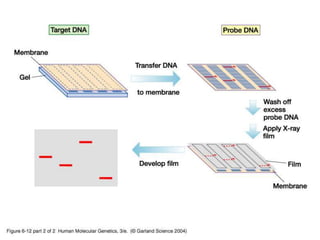
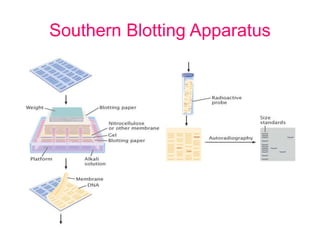
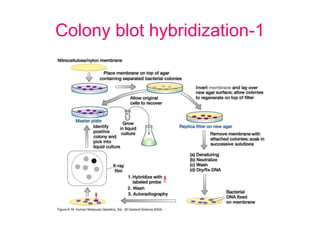
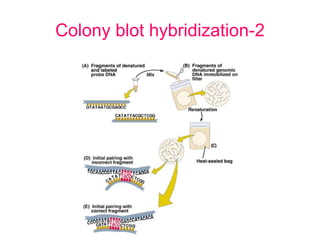
Nucleic acid hybridization uses labeled probes to identify related DNA or RNA molecules in a complex mixture. It relies on base complementarity between the probe and target molecules to form double-stranded hybrids. Probes can be radioactively or nonradioactively labeled. Hybridization is affected by factors like temperature, salt concentration, and mismatches. Southern blotting uses hybridization to detect specific DNA sequences separated by gel electrophoresis and transferred to membranes.