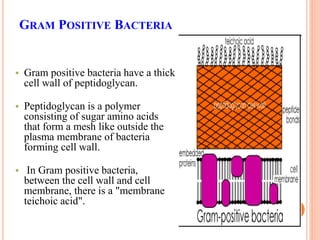
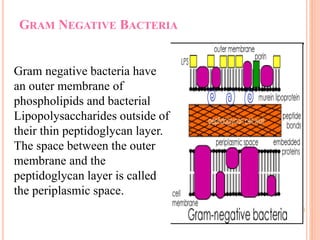
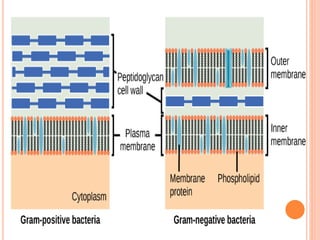
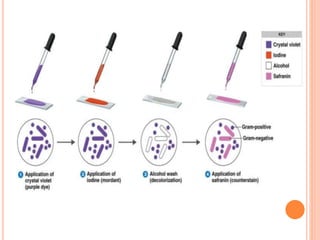
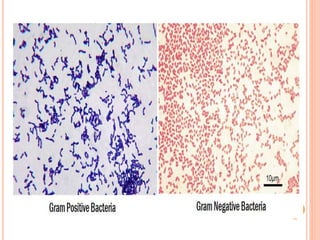
The document summarizes Gram staining, a method developed by Hans Christian Gram in 1883 to differentiate between bacterial species. Gram staining uses crystal violet dye and iodine to stain bacteria, then decolorizes them with acetone or alcohol. Gram-positive bacteria retain the crystal violet dye after decolorization due to their thick peptidoglycan cell wall, appearing purple or blue. Gram-negative bacteria's thinner cell wall is unable to retain the dye after decolorization but can be counterstained pink with safranin. This differentiation depends on differences in bacterial cell wall composition and structure.