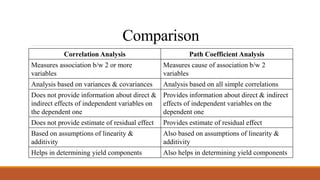
1. Association analysis includes correlation coefficient analysis and path coefficient analysis to study the relationship between two or more variables.
2. Correlation coefficient analysis measures the degree and direction of association between variables on a scale of -1 to 1, where 1 is total positive correlation and -1 is total negative correlation.
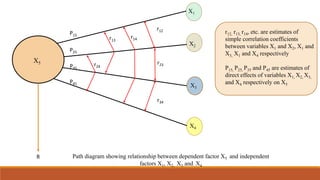
3. Path coefficient analysis splits the correlation coefficients into measures of direct and indirect effects to determine the direct and indirect contribution of independent variables to a dependent variable like yield.