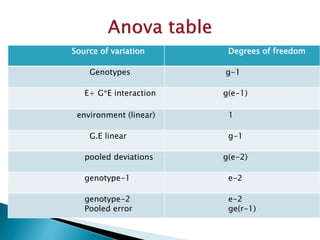
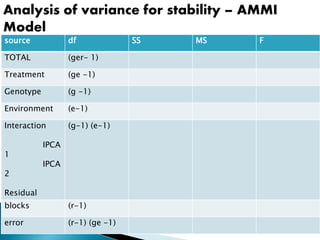
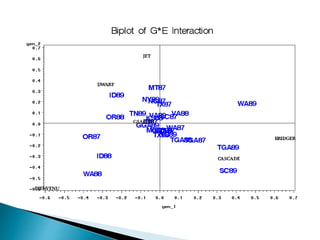
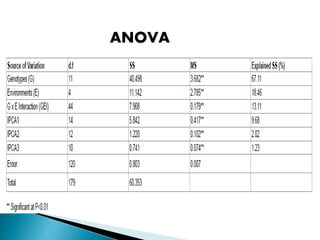
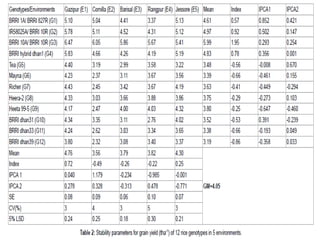
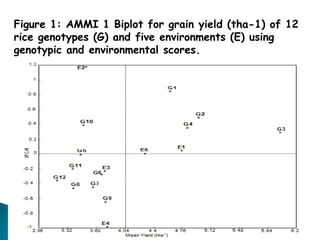
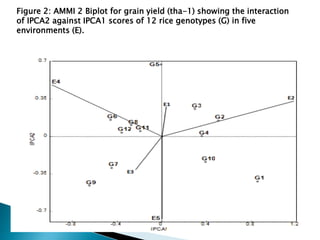
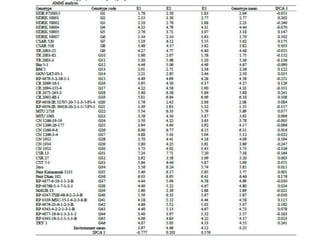
The document discusses the interaction between genotype and environment affecting phenotype, emphasizing the importance of genotype x environmental interaction (GxE) in crop performance. It details various models for analyzing GxE interactions, notably the AMMI model, which combines ANOVA and principal components analysis to assess genotype stability across different environments. The findings include experiments conducted on rice genotypes in varying agro-ecological zones and reveal significant GxE interaction, underscoring the need for stable genotypes adaptable to diverse conditions.