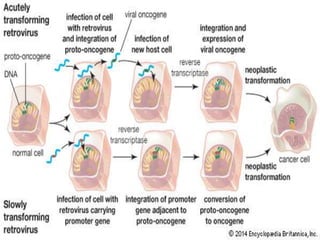
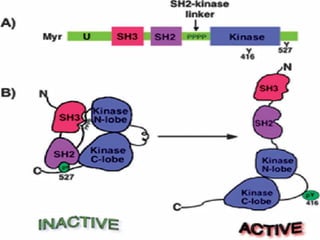
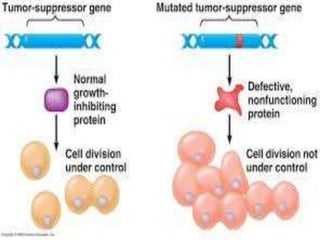
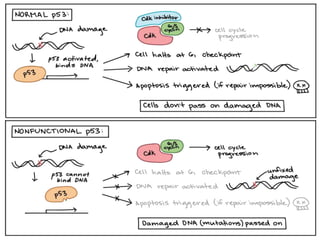
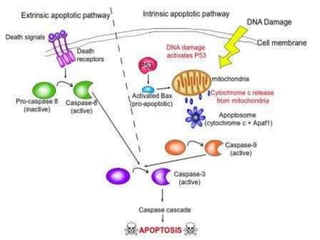
This document discusses proto-oncogenes, oncogenes, and tumor suppressor genes. Proto-oncogenes are normal genes that regulate cell growth, while oncogenes are mutated proto-oncogenes that contribute to uncontrolled cell growth. The discovery of oncogenes began with Rous sarcoma virus, which was found to contain the viral oncogene v-src. The tumor suppressor gene p53 plays a key role in DNA repair and apoptosis, preventing cancer formation. When mutated, p53 cannot regulate the cell cycle properly, allowing damaged DNA to be passed on and tumors to form.