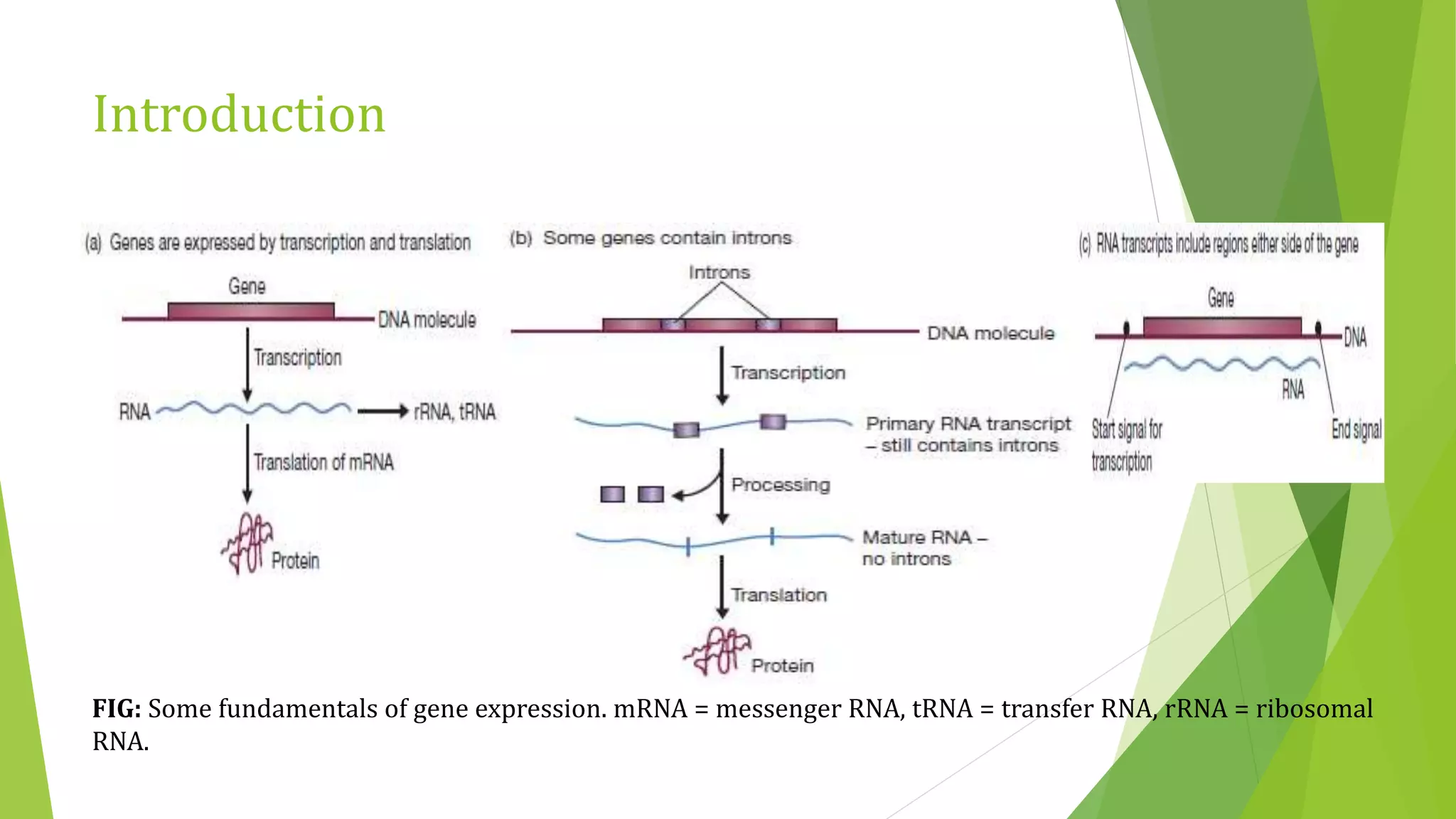
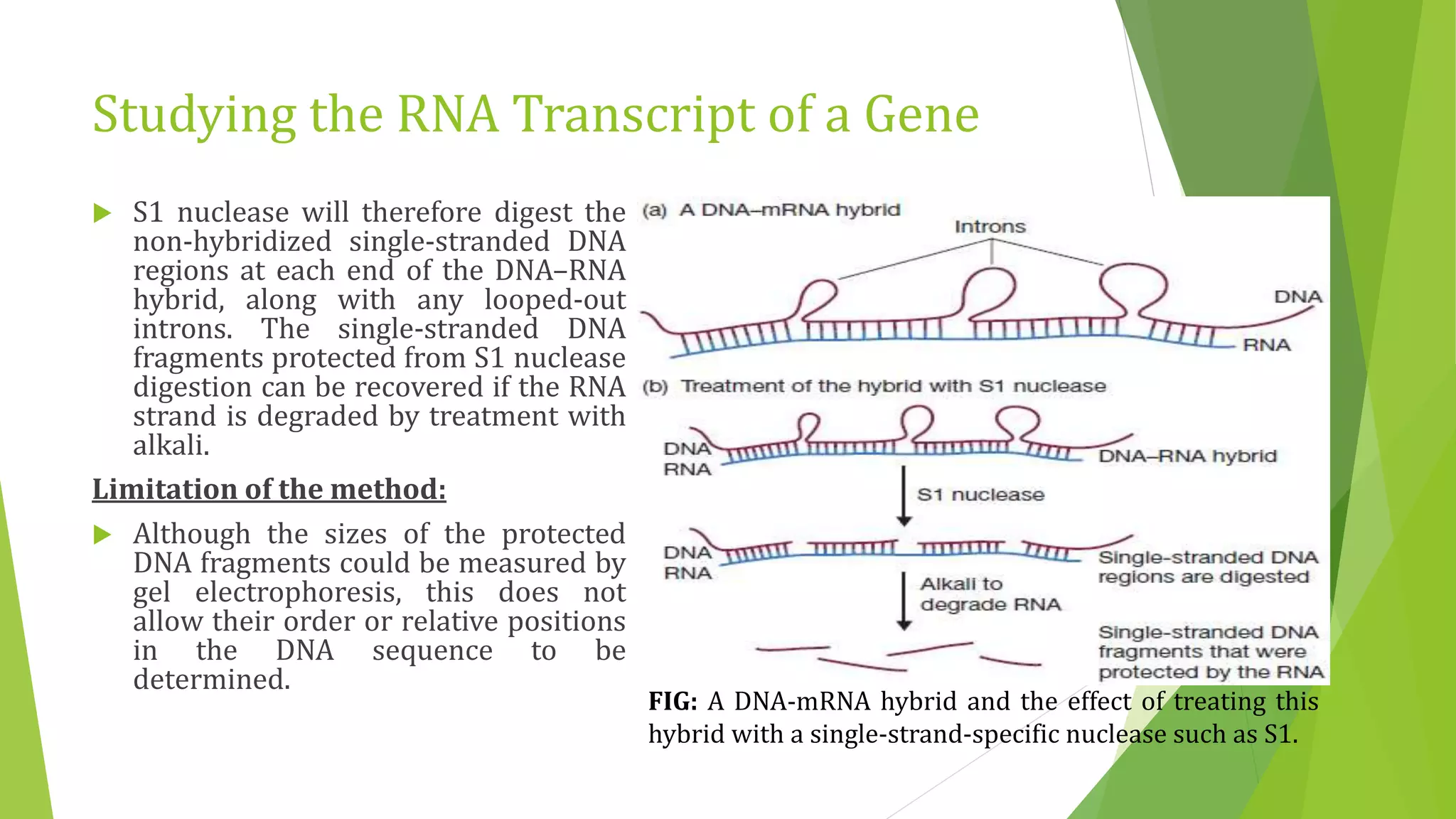
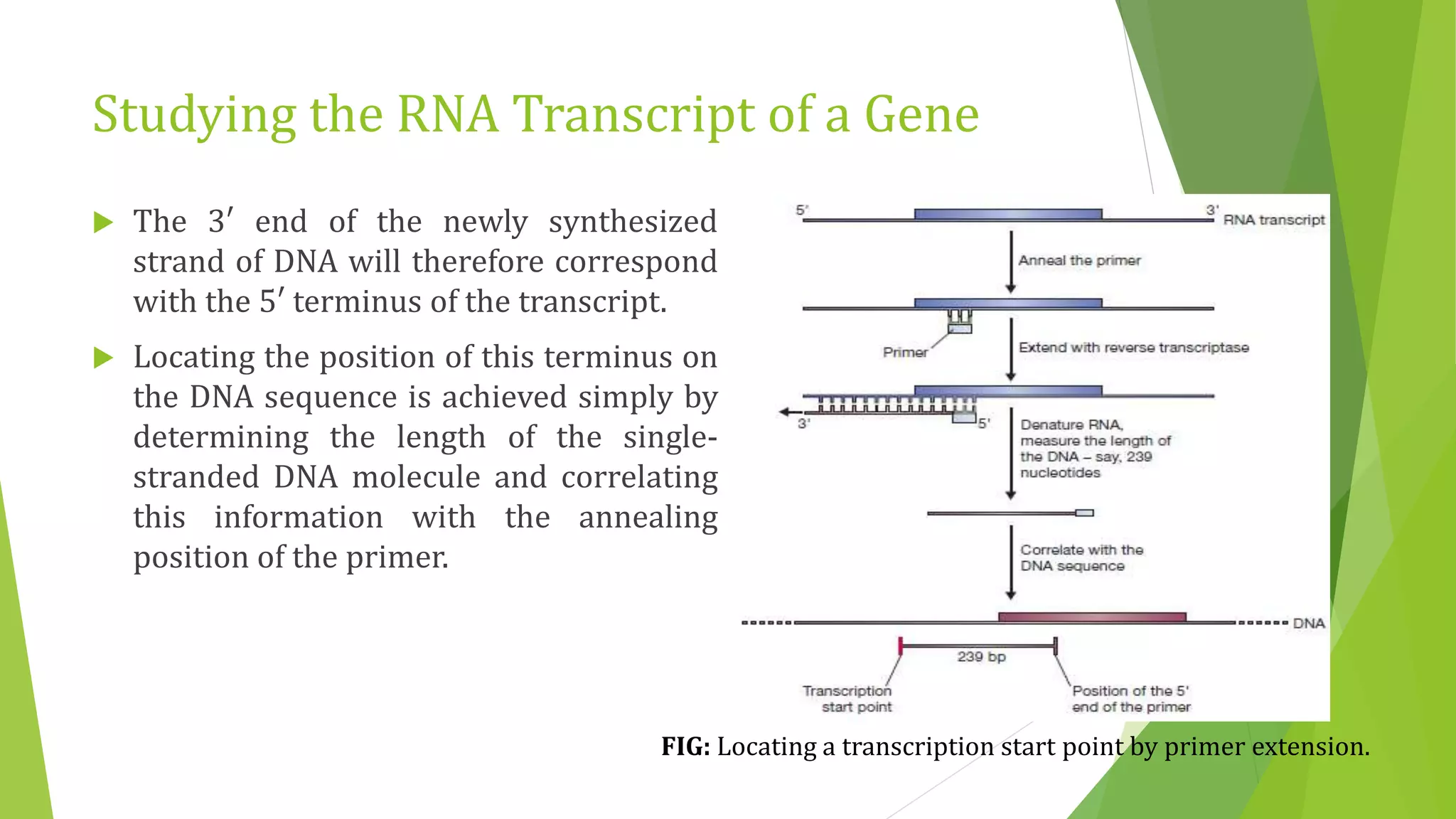
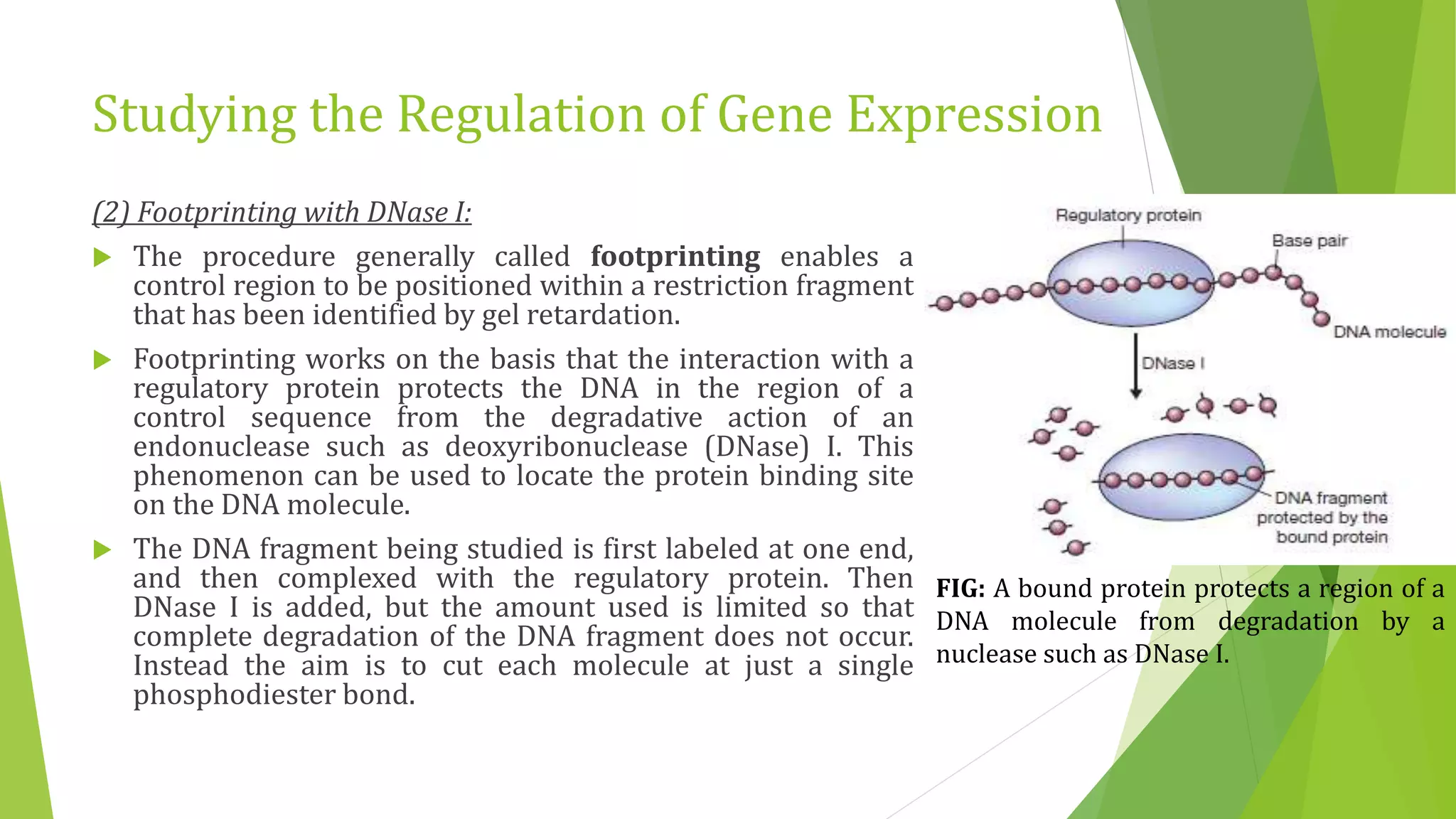
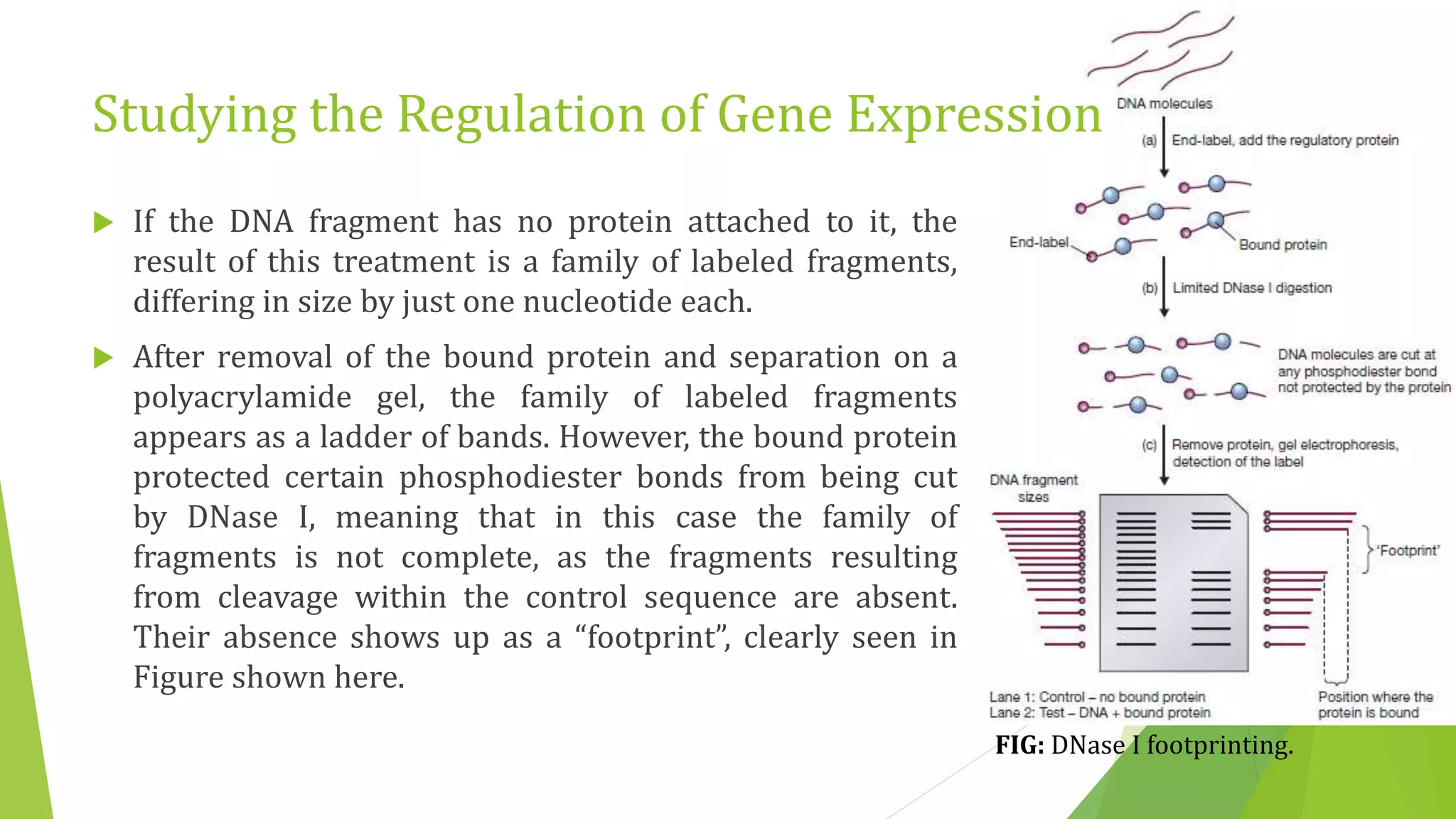
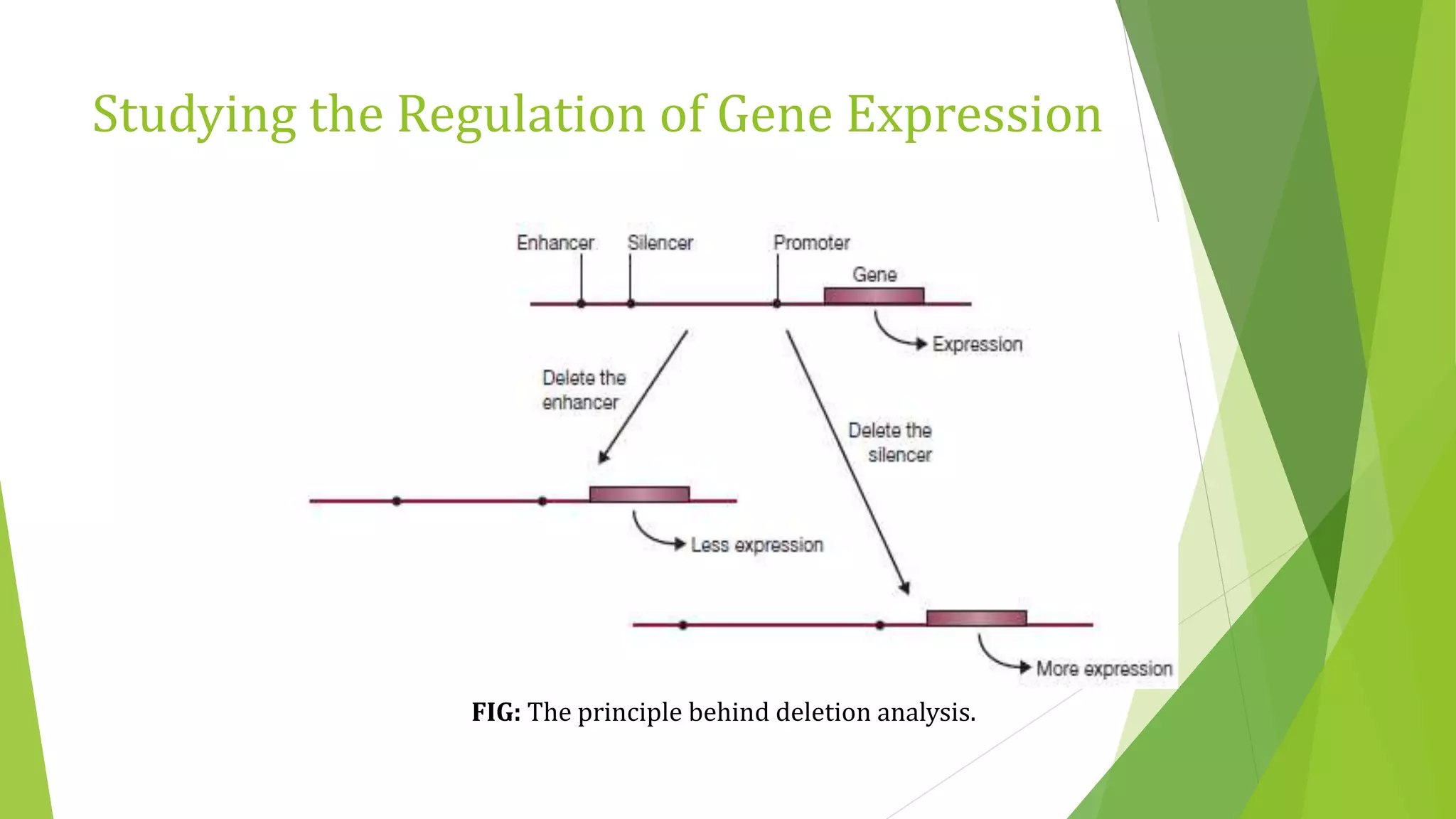
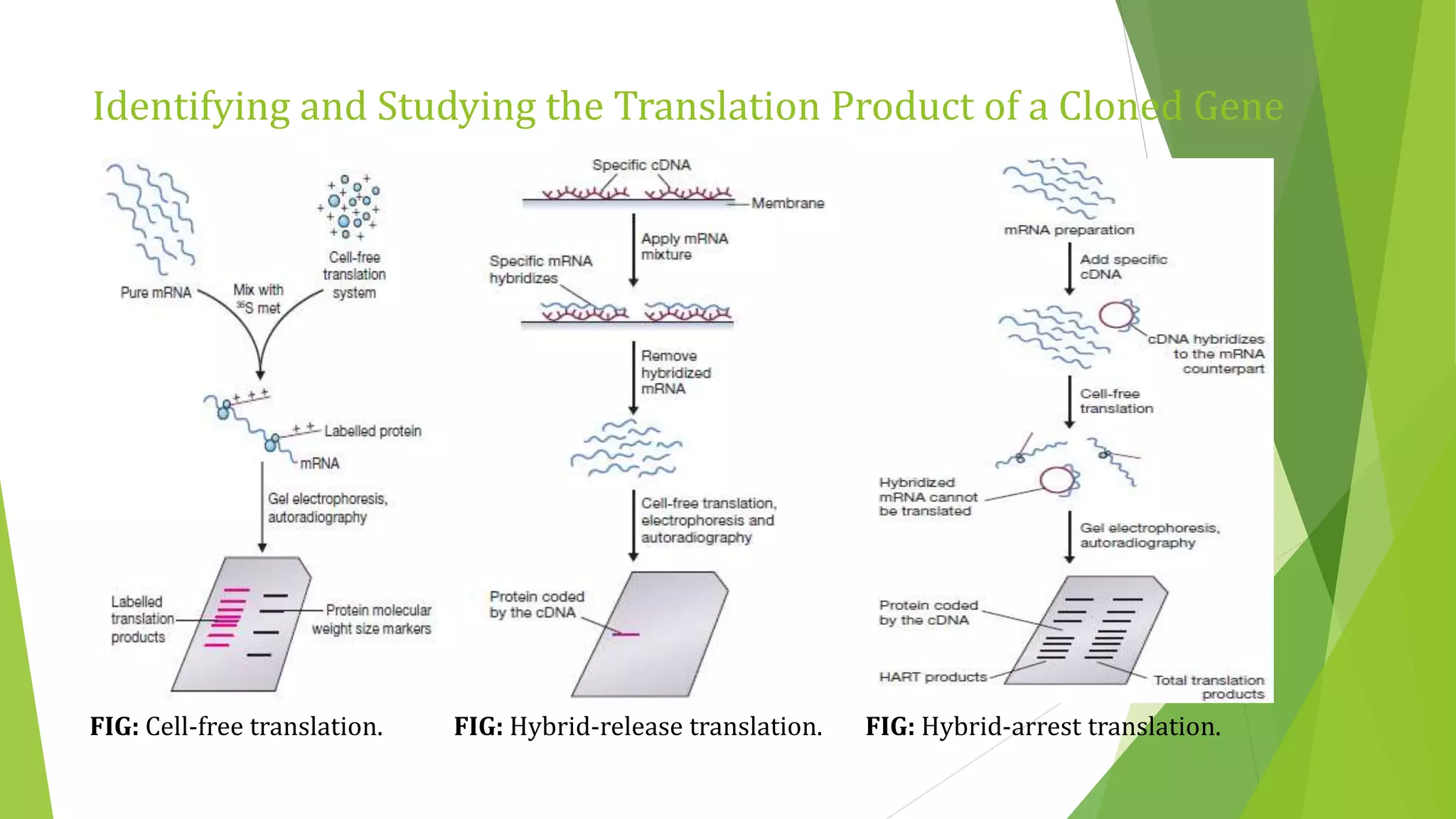
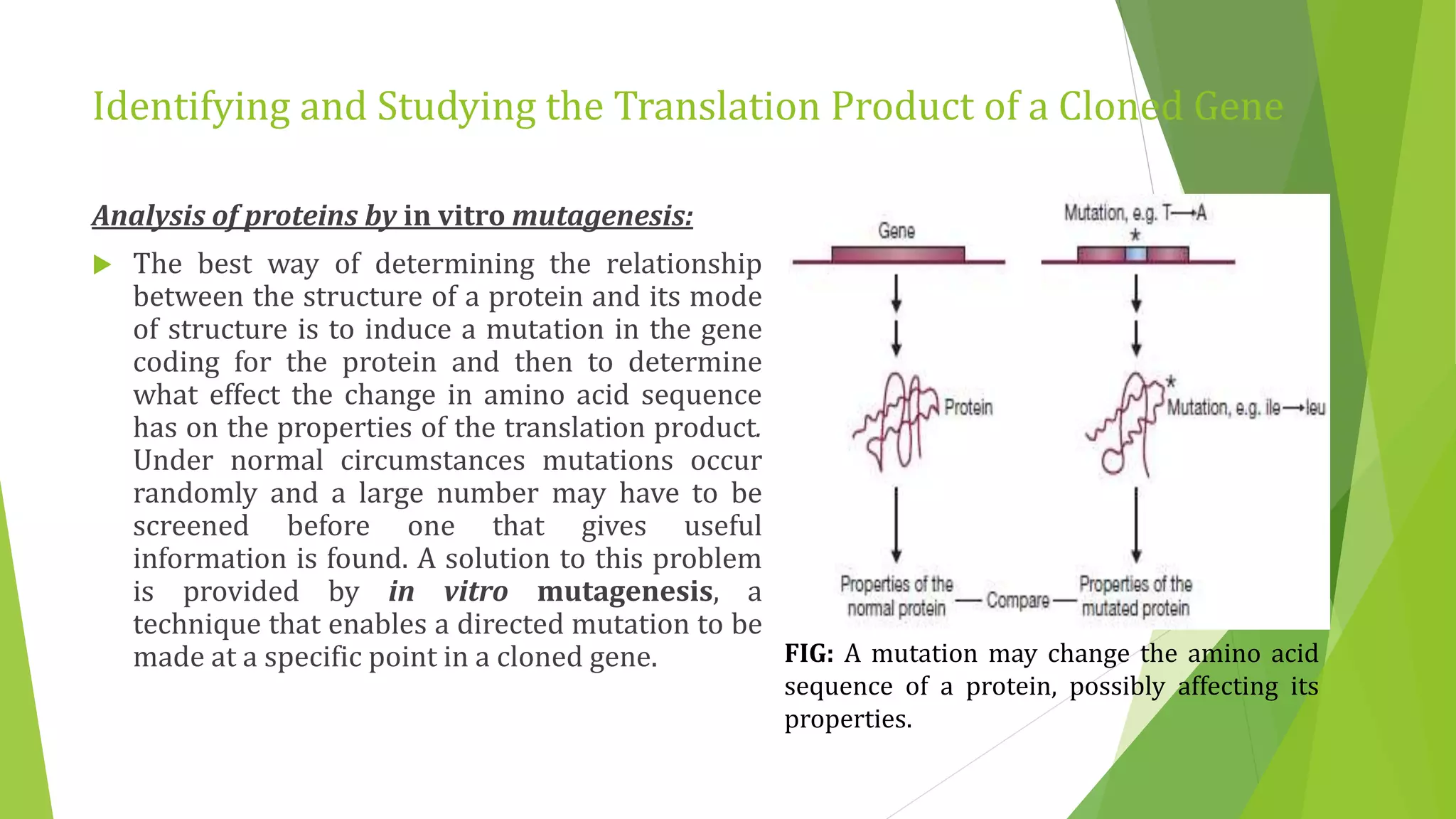
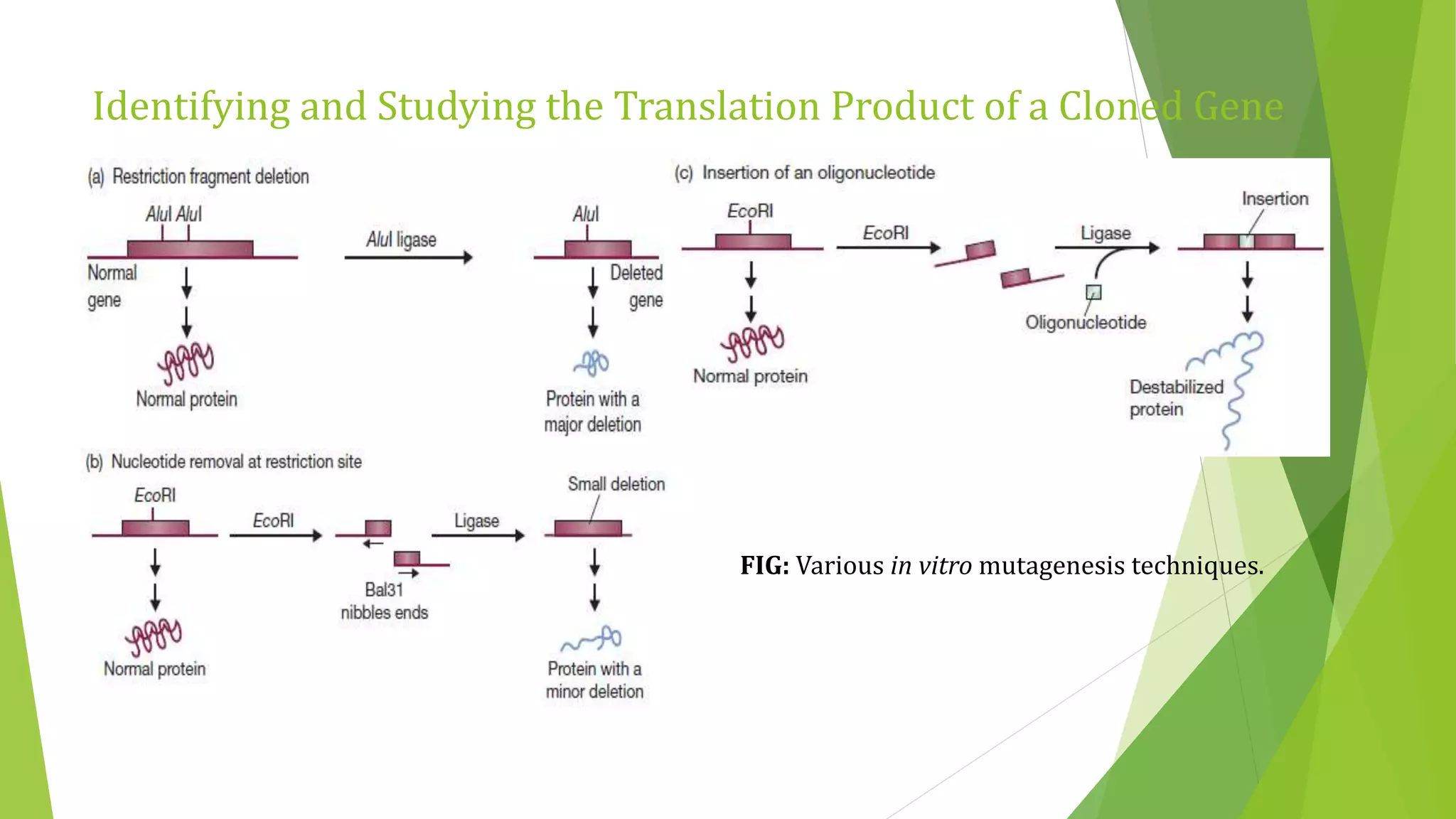
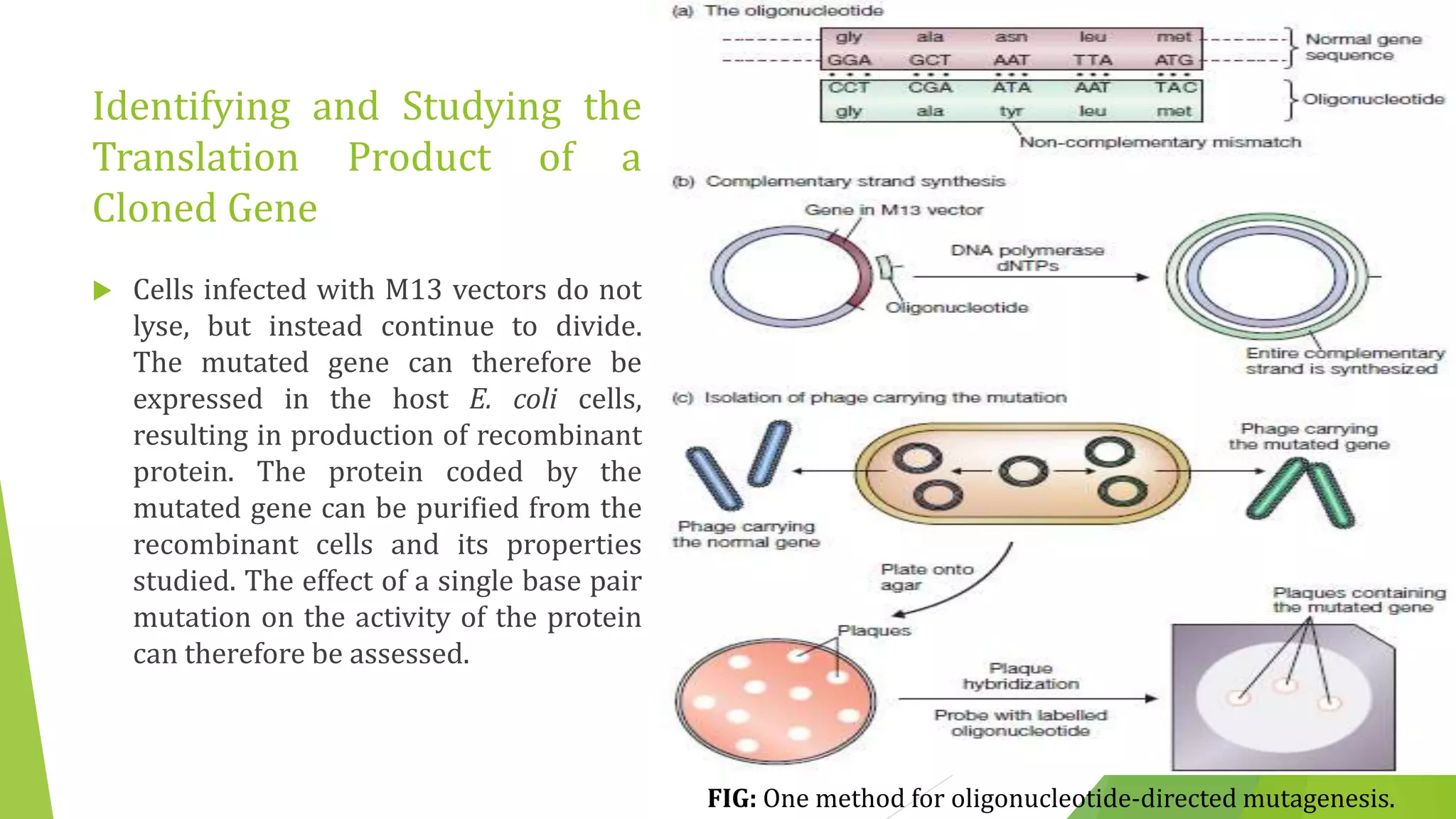
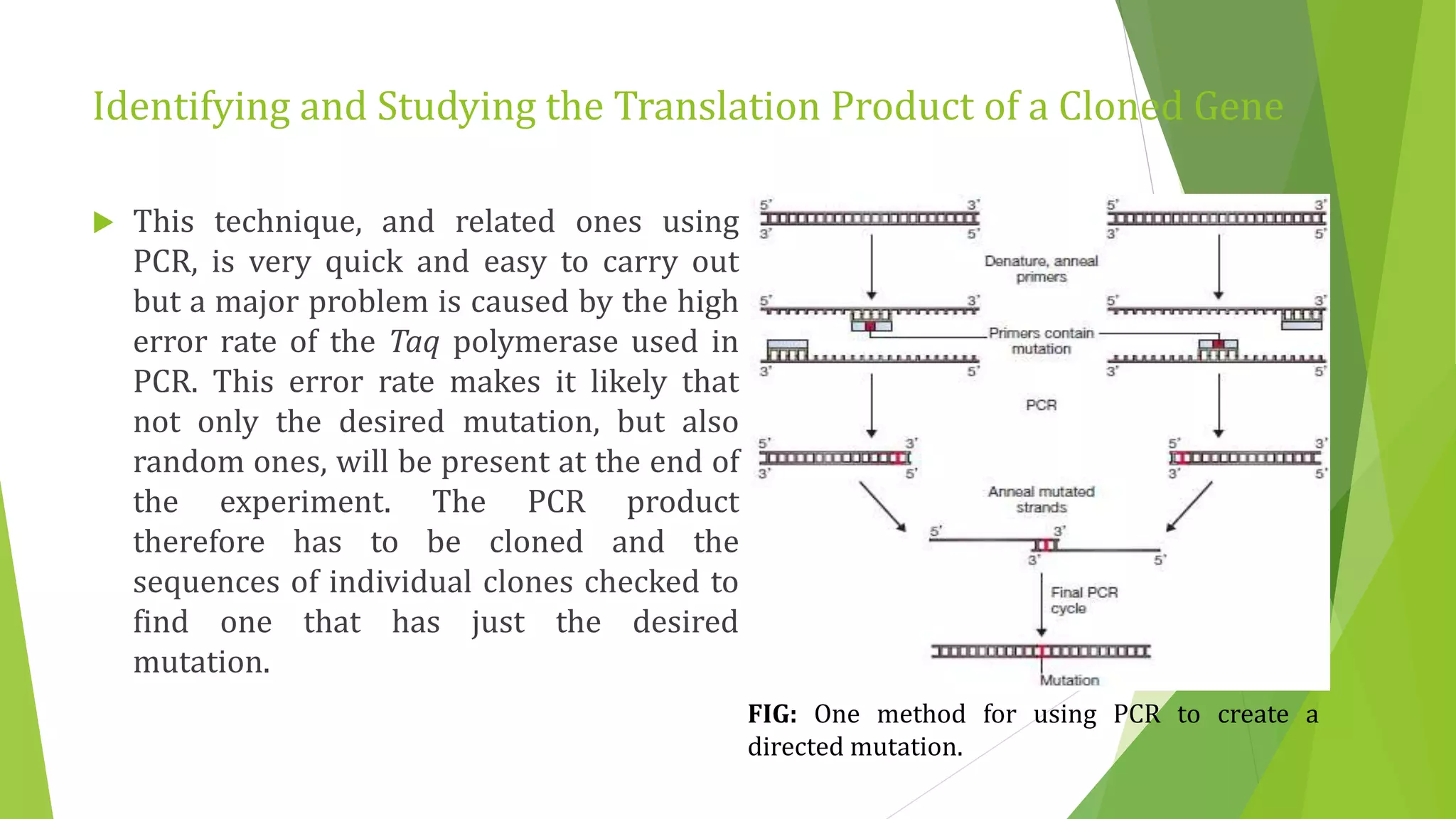
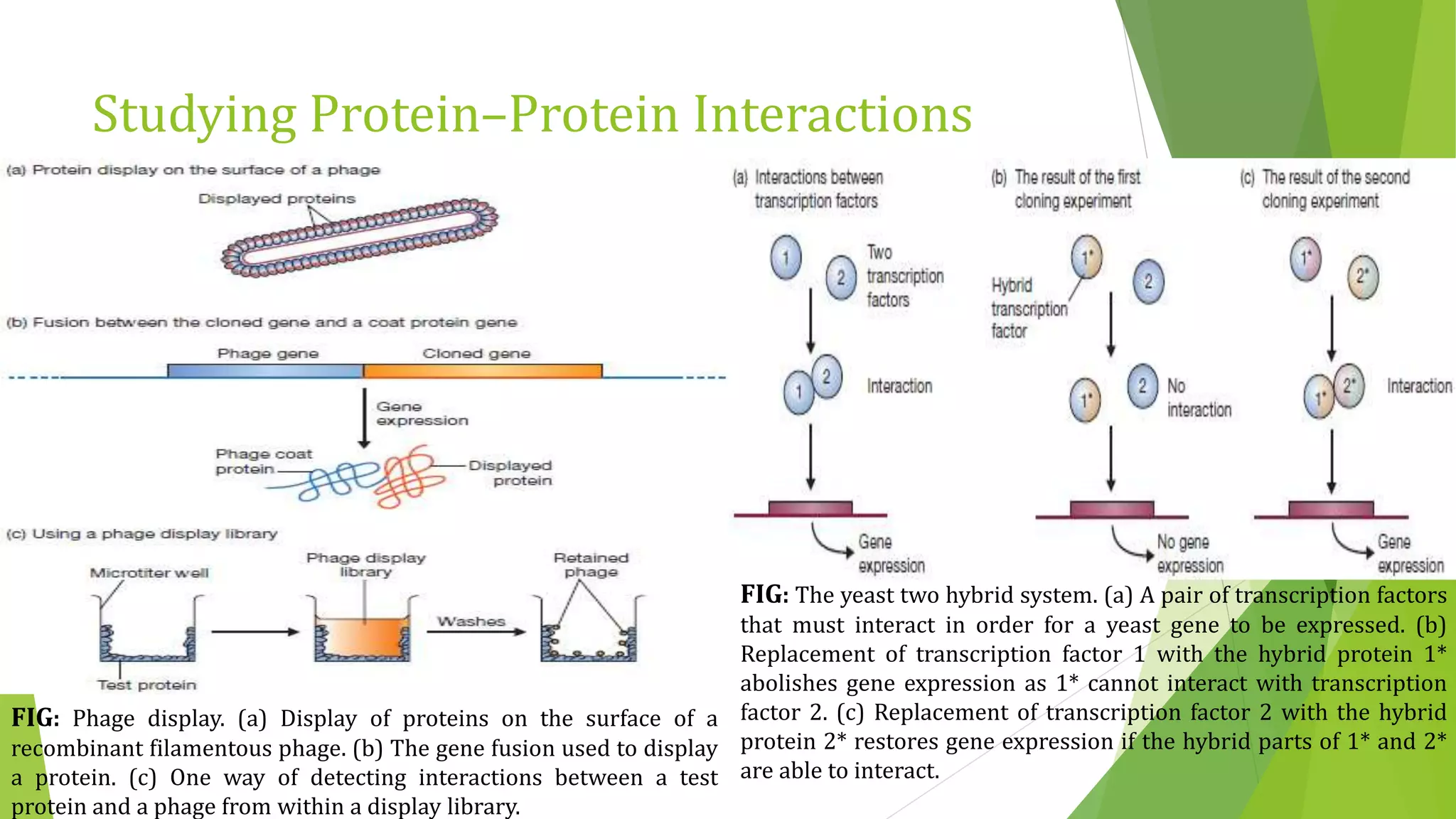
The document discusses various methods for studying gene expression and function, including analyzing RNA transcripts. It describes techniques like northern hybridization, DNA-mRNA hybridization, S1 nuclease mapping, primer extension, and PCR that can be used to study transcripts and locate start/stop points. The document also covers methods for studying gene regulation, such as identifying protein binding sites through gel retardation assays and footprinting, and using deletion analysis to identify control sequences.