Embed presentation
Downloaded 82 times















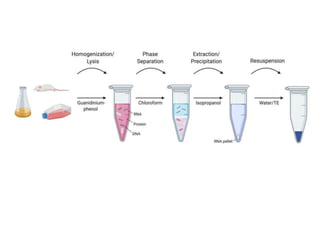


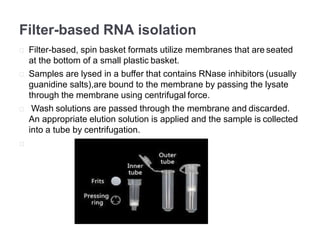
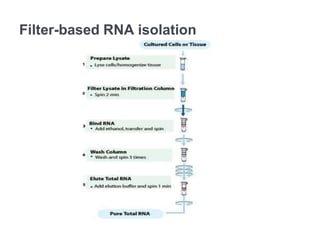


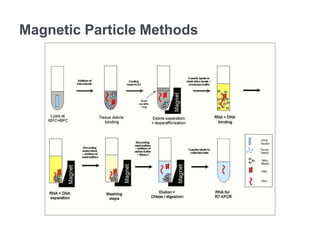
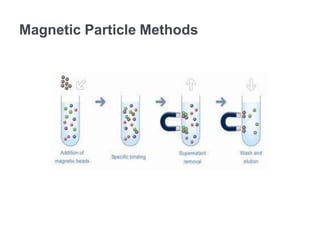

RNA isolation is the purification of RNA from biological samples. It requires strict precautions to avoid degradation by RNases, enzymes that naturally degrade RNA. The TRIzol method uses phenol and chloroform to separate RNA, DNA and proteins during cell lysis and homogenization. Chloroform separates the mixture into aqueous and organic phases, with RNA remaining in the aqueous phase. RNA is then precipitated with isopropanol and washed with ethanol before being re-dissolved in water. RNA concentration and purity is determined spectroscopically. Filter-based and magnetic particle methods provide alternative RNA isolation techniques.


























