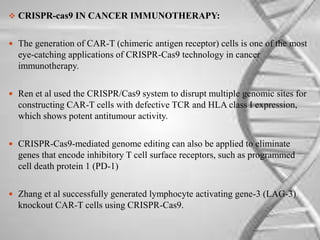
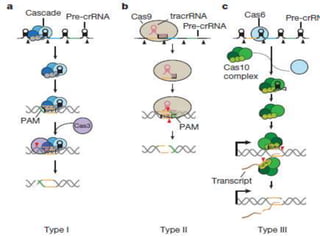
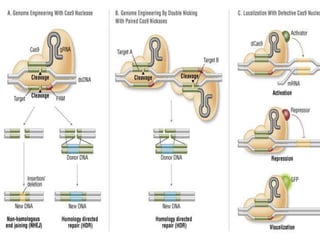
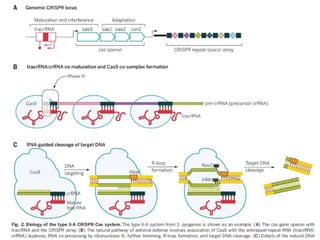
CRISPR-Cas is a natural defense system in bacteria that uses CRISPR sequences and Cas proteins to target and degrade foreign DNA such as from viruses. It has been adapted for genome editing in other organisms using a Cas9 protein guided by a synthetic single guide RNA to introduce targeted double-strand breaks. This system allows for precise genome modifications and has applications in biomedical research, disease treatment, and engineering of plants and other organisms. However, off-target effects and delivery methods require further optimization.









![APPLICATIONS
GENOME EDITING OF IMPORTANT MODEL ORGANISMS AND
CELL TYPES:
Groundbreaking research was begun by Jinek et al. [2012] when they
investigated the mature crRNA that forms a pair with the transactivating
crRNA (tracrRNA)
Reengineered dual-RNA:Cas9 specificity by changing the nucleotides of the
crRNAs to make single and multiple nucleotide changes. They used two
crRNAs to simultaneously create the desired mutations in Streptococcus
pneumoniae.
Hwang et al. [2013] used a CRISPR-Cas9 system in vivo for targeted
genome editing in zebrafish embryos.
Mali et al. [2013] engineered a (sgRNAs) and a human codon-optimized
Cas9 for targeted genome editing.](https://image.slidesharecdn.com/crispr-cas-210414100041/85/Crispr-cas-15-320.jpg)
![ CRISPR-Cas9 AS SMART ANTIMICROBIALAGENTS AGAINST
MDR PATHOGENS:
In order to test the multiplexity of CRISPR-Cas9, they designed two
sgRNAs to target the superantigen enterotoxin sek gene and a region of the
mecA gene.
They observed that target strains had been killed with comparable efficiency
CRISPR-Cas9 FOR THE ERADICATION OF HUMAN
VIRUSES:
The hepatitis viruses include hepatitis A, hepatitis B, and hepatitis C, three
distinct viruses that cause acute and chronic infections in human [Zhang et
al., 2014; Zeng, 2014]. CRISPR-Cas9 has been developed to eliminate HBV
through the targeting of P1 and XCp sites.
Used to eliminate HPV, HSV, HIV](https://image.slidesharecdn.com/crispr-cas-210414100041/85/Crispr-cas-16-320.jpg)
![ CRISPR-cas9 IN THE NEUROSCIENCES:
Recently CRISPR‐Cas9 technology has been used for creating a new in vitro
and in vivo animal model for testing and characterizing the nervous system
and neurological diseases.
CRISPR‐Cas9 interruptions in and around the DISC1 gene - knocked‐down
levels of the protein expression. Further study is required to see whether or
not DISC1 can be corrected in vivo to overcome the schizophrenia.
[Srikanth et al.,2015]
CRISPR‐Cas9 have been applied in iPSC‐based disease models to explore
the mechanism of epilepsy caused by SCN1A loss‐of‐function mutations
[Liu et al., 2016]
Wang et al. [2015] have made mutations in CHD8 gene, replicating autism
spectrum disorders (ASDs).](https://image.slidesharecdn.com/crispr-cas-210414100041/85/Crispr-cas-17-320.jpg)
![ CRISPR-cas9 IN HUMAN DISEASE THERAPY:
CRISPR‐Cas9 has been revolutionized for combating the cardiovascular
diseases. Mutation in the PCSK9 gene results in a reduction in (LDL‐C),
which can improve the heart health and reduce the cardiovascular disease.
Ding et al. [2014] have used CRISPR‐Cas9 for in vivo mutation of
the PCSK9 gene in mouse liver.
Yang et al. [2016] have developed an adeno‐associated virus delivery system
for CRISPR‐Cas9 toward the correction of OTC gene in newborn mice –
hyperammonemia.
CRISPR‐Cas9 has been used to correct the mutated dmd (Duche nne
muscular dystrophy) gene in the mdx mice germ line and found 2–100%
corrections efficiency Long et al., 2014; Nelson et al., 2016].](https://image.slidesharecdn.com/crispr-cas-210414100041/85/Crispr-cas-18-320.jpg)