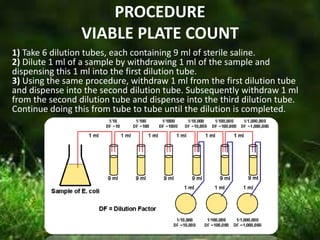
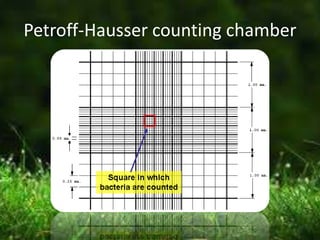
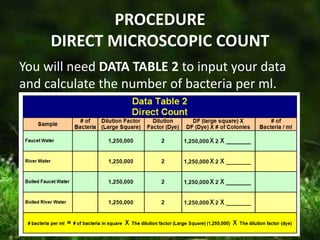
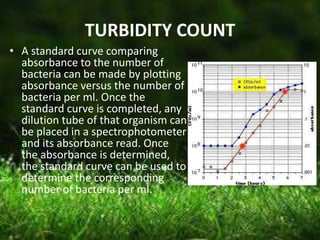
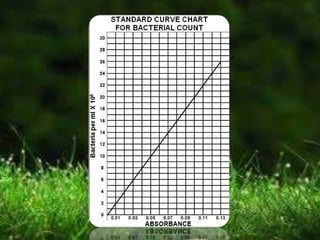
This document describes several methods for enumerating or counting bacteria in a sample, including viable plate count, direct microscopic count, and turbidity count. The viable plate count method involves making serial dilutions of a sample and counting the number of colonies that grow on an agar plate, then multiplying by the dilution factor to determine the concentration in the original sample. The direct microscopic count uses a counting chamber to directly view and count bacteria under a microscope. The turbidity count uses spectrophotometry to measure the turbidity or cloudiness of diluted samples, which correlates to the number of bacteria present based on a generated standard curve. Procedures for each method are provided.