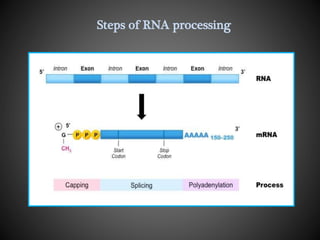
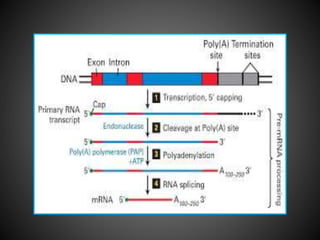
The document discusses the key differences in RNA processing between prokaryotes and eukaryotes, highlighting the necessity of processing in eukaryotes including capping, polyadenylation, and splicing to produce mature mRNA. It describes the roles of various components such as the spliceosome in removing introns and the importance of modifications for mRNA stability and translation. Additionally, it addresses alternative splicing, which enables the generation of different proteins from a single gene by combining various exons.