RNA PROCESSING
- 1. RNA PROCESSING PRESENTEDTO Dr. UDITATIWARI PRESENTED BY MOHD NADEEM M.Sc. BIOCHEMISTRY SEM-II
- 2. RNA Processing Post-transcriptional Processing or RNA processing is a set of biological processes common to mosteukaryotic cells by which anRNA primary transcript is chemically altered following transcription from a gene to produce a mature, functional RNA molecule that can then leave the nucleus and perform any of a variety of different functions in the cell. There are many types of post-transcriptional modifications achieved through a diverse class of molecular mechanisms. Phillip Sharp and RichardRoberts were awarded the 1993 Nobel Prize in Physiology or Medicine for their discovery of introns and the splicing process
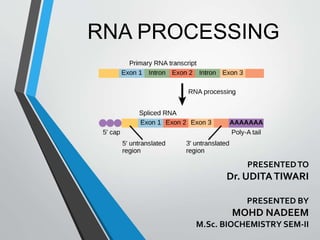
- 4. • In prokaryotes, no RNA processing is necessary: – the nascent RNA is usually the mRNA. • In eukaryotes, the nascent RNA is called primary transcript-RNA – needs to be processed – and transported to the cytoplasm for translation to occur. • The processing steps are: – Addition of a 5’ 7-methyl guanosine cap (capping). – Addition of a poly-A tail at the 3’ end (polyadenylation) – RNA splicing to remove intervening sequences (remove introns
- 7. When the RNA chain is about 30 nucleotides long, the 5’ ends are modified by the addition of a guanine group in the opposite orientation: – involves a 5’-5’ triphosphate linkage
- 8. Guanylate base is methylated at N-7
- 9. 2-Hydroxyl groups of last two riboses may also be methylated
- 10. Eukaryotic RNA Processing:Polyadenylation • nascent RNA is cleaved downstream from the AAUAAA conserved sequence. – By ribonuclease • The enzyme poly(A) polymerase adds adenine ribonucleotides – up to 200 bases long at the 3’ end of the RNA. • The poly(A) tail – enhances the stability of eukaryotic mRNA and – regulates its transport to the cytoplasmic compartment TAILING
- 11. Polyadenylation: The Proteins Proteins required in mammals for cleavage and polyadenylation of a new transcript. Proteins required for efficient cleavage of pre-mRNA: 1. CPSF (cleavage & polyadenylation specificity factor), binds the AAUAAA 2. CstF (cleavage stimulation factor) binds to the G/U rich region cooperatively with CPSF 3. CFI and CFII (cleavage factors I and II), RNA-binding proteins 4. PAP (polyA polymerase) 5. nRNAP II (the CTD of the very large RPB1 subunit) stimulates cleavage
- 12. CPSF specificity factor CFI and CFII PAP II CstF stimulation factor PAB II
- 17. The Spliceosome A spliceosome is a large and complex molecular machine found primarily within the nucleus of eukaryotic cells.The spliceosome is assembled from small nuclear RNAs (snRNA) and approximately 80 proteins. snRNAs (U1, U2, U4, U5 and U6) and associated proteins = snRNPs • U1 binds to the GU sequence at the 5' splice site, along with accessory proteins/enzymes, • U2 binds to the branch site, and ATP is hydrolyzed; • U5/U4/U6 trimer binds, and the U5 binds exons at the 5' site, with U6 binding to U2; • U1 is released, U5 shifts from exon to intron and the U6 binds at the 5' splice site; • U4 is released, U6/U2 catalyzes transesterification, U5 binds exon at 3' splice site, and the 5' site is cleaved, resulting in the formation of the lariat; • U2/U5/U6 remain bound to the lariat, and the 3' site is cleaved and exons are ligated usingATP hydrolysis.The spliced RNA is released and the lariat debranches.
- 18. Three types of short sequences dictate the precise cutting of the intron/exon boundaries - called splice junctions. – Splice donor: 5’ end of intron: exon-G-U – SpliceAcceptor: 3’ end of intron: A-G-exon – Branch site: within the intron, about 30 nucleotides upstream of the splice acceptor, has an AT rich region with at least one A.
- 20. Alternative Splicing → ProteinDiversity Splicing Mutations → Disease