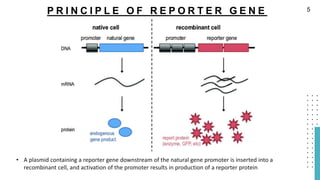
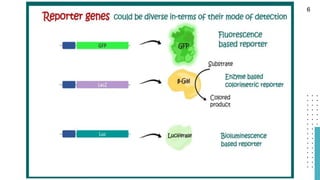
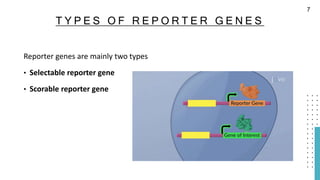
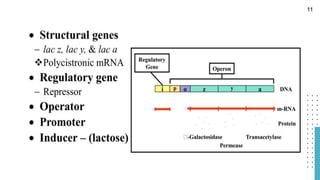
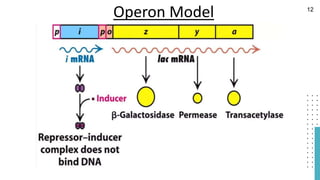
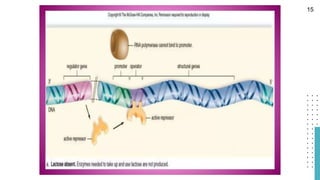
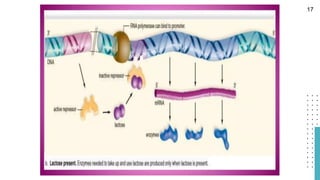
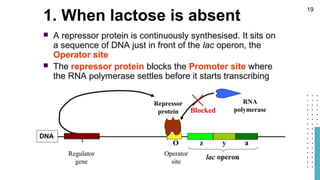
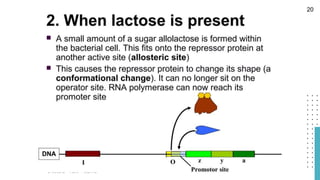
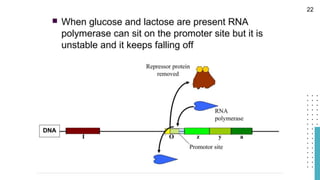
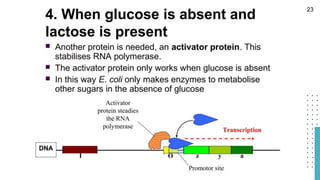
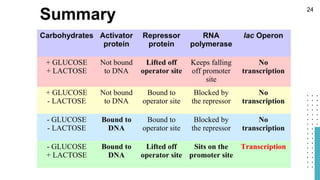
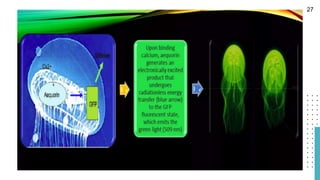
The document discusses reporter genes, which can differentiate transformed cells from non-transformed ones through specific phenotypes. It explains two main types of reporter genes: selectable (e.g., lacZ) that allow cells to survive under selective conditions, and scorable (e.g., GFP) that produce quantifiable phenotypes for studying gene expression. Additionally, the document details the lac operon's functioning and the characteristics needed for an ideal reporter gene.