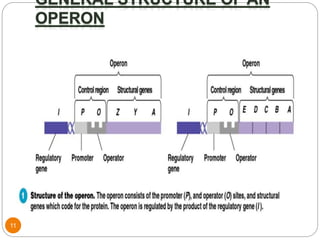
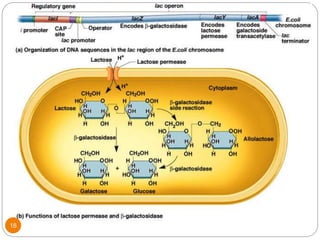
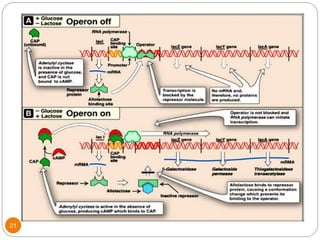
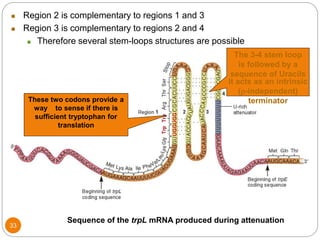
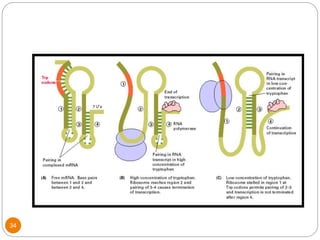
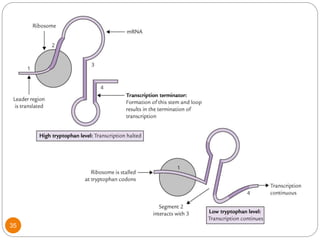
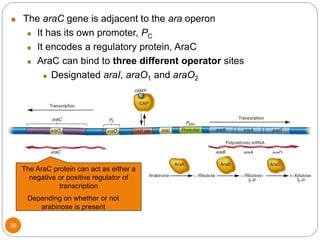
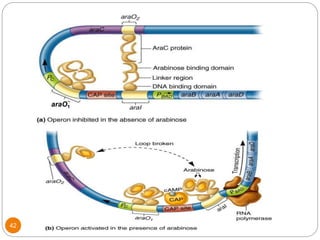
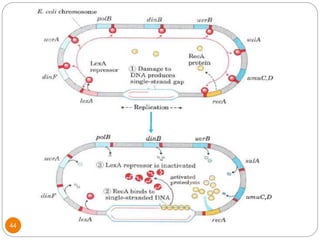
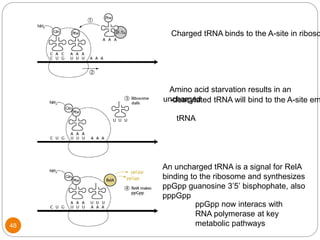
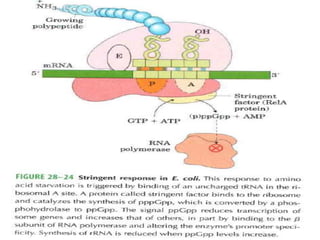
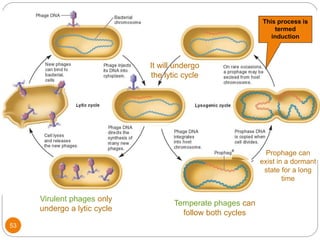
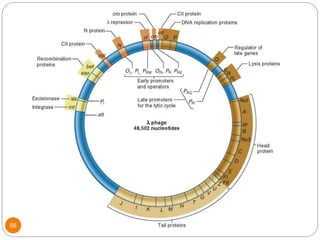
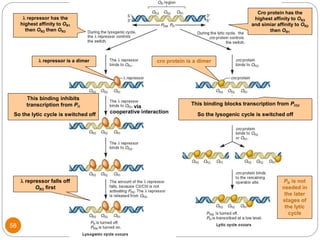
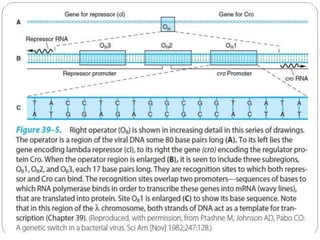
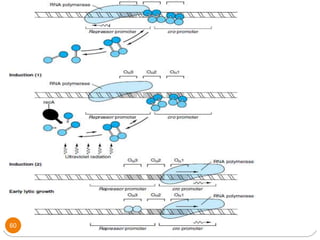
This document discusses gene regulation in prokaryotes. It begins by introducing the concept of gene regulation and how only a fraction of genes are expressed at any given time. It then discusses several key examples of gene regulation in prokaryotes, including the lac, trp, and arabinose operons. The lac operon regulates lactose metabolism in E. coli in response to glucose and lactose availability. The trp operon regulates tryptophan synthesis using both repression and attenuation. The arabinose operon regulates arabinose metabolism using both positive and negative regulation by the AraC protein. The document also briefly discusses the SOS response, stringent response, and gene regulation in bacteriophage lambda.