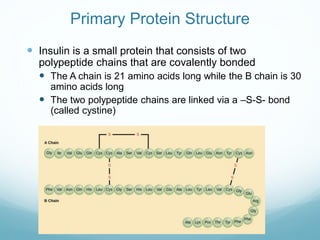
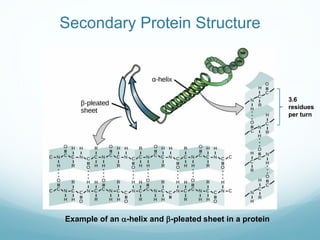
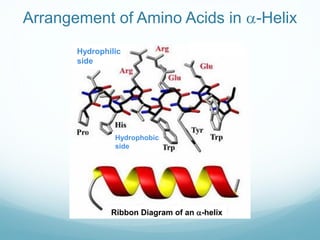
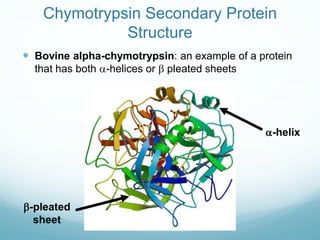
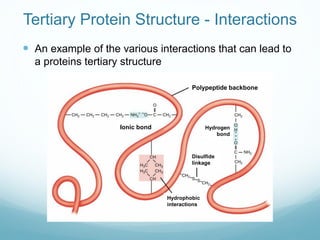
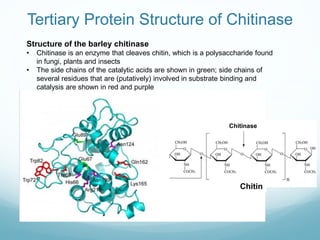
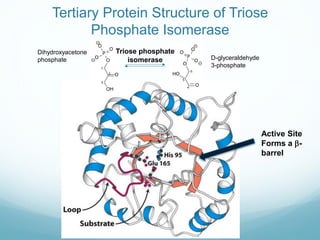
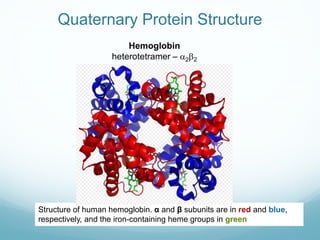
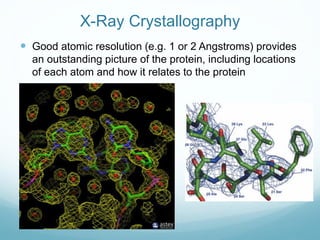
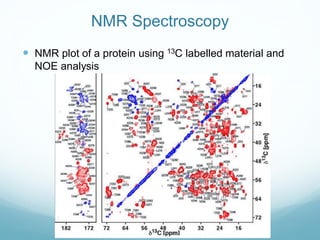
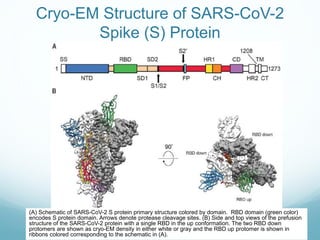
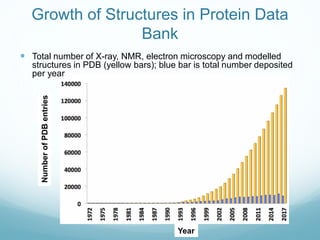
Protein structure is hierarchical, proceeding from primary to quaternary structure. Primary structure refers to the linear sequence of amino acids. Secondary structure involves folding into alpha helices and beta sheets. Tertiary structure describes the overall three-dimensional shape of a polypeptide. Quaternary structure refers to the arrangement of multiple protein subunits. Several methods can determine protein structure at high resolution, including X-ray crystallography, NMR spectroscopy, cryo-electron microscopy, and X-ray free electron lasers.