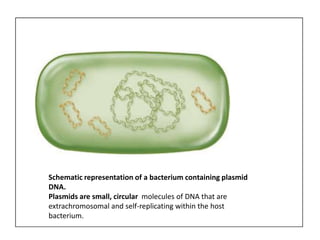
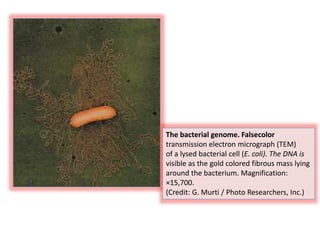
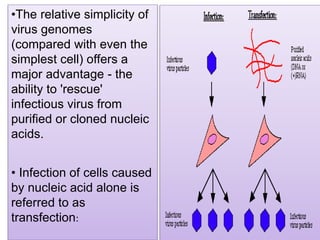
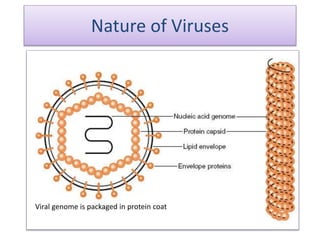
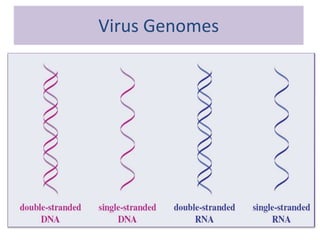
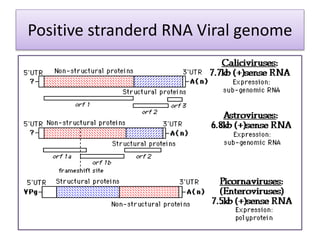
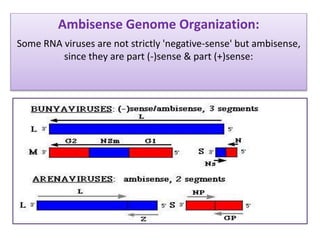
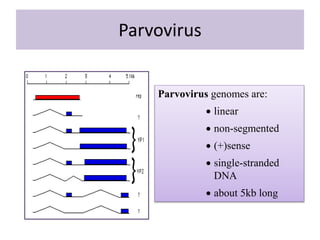
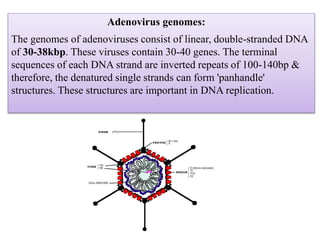
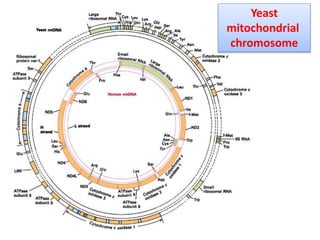
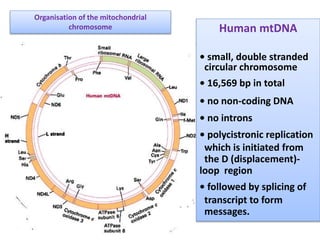
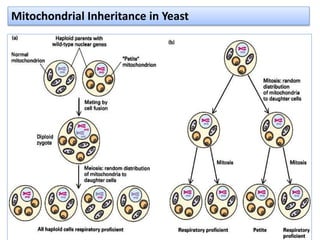
Plasmids are small, circular DNA molecules that are self-replicating and carried by bacteria. They range in size from 2-100kb and can contain genes for antibiotic resistance. Bacterial genomes exist as a single circular chromosome that is highly condensed and packaged. Viruses have RNA or DNA genomes that are either single or double-stranded. Their genomes must be able to be recognized and expressed by their host cell. Mitochondria and chloroplasts originated from endosymbiotic bacteria and contain their own genomes that are maternally inherited and range in size and structure between species. Plant mitochondrial DNA can be much larger than animals.