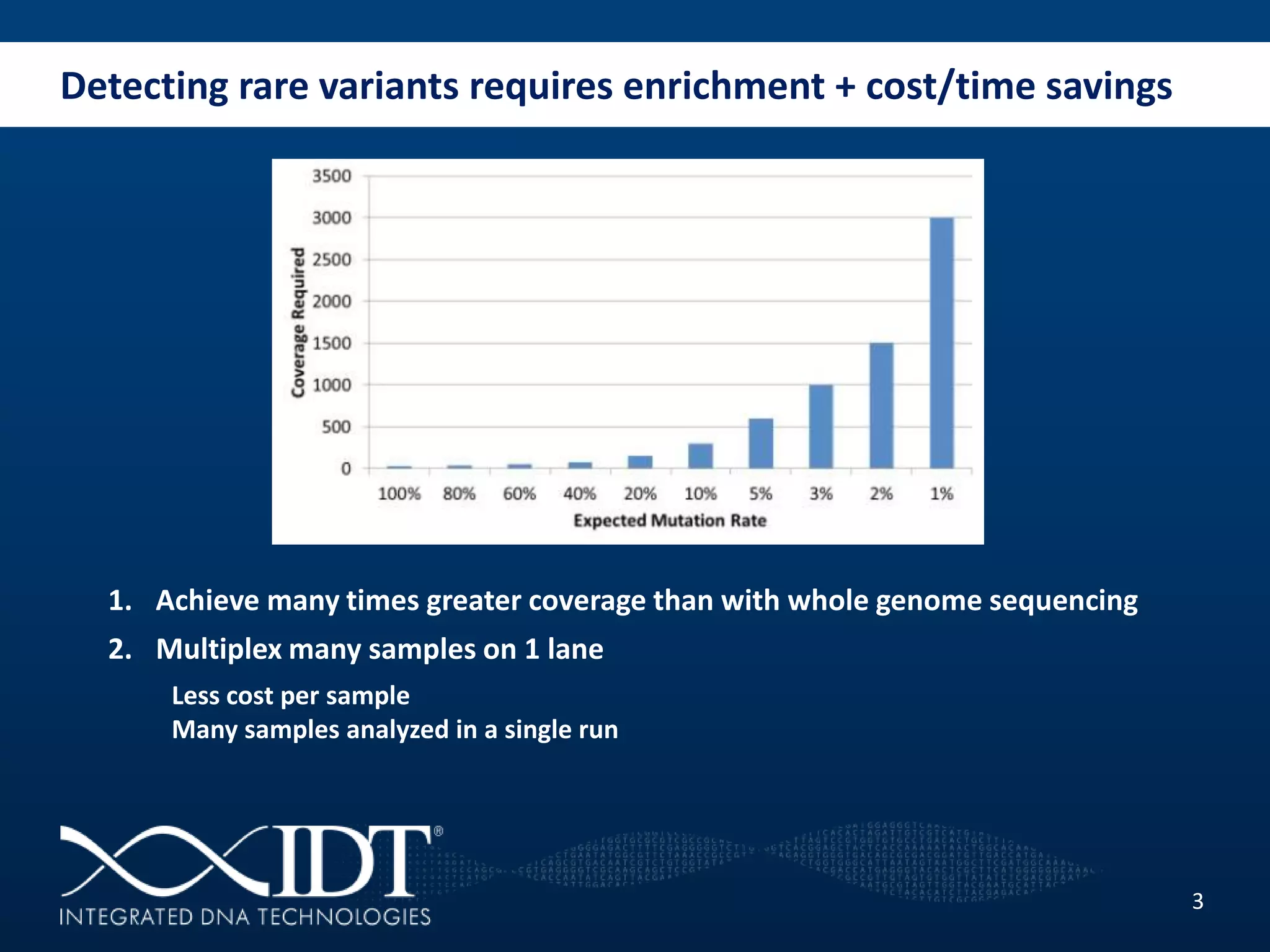
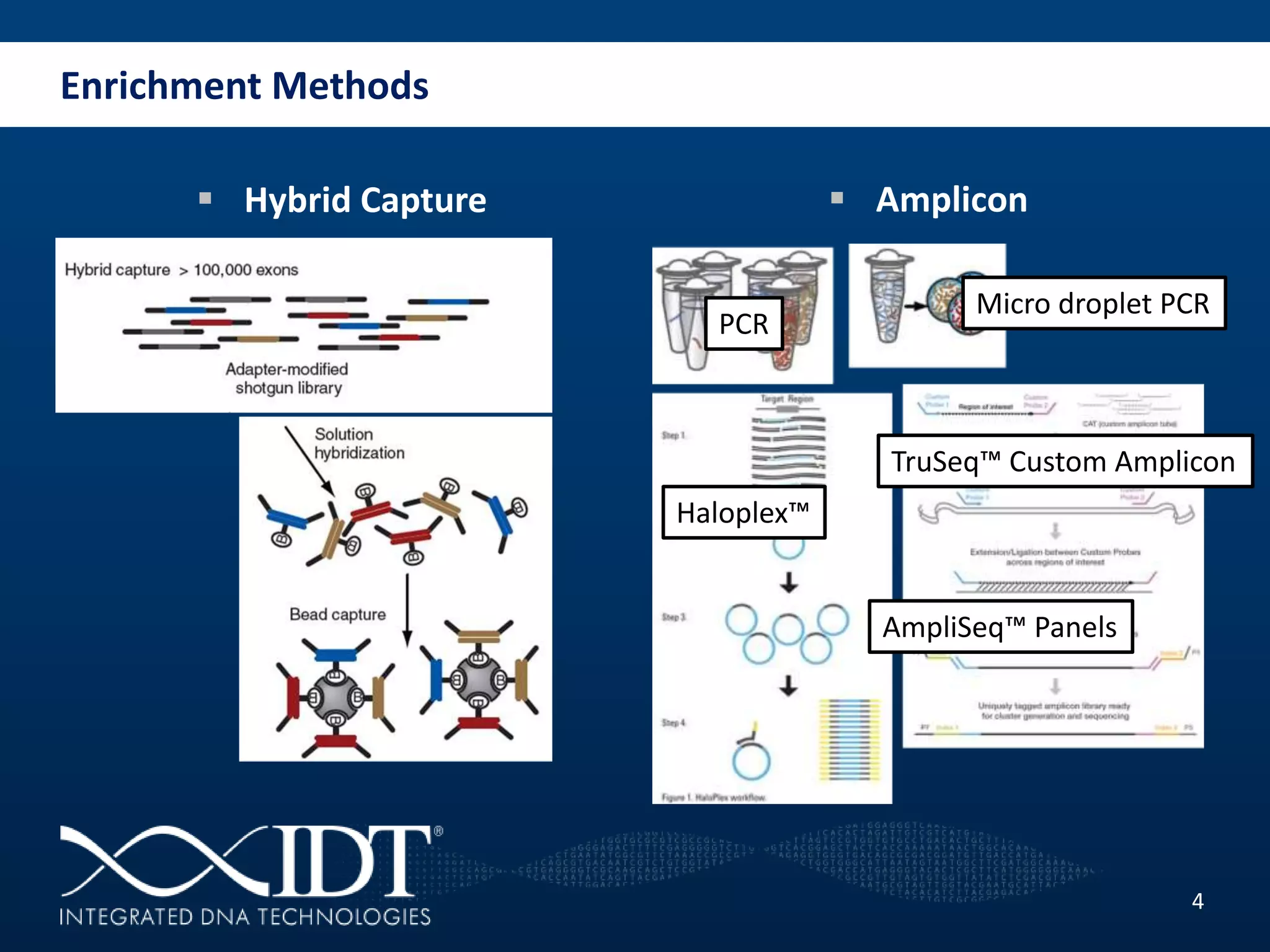
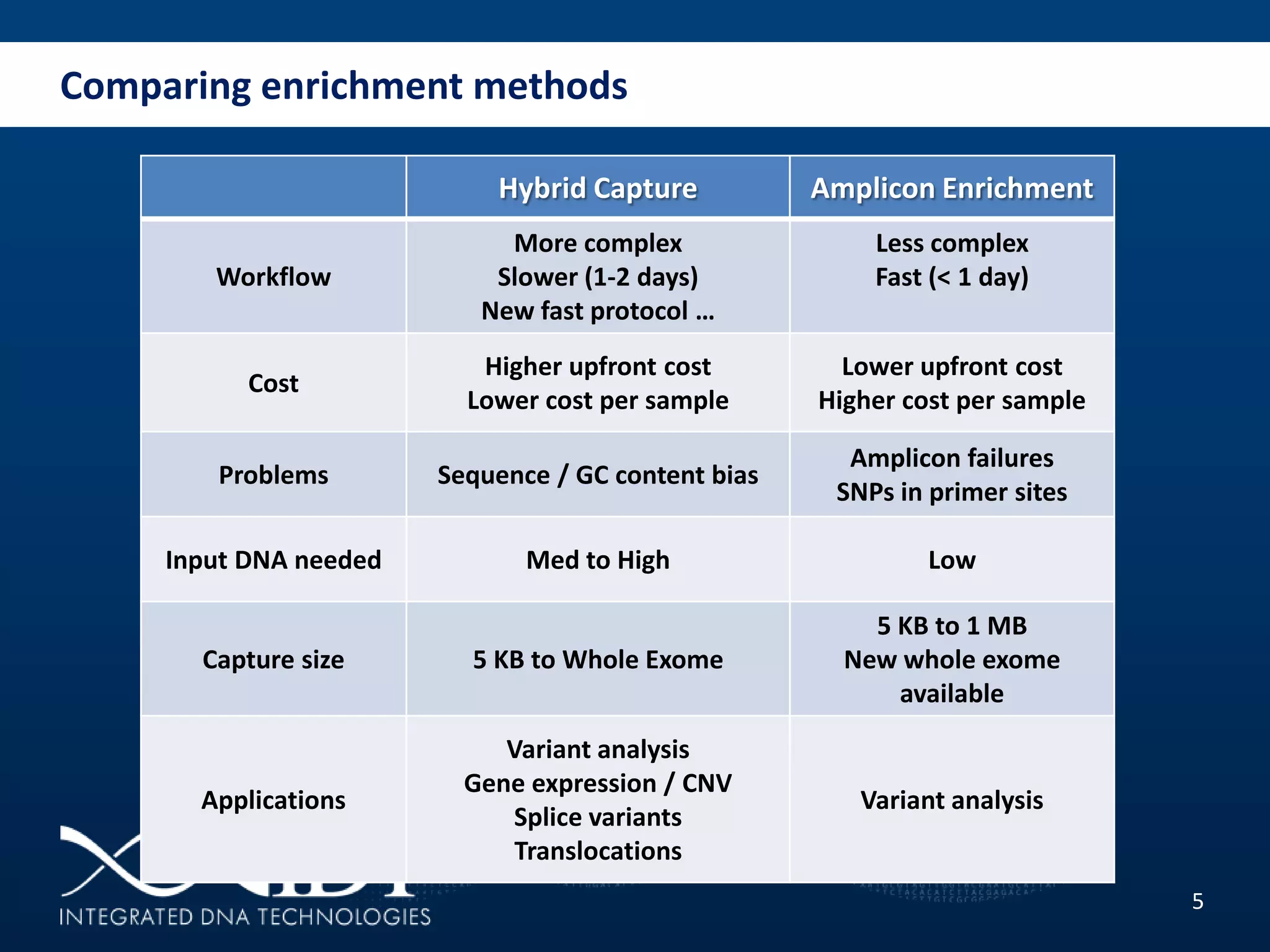
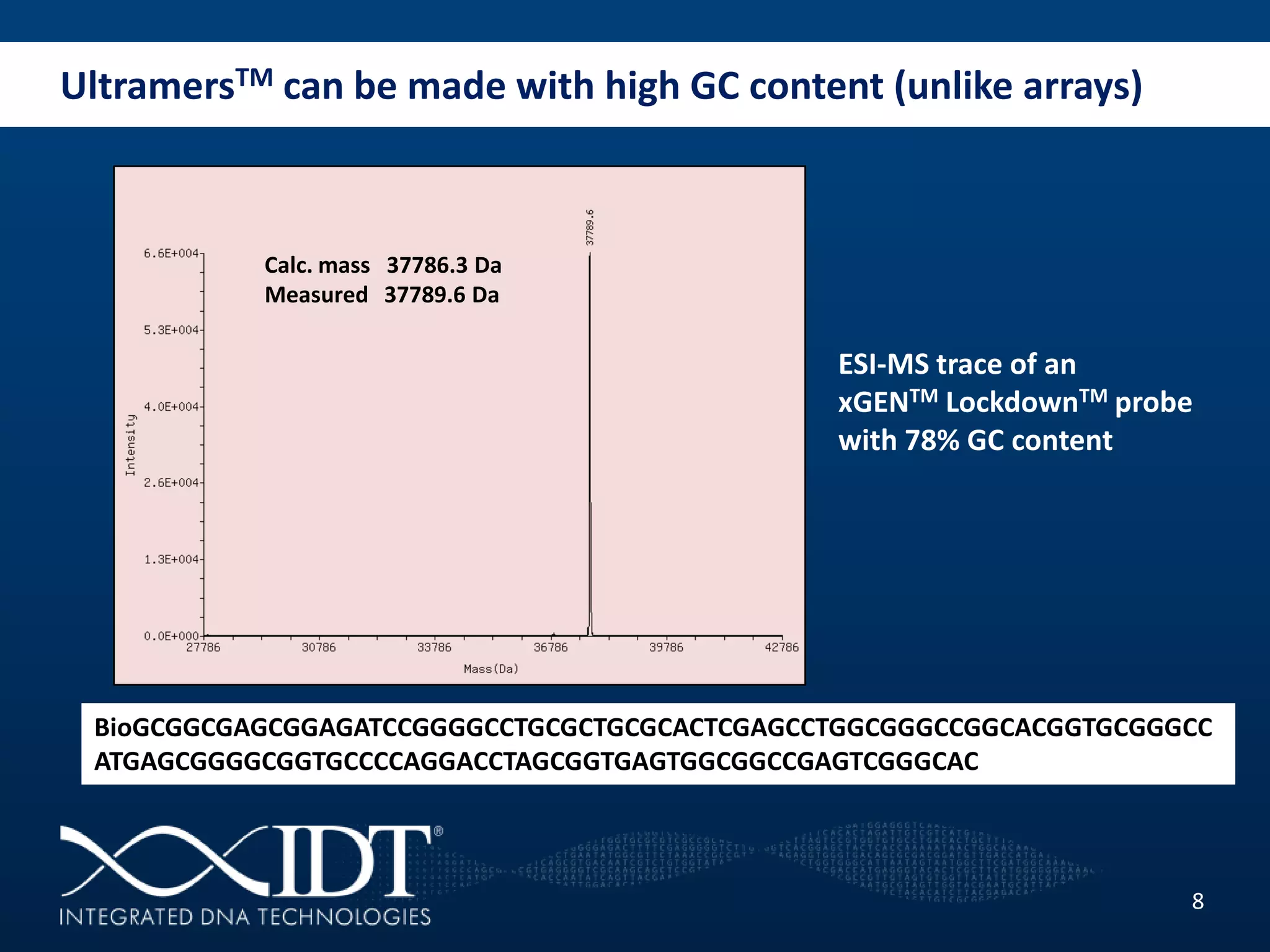
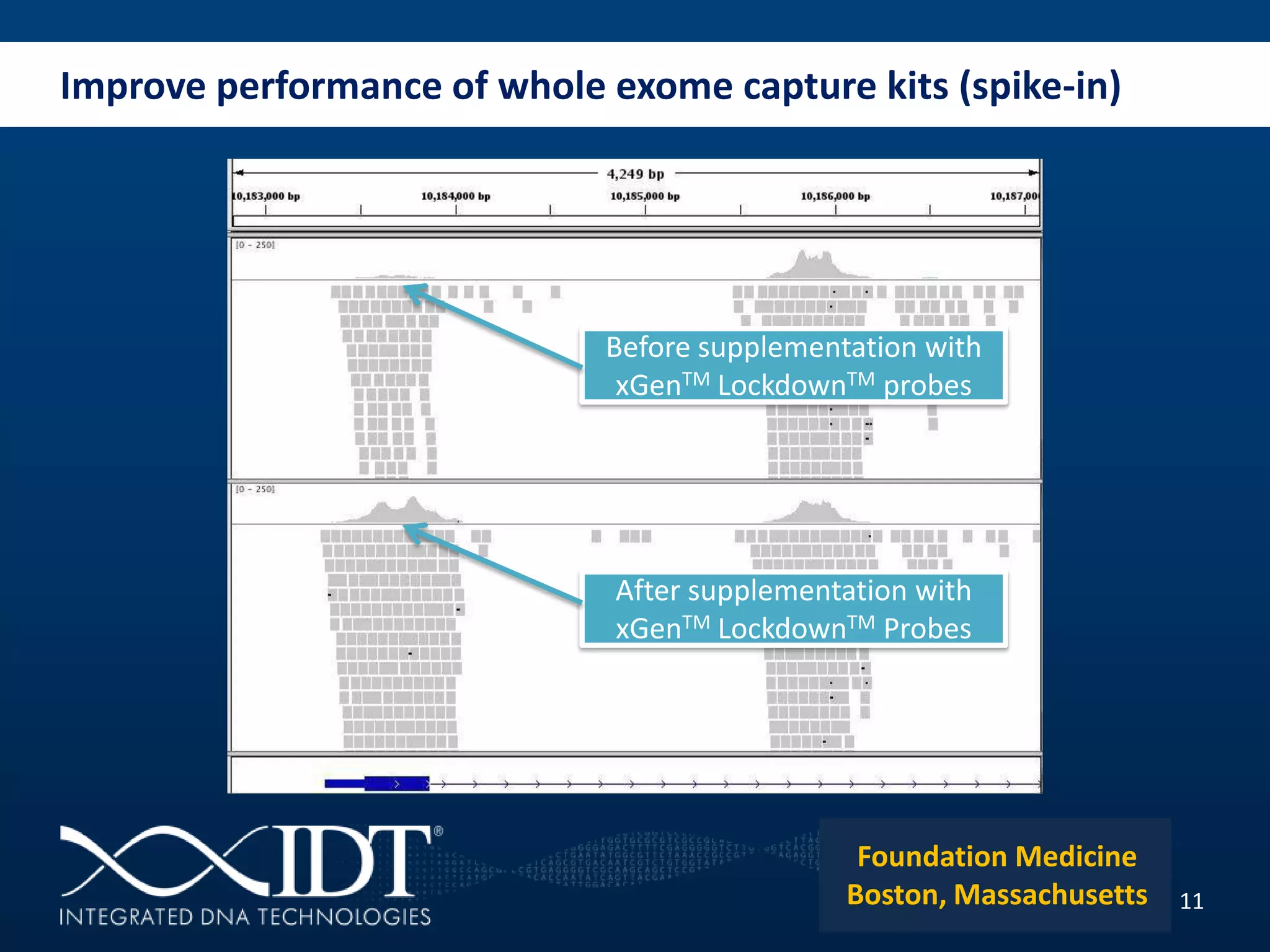
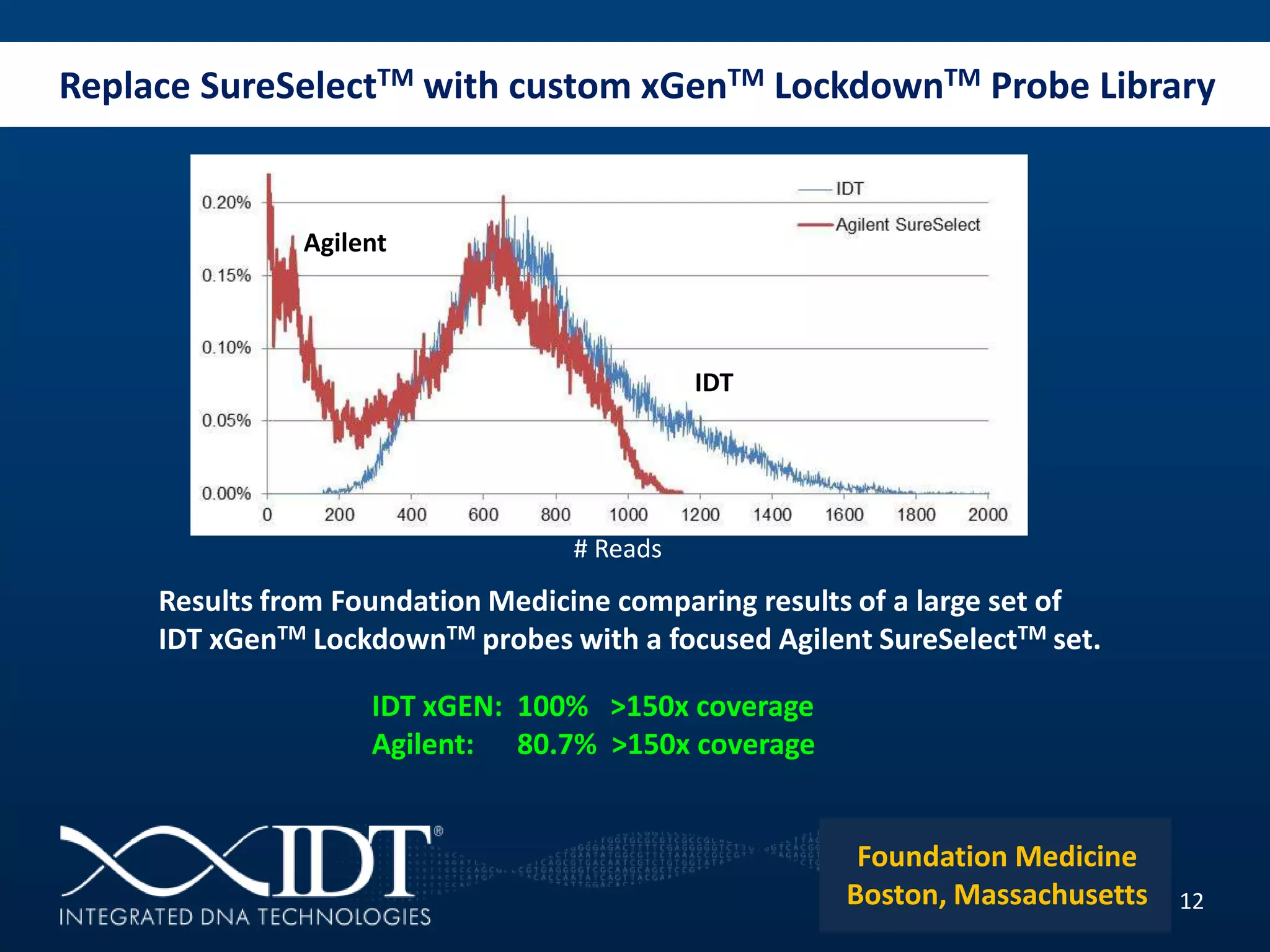
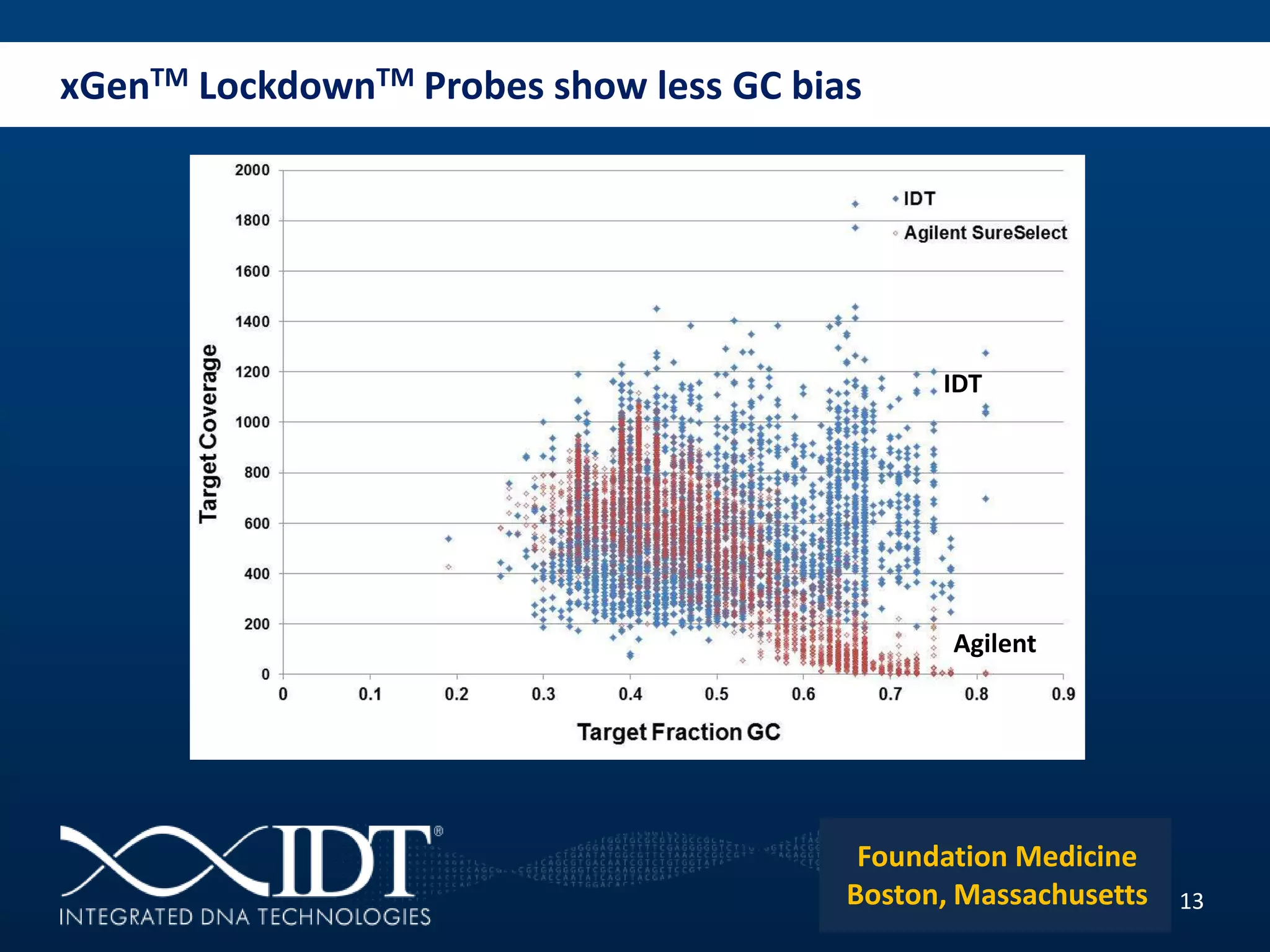
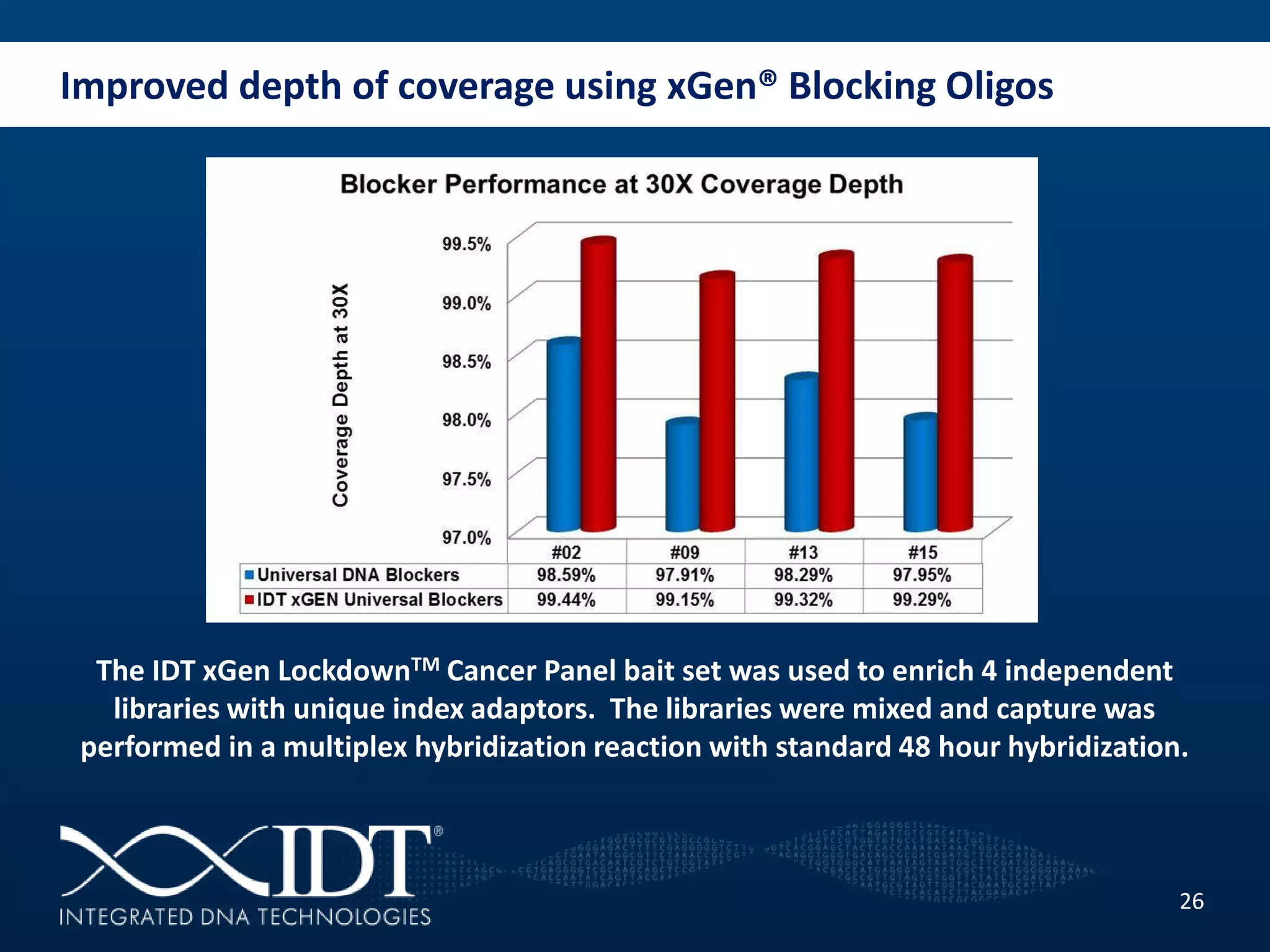
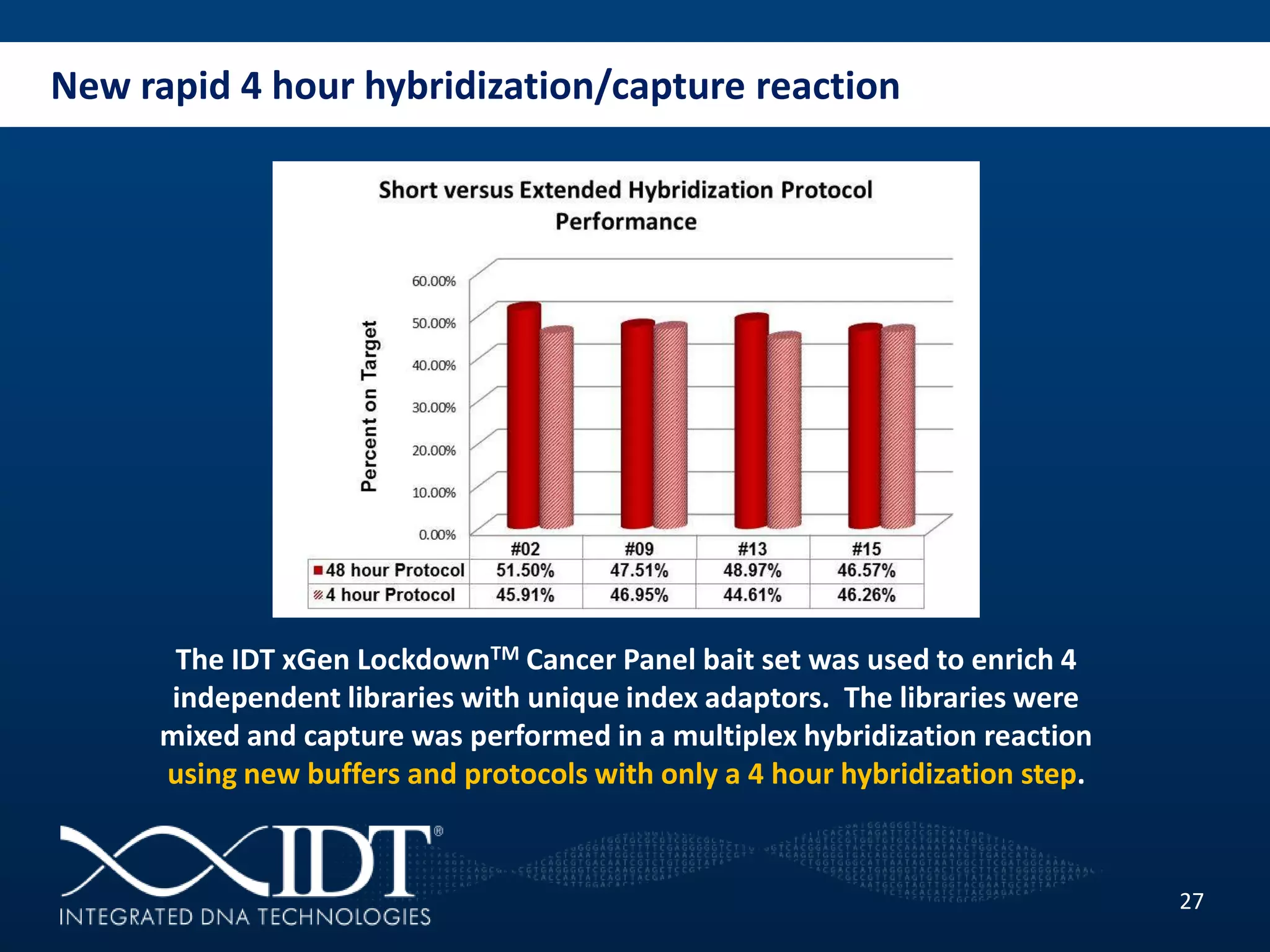
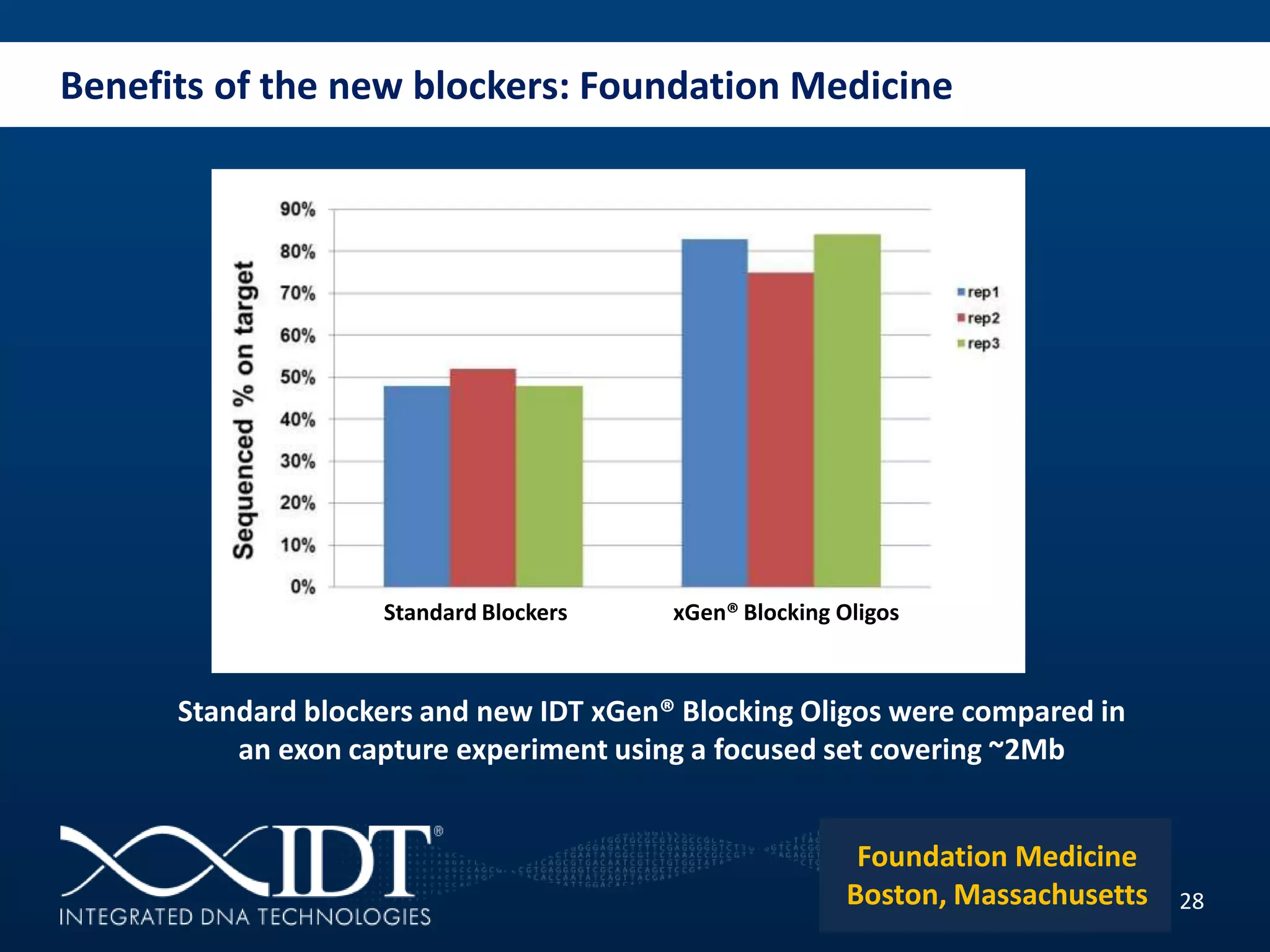
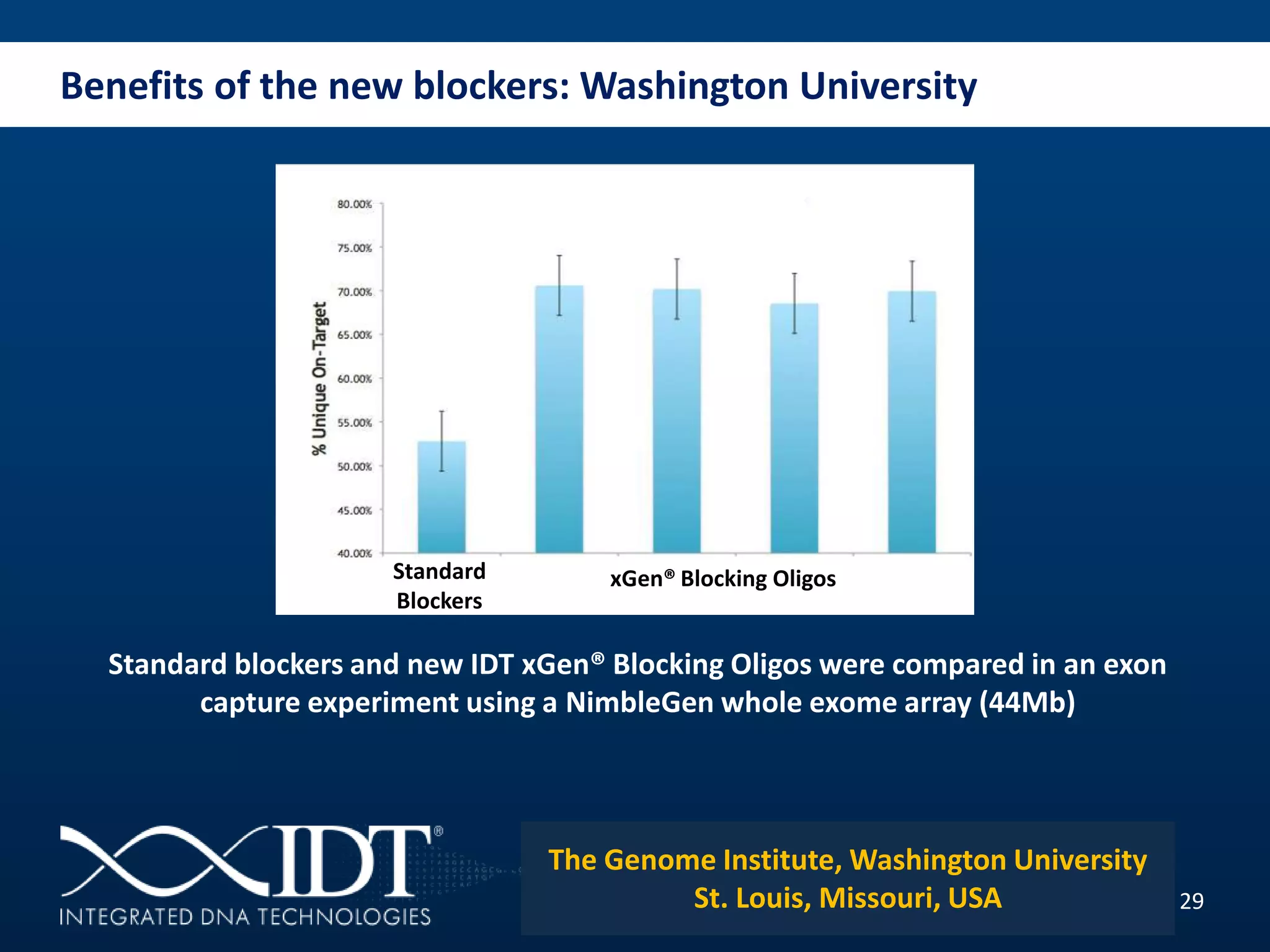
The presentation discussed advancements in next-generation sequencing, specifically focusing on improved reagents and methods for target enrichment. It compared various enrichment methods, highlighting the benefits of IDT's xgen lockdowntm probes over traditional approaches like Agilent SureSelect. The new xgen blocking oligos further enhance capture efficiency by addressing issues with index mismatches, improving on-target rates, and allowing for rapid hybridization protocols.

![General NGS Workflow
2
DNA DNA Shearing
Adaptor and
Barcode [opt]
Attachment
Enrichment [optional]
Template/Library Preparation
Sequencing Analysis
Why enrich?](https://image.slidesharecdn.com/2013-07-30behlkengstargetenrichmentproductstokyo-130808100449-phpapp02/75/Improved-Reagents-Methods-for-Target-Enrichment-in-Next-Generation-Sequencing-2-2048.jpg)