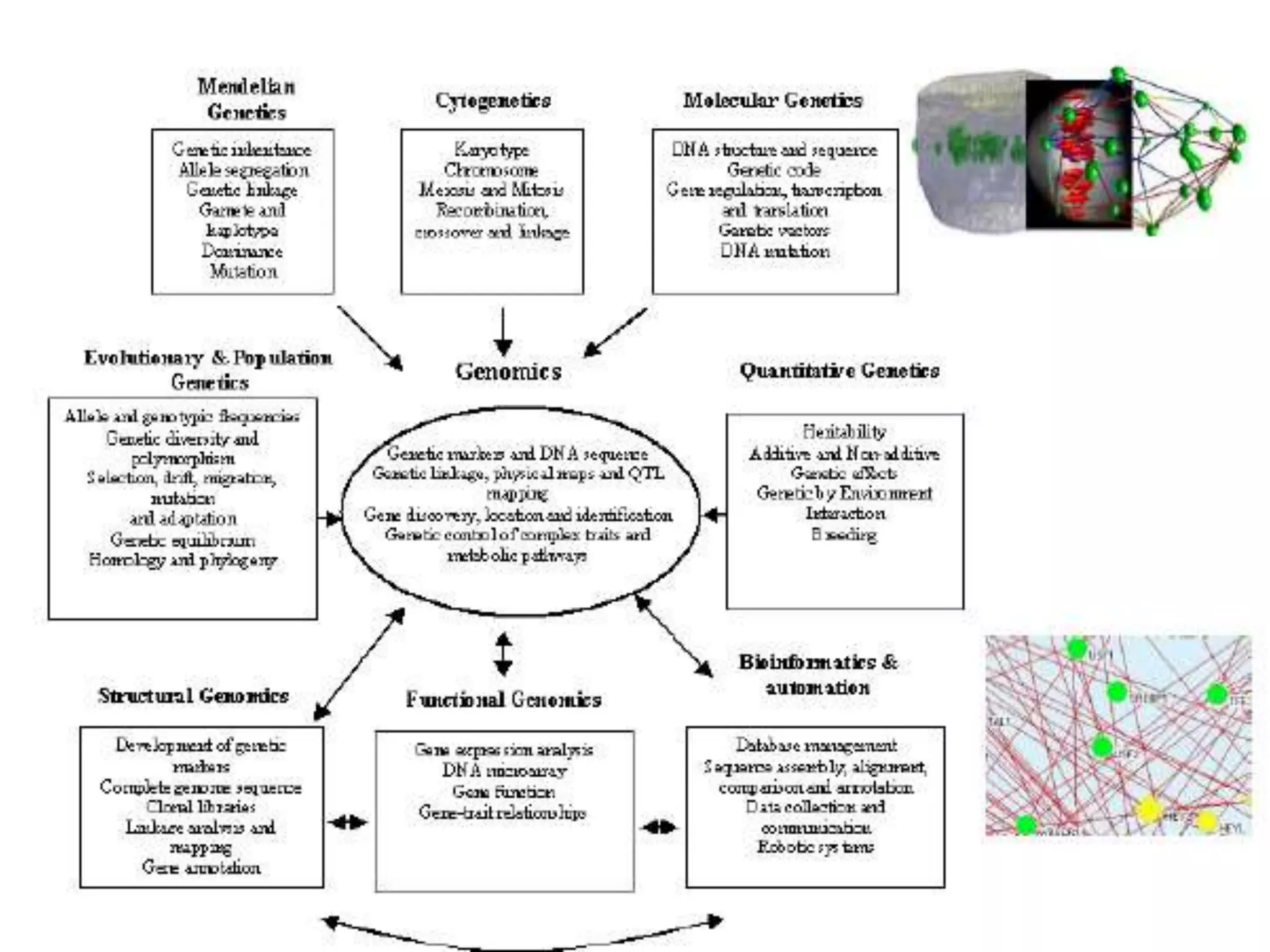
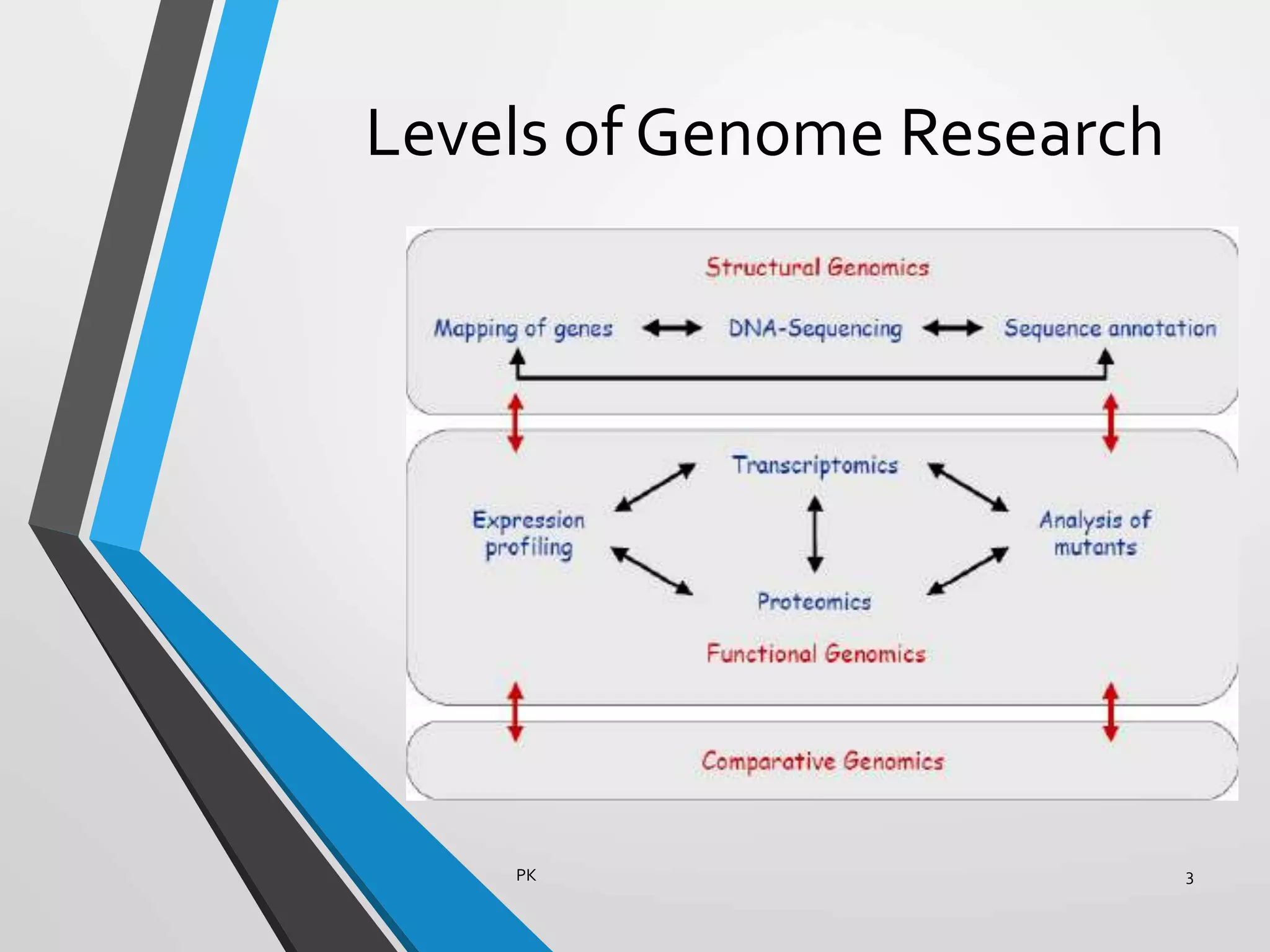
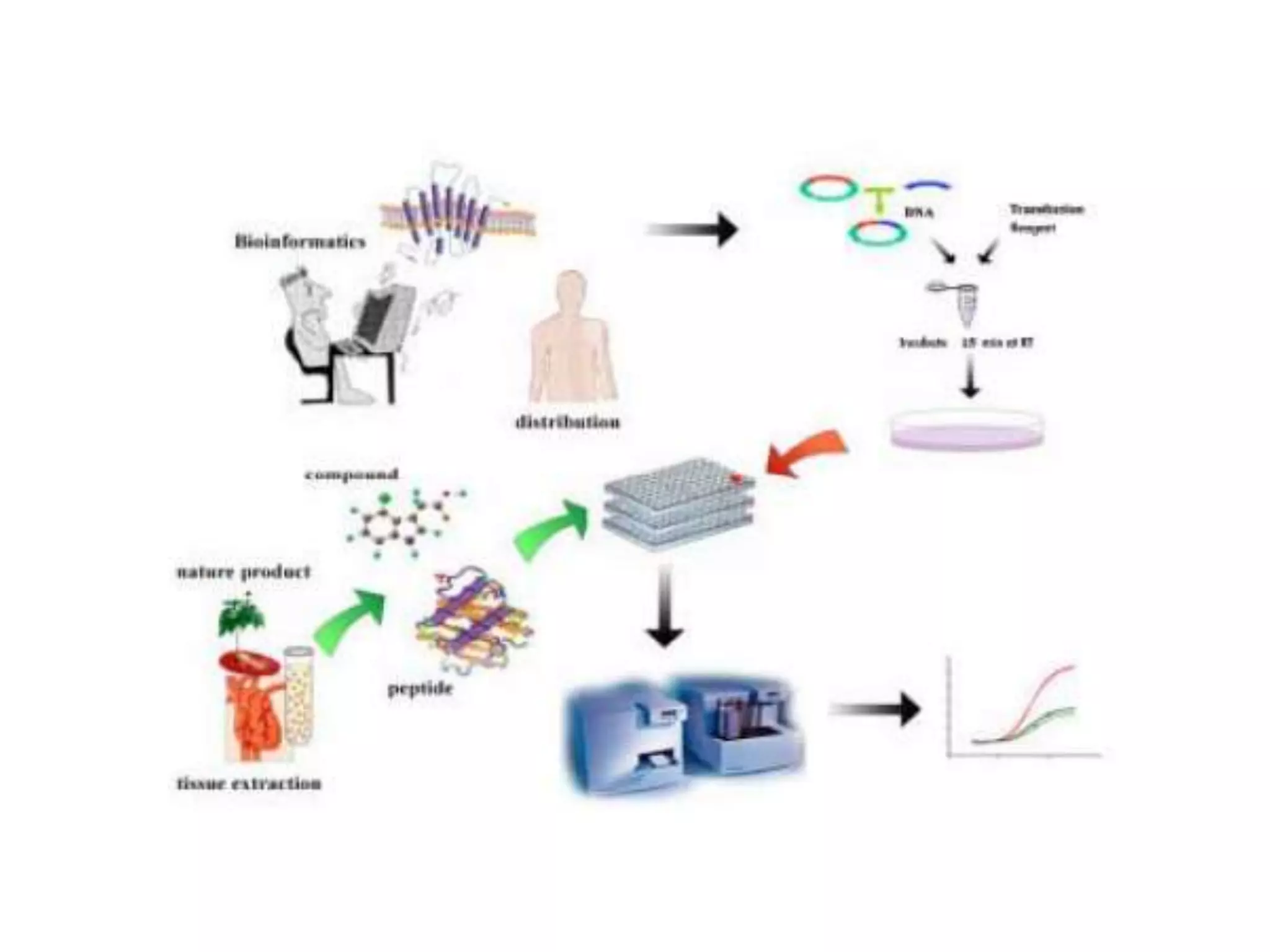
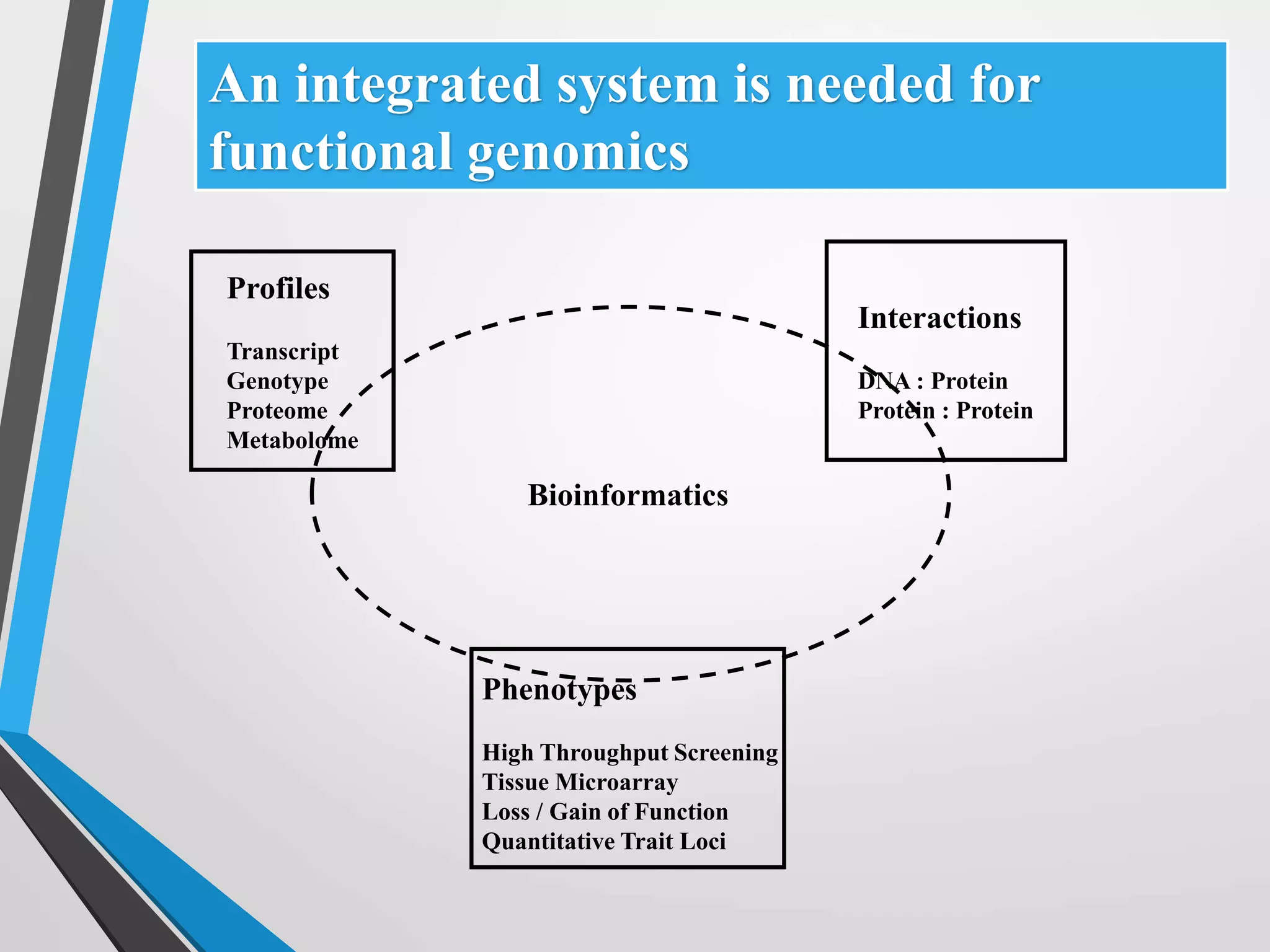
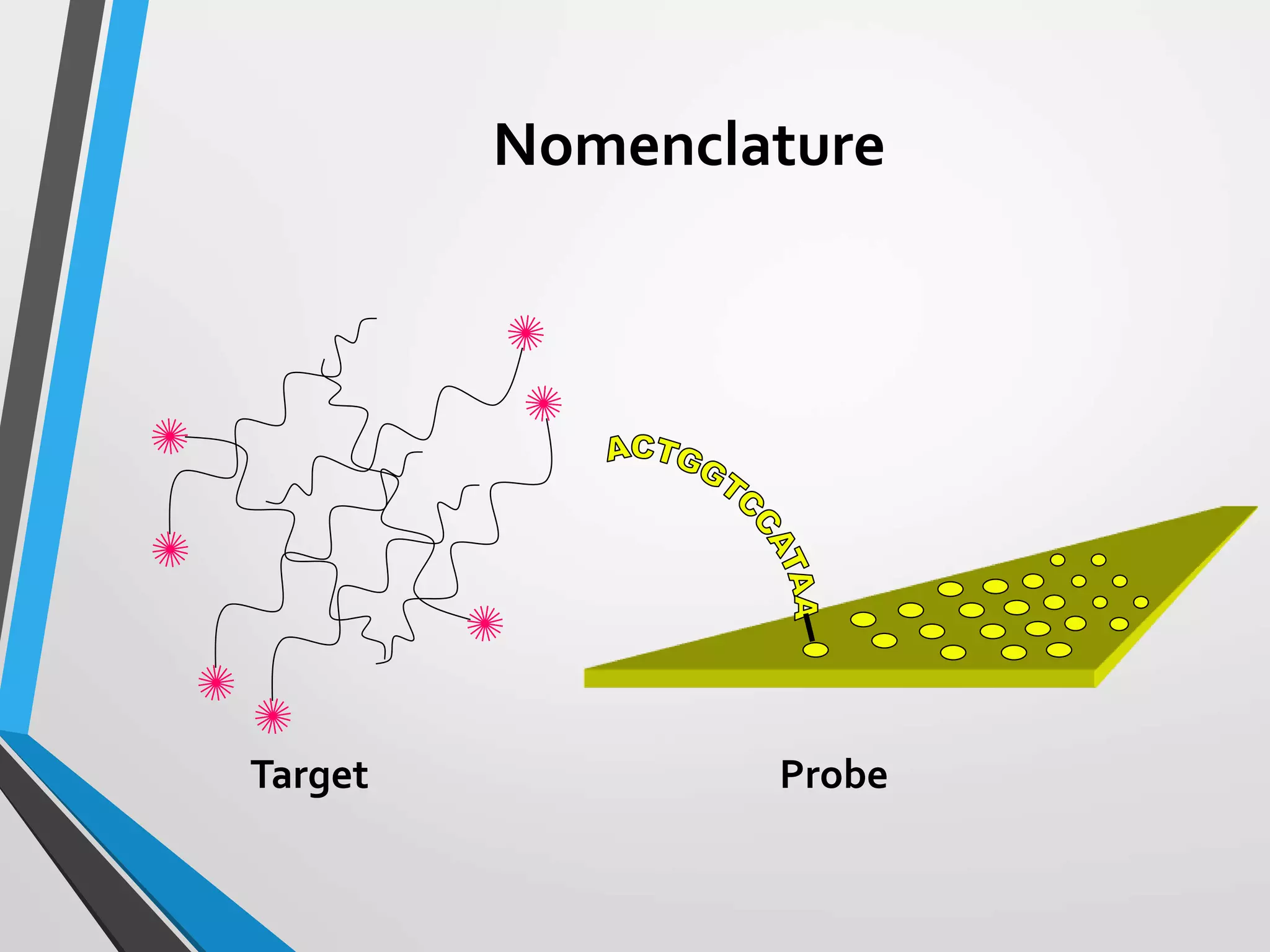
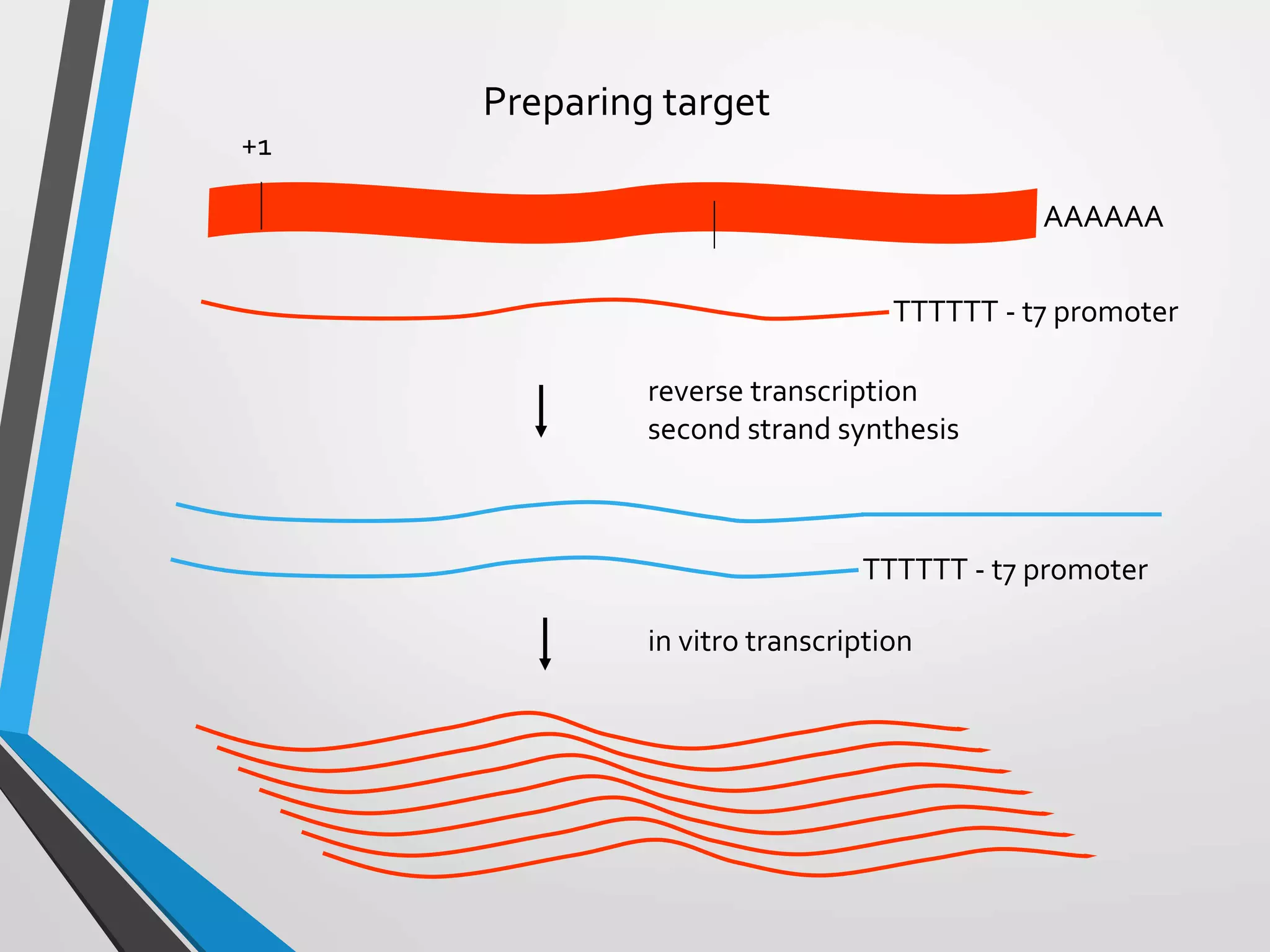
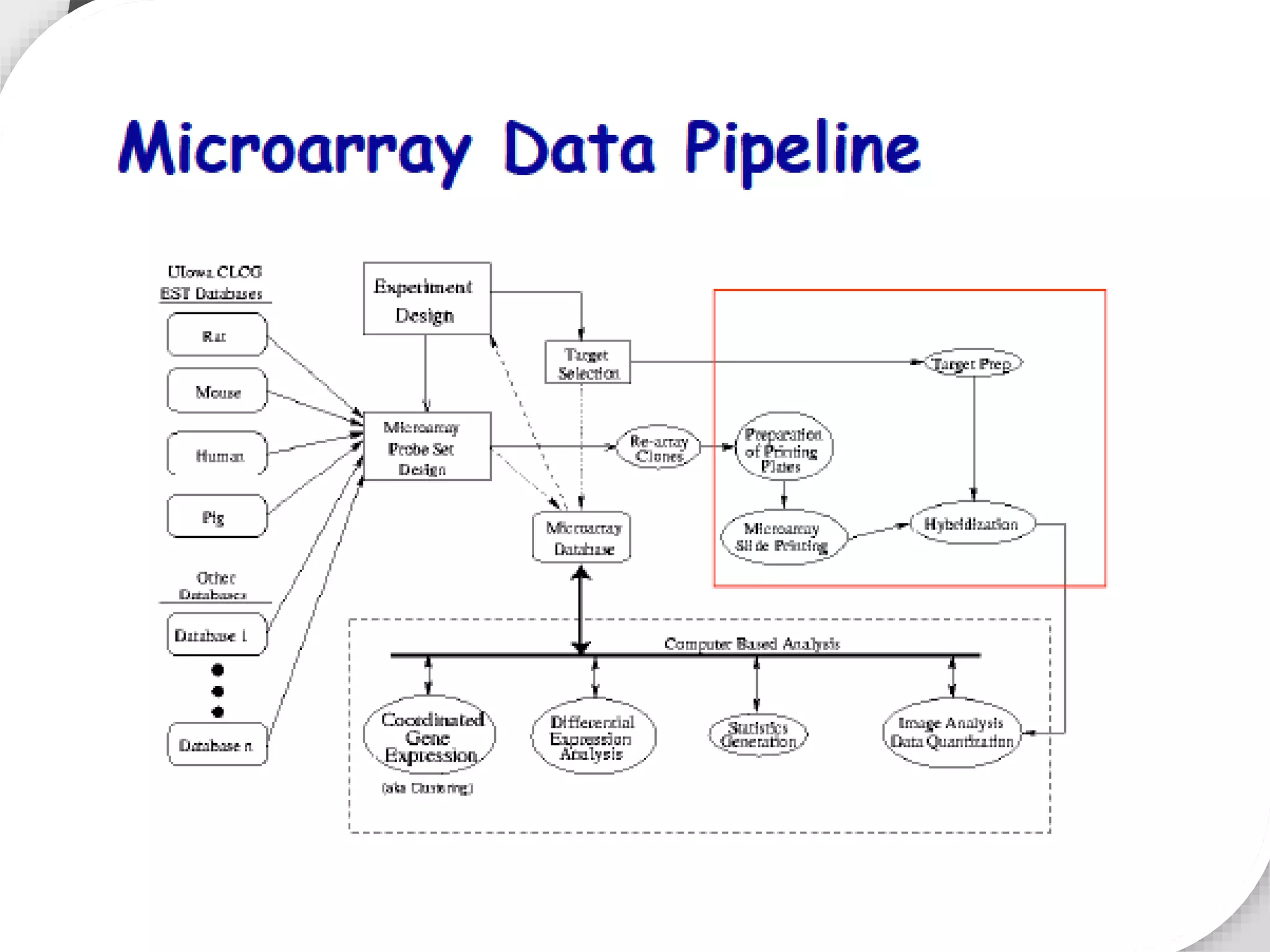
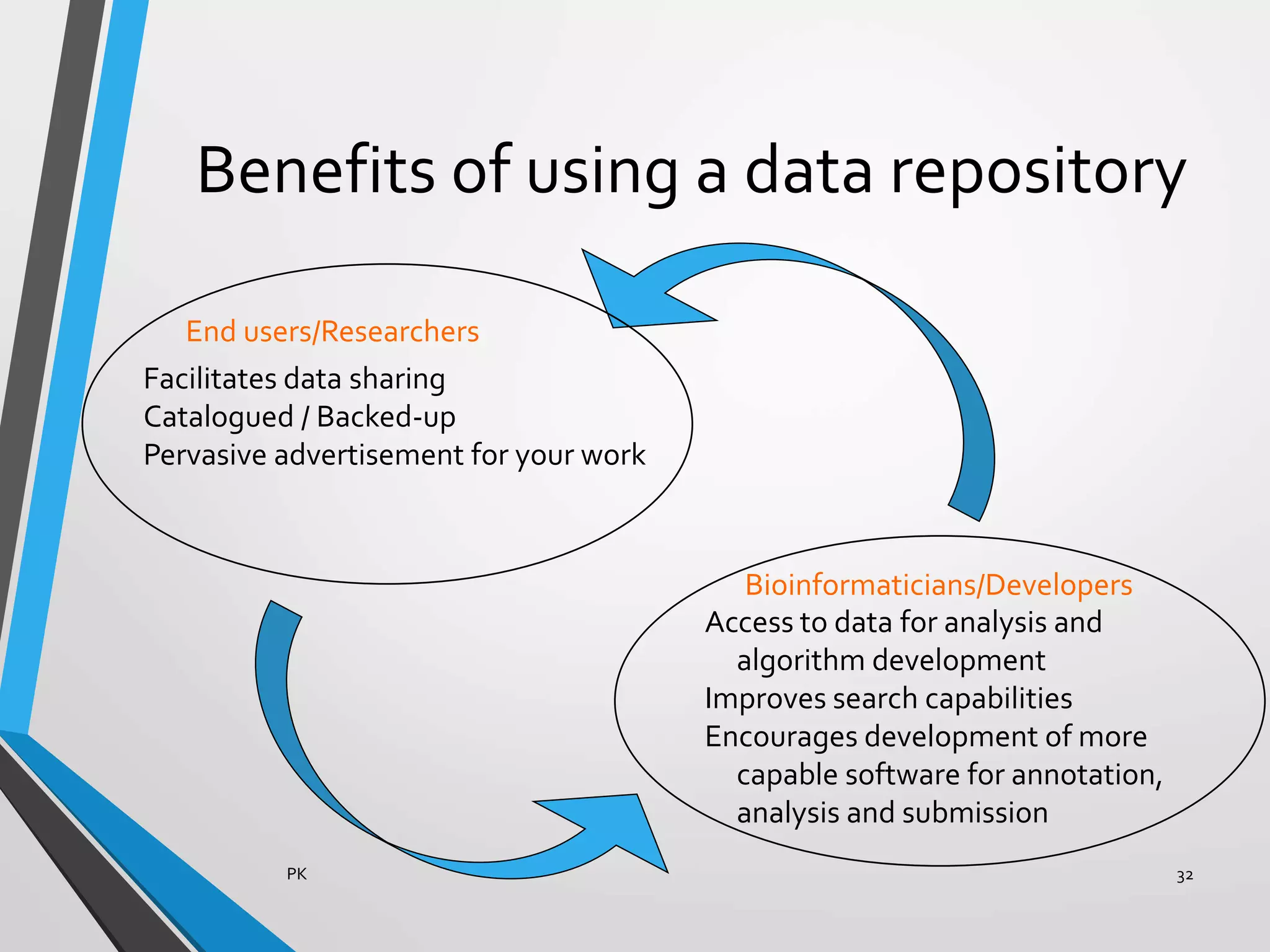
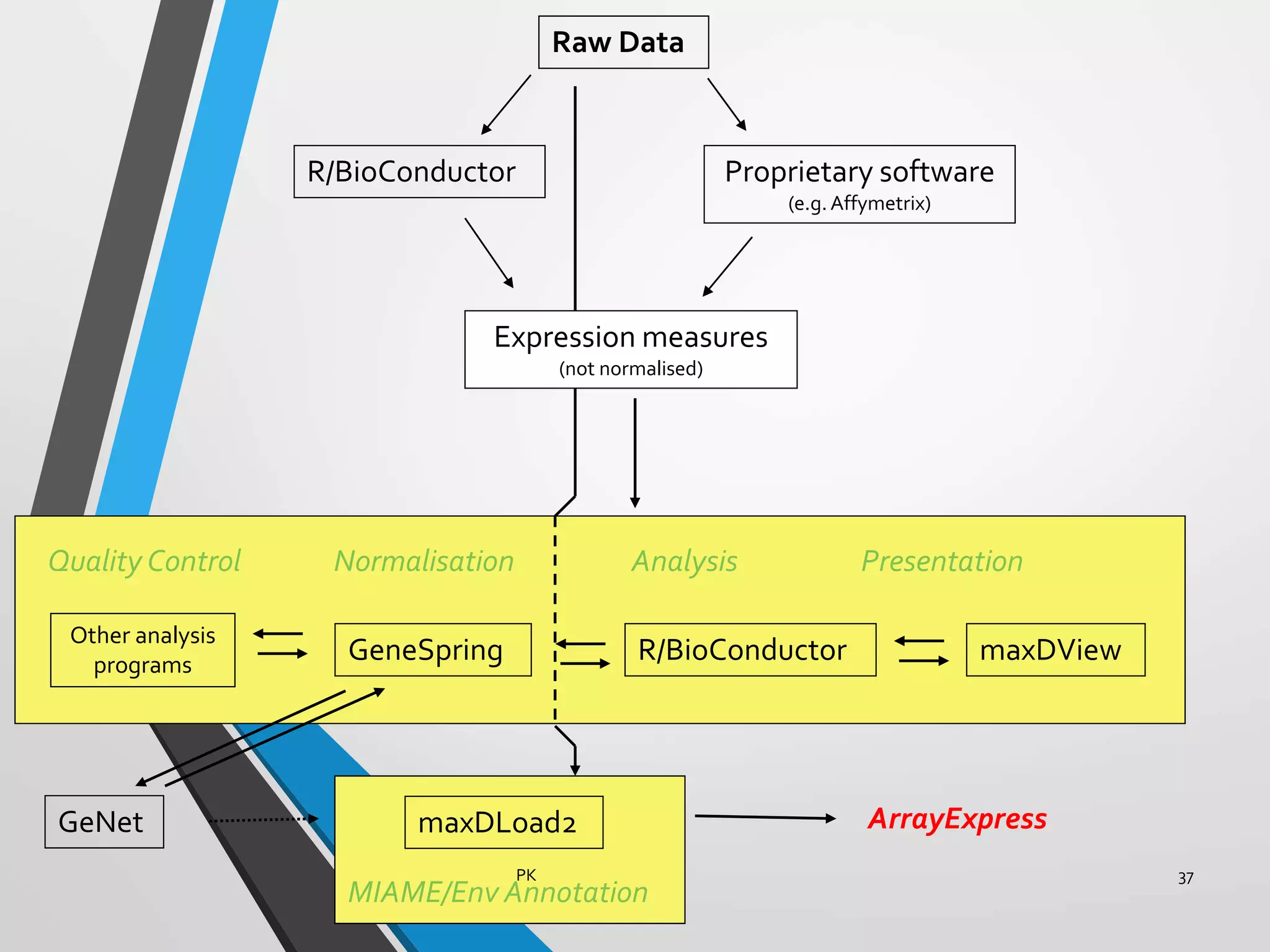
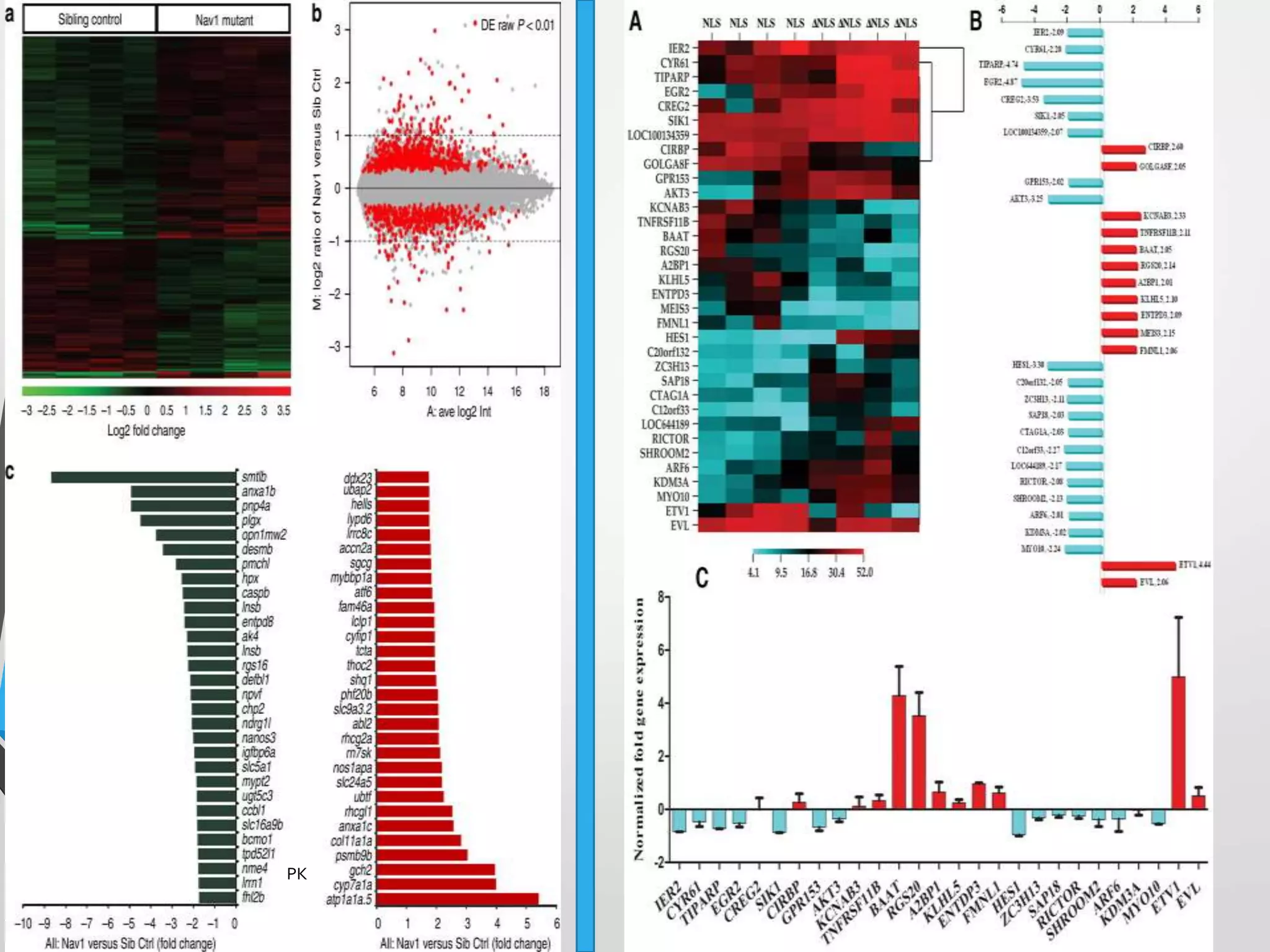
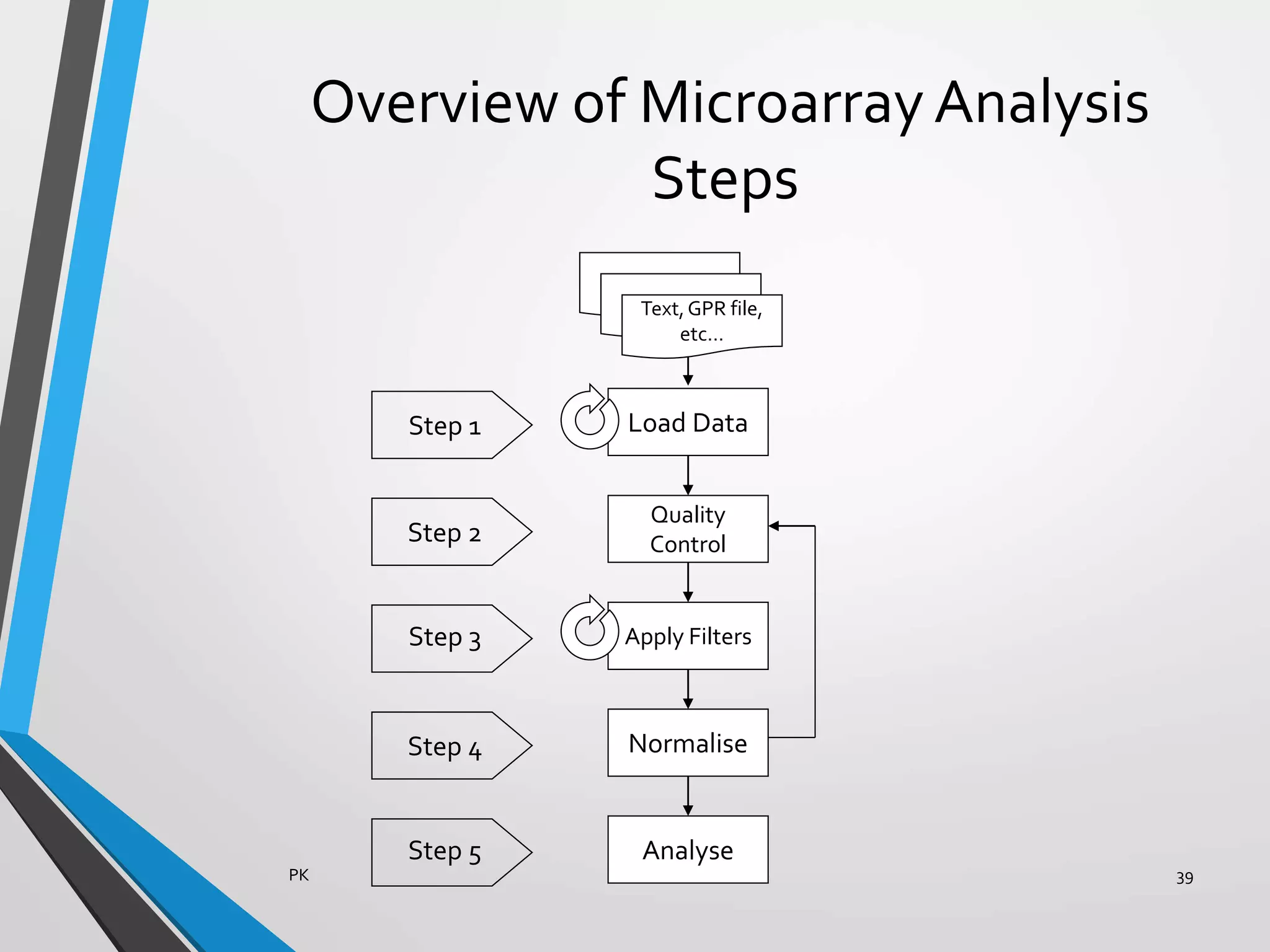
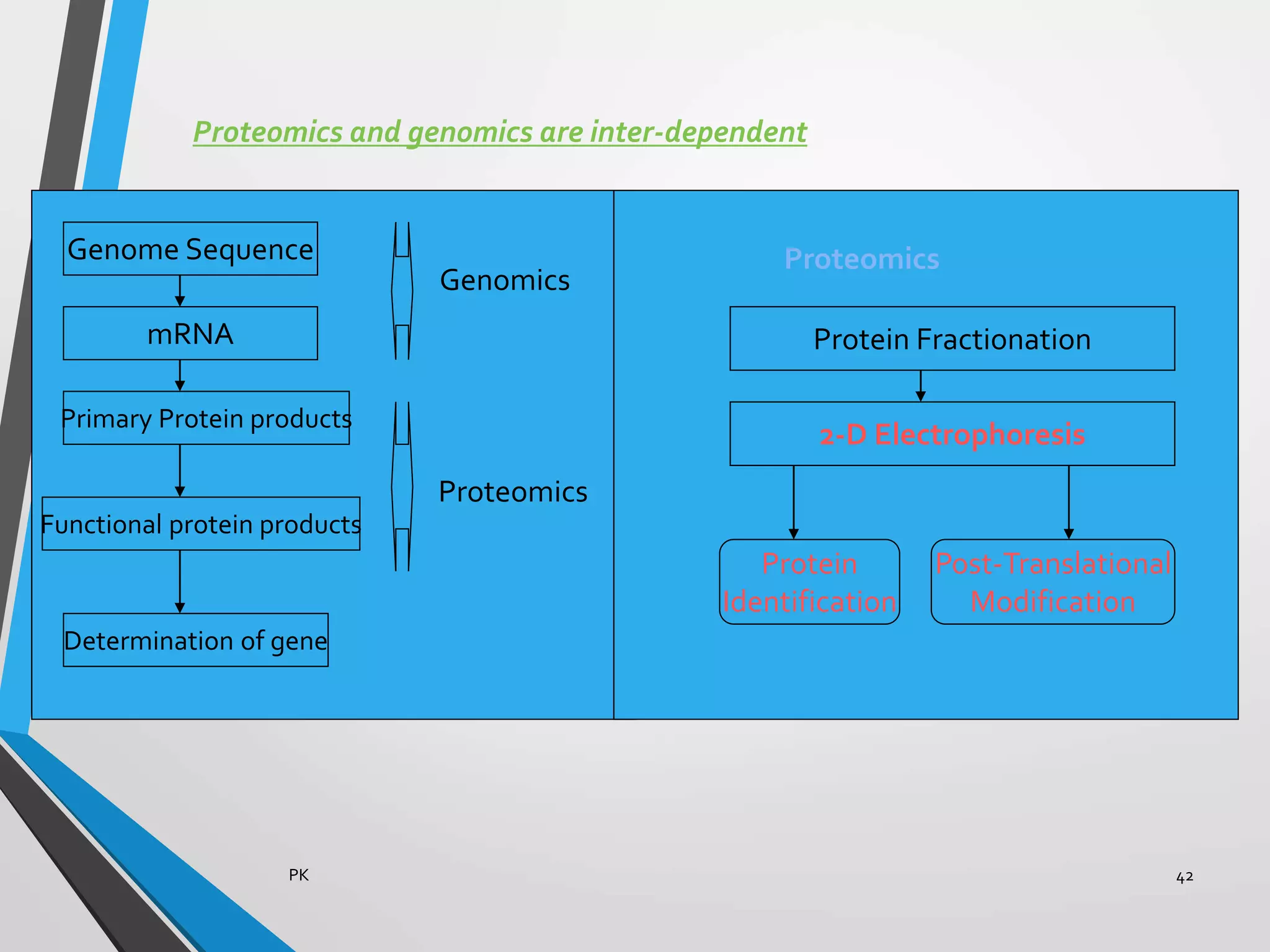
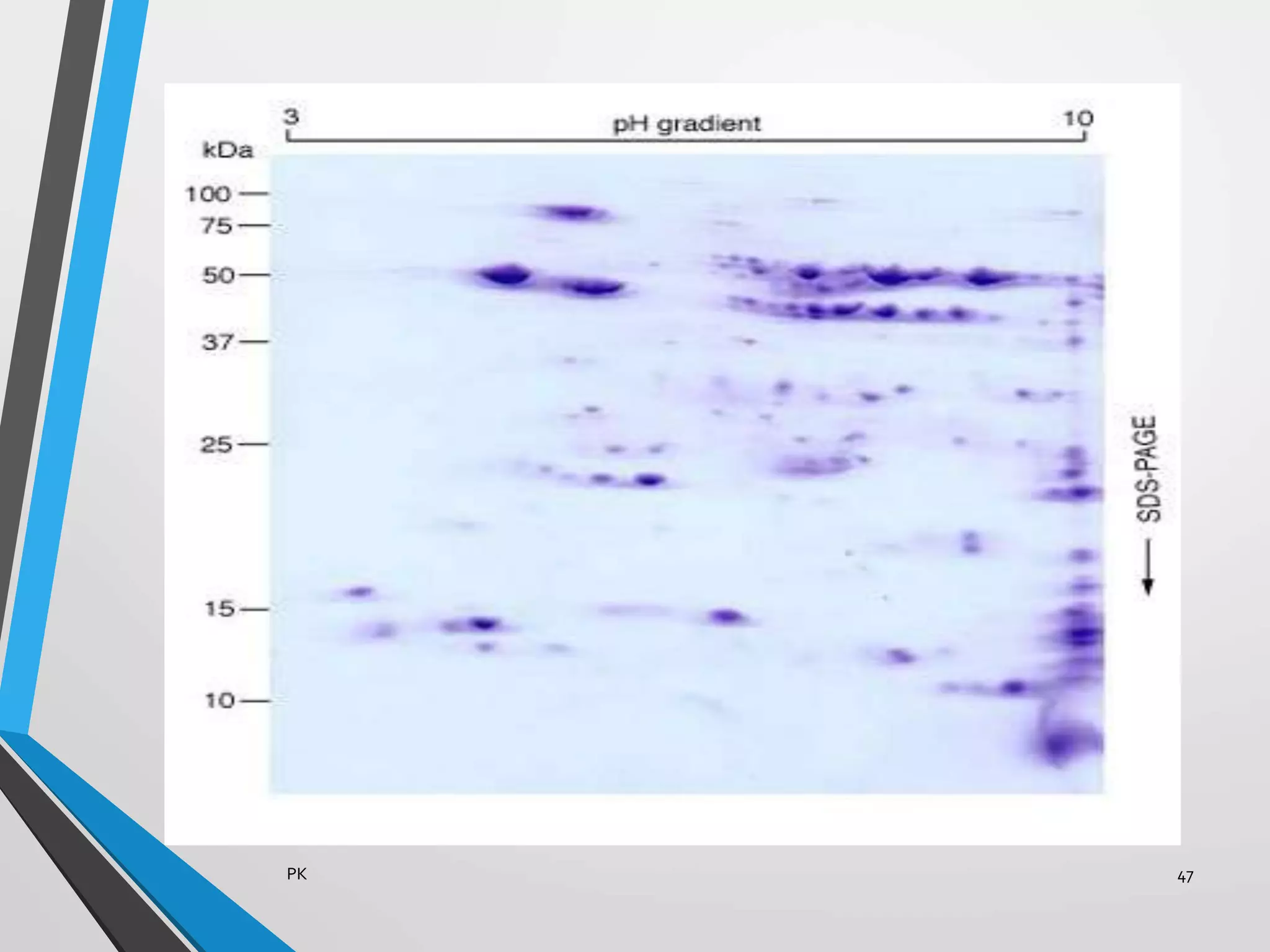
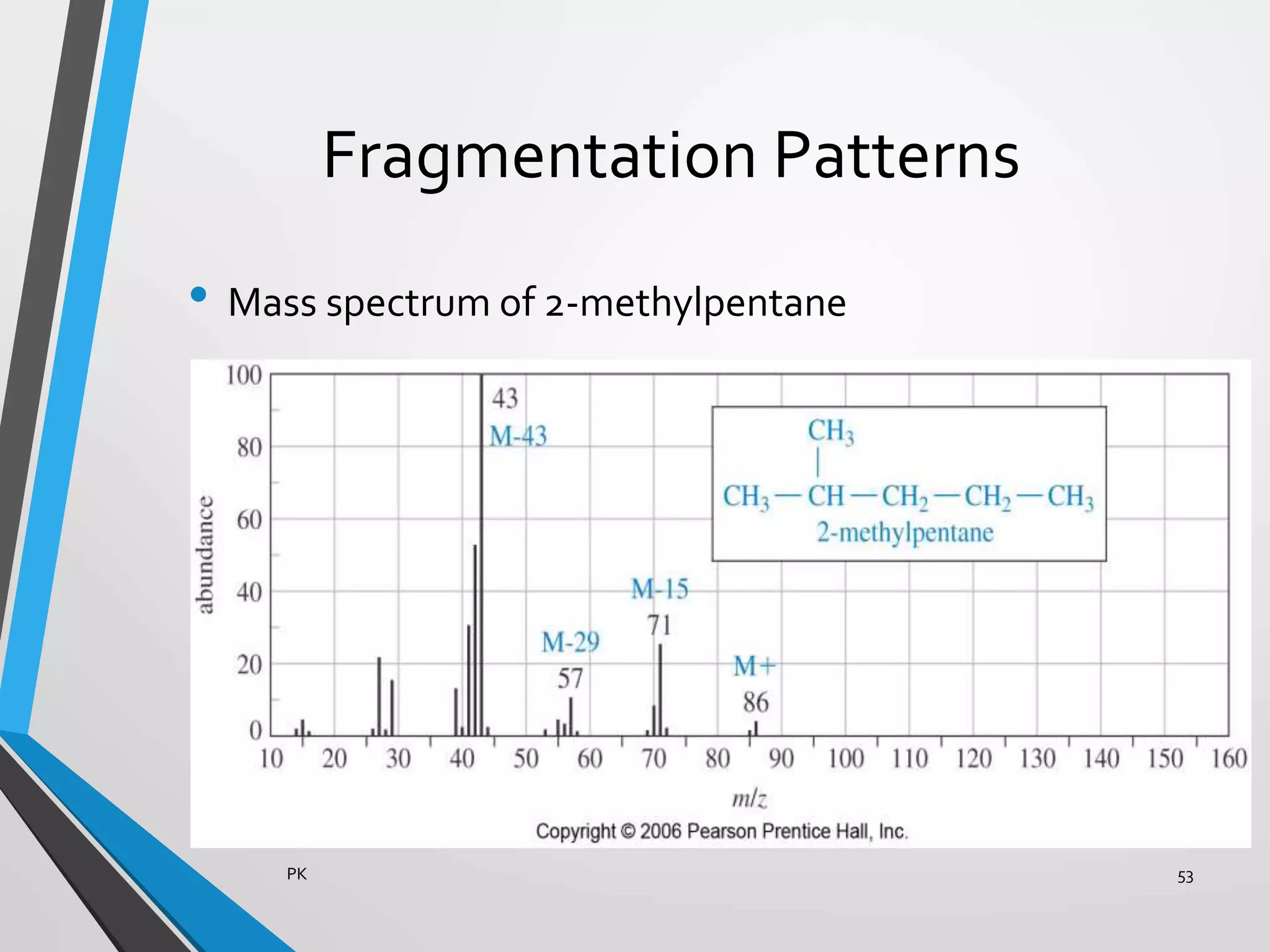
Functional genomics uses genome-wide experimental approaches to assess gene function on a large scale. It analyzes gene expression through techniques like transcriptomics and proteomics. Transcriptomics analyzes gene expression profiles through RNA sequencing or microarray analysis. Microarray analysis involves hybridizing fluorescently-labeled cDNA or cRNA to microarrays containing DNA probes to measure gene expression levels across thousands of genes simultaneously. Functional genomics provides a global understanding of gene function and molecular interactions through integrated omics approaches.