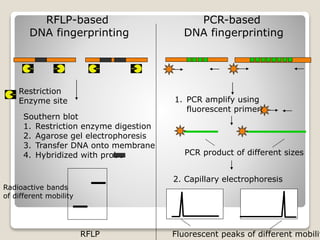
The document discusses DNA fingerprinting techniques for identifying plants at the molecular level. It describes methods such as PCR-based techniques like RAPD, ISSR, and SSR, as well as non-PCR based RFLP. The key steps are DNA isolation from plant tissues, amplification using PCR, restriction enzyme digestion, gel electrophoresis, and comparing banding patterns to determine differences between individuals. DNA fingerprinting can be used for applications like criminal investigations by matching DNA from crime scenes to suspects.