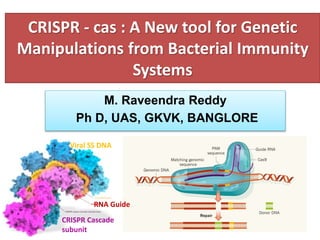
This document provides an overview of the CRISPR-Cas immune system in bacteria and its applications. It discusses:
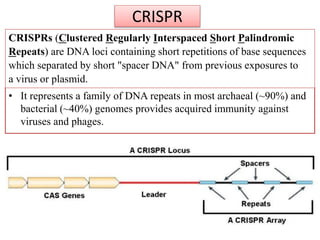
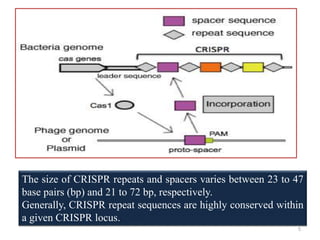
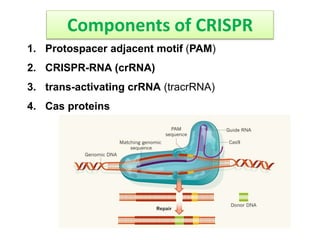
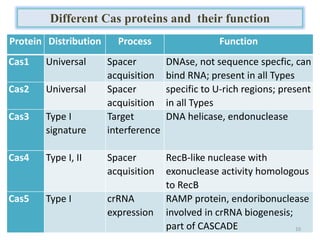
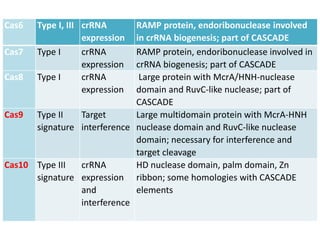
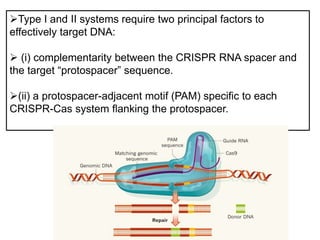
1. The history and components of the CRISPR-Cas system, including Cas proteins, CRISPR RNA, and protospacer adjacent motifs.
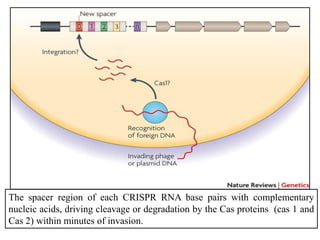
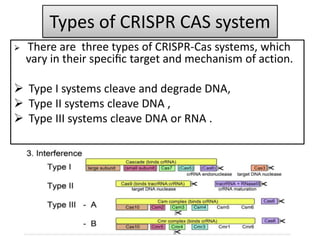
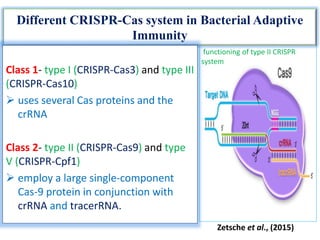
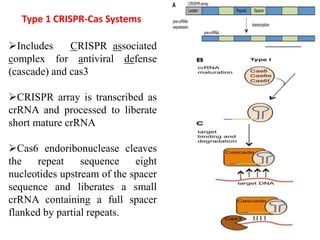
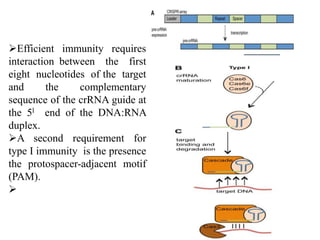
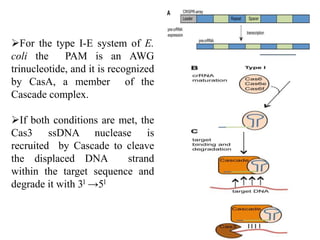
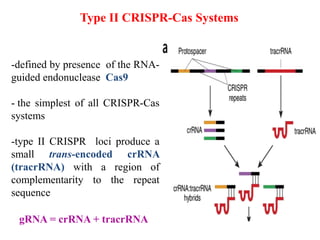
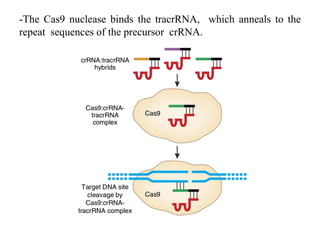
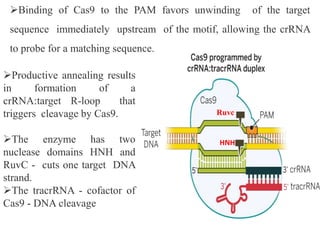
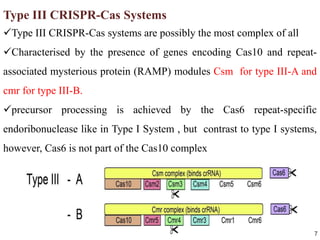
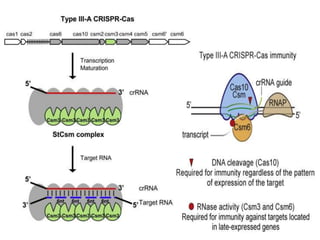
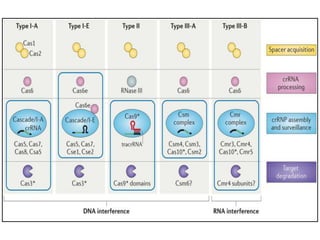
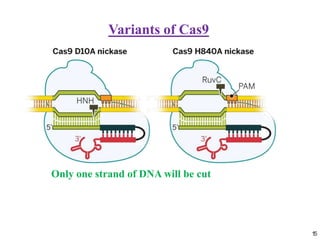
2. The three main types of CRISPR-Cas systems and their mechanisms of targeting DNA or RNA.
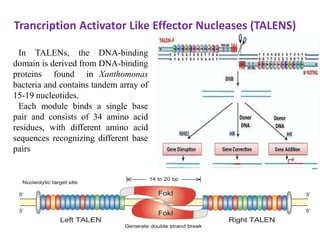
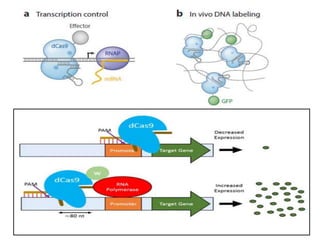
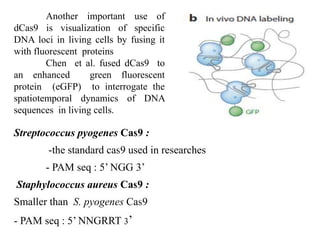
3. How engineered versions of Cas9 can be used for targeted genome editing and modulation of gene expression in mammalian cells.
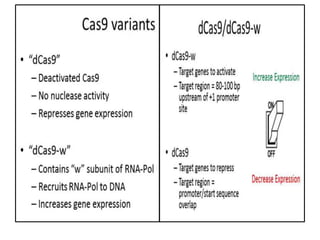
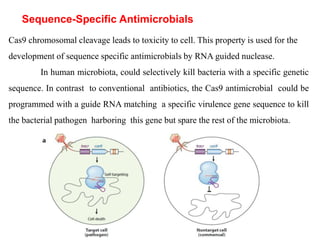
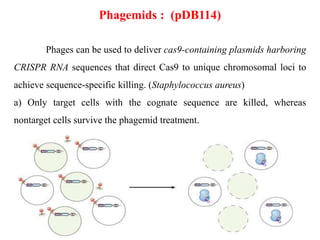
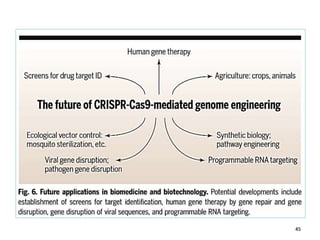
4. Applications of CRISPR-Cas in microbiology such as genetic engineering of bacteria, gene repression/activation using deactivated Cas9, and developing sequence-specific antimicrobials.