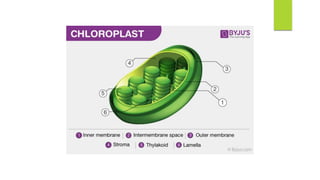
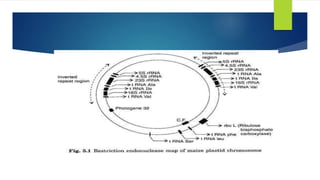
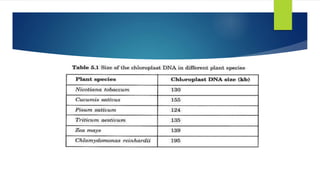
Chloroplasts are double-membrane organelles found in plant cells that contain chlorophyll and are the site of photosynthesis. Chloroplast DNA is circular and ranges in size from 120,000 to 170,000 base pairs. It contains approximately 120 genes, including genes that encode proteins involved in photosynthesis and the transcription and translation machinery. Chloroplast DNA replication is semi-conservative and there are typically multiple copies of the chloroplast genome within each chloroplast.



























![Case studies
Five Complete Chloroplast Genome Sequences from Diospyros: Genome Organization and
Comparative Analysis- journals.plos.org
Jianmin Fu et al[2016], https://doi.org/10.1371/journal.pone.0159566
Diospyros is the largest genus in Ebenaceae, comprising more than 500 species with remarkable economic value, especially Diospyros kaki Thunb., which has
traditionally been an important food resource in China, Korea, and Japan. Complete chloroplast (cp) genomes from D. kaki, D. lotus L., D. oleifera Cheng., D.
glaucifolia Metc., and Diospyros ‘Jinzaoshi’ were sequenced using Illumina sequencing technology. This is the first cp genome reported in Ebenaceae. The cp genome
sequences of Diospyros ranged from 157,300 to 157,784 bp in length, presenting a typical quadripartite structure with two inverted repeats each separated by one large
and one small single-copy region. For each cp genome, 134 genes were annotated, including 80 protein-coding, 31 tRNA, and 4 rRNA unique genes. In all, 179 repeats
and 283 single sequence repeats were identified. Four hypervariable regions, namely, intergenic region of trnQ_rps16, trnV_ndhC, and psbD_trnT, and intron of ndhA,
were identified in the Diospyros genomes. Phylogenetic analyses based on the whole cp genome, protein-coding, and intergenic and intron sequences indicated that D.
oleifera is closely related to D. kaki and could be used as a model plant for future research on D. kaki; to our knowledge, this is proposed for the first time. Further,
these analyses together with two large deletions (301 and 140 bp) in the cp genome of D. ‘Jinzaoshi’, support its placement as a new species in Diospyros. Both
maximum parsimony and likelihood analyses for 19 taxa indicated the basal position of Ericales in asterids and suggested that Ebenaceae is monophyletic in Ericales.](https://image.slidesharecdn.com/chloroplastgenomeorganisation-210531151521/85/Chloroplast-genome-organisation-33-320.jpg)
![The complete nucleotide sequence of the
tobacco chloroplast genome: its gene
organization and expression-embopress.org
K. Shinozaki et al[1986] https://doi.org/10.1002/j.1460-2075.1986.tb04464.x
The complete nucleotide sequence (155 844 bp) of tobacco (Nicotiana tabacum var. Bright Yellow 4) chloroplast DNA has been determined. It
contains two copies of an identical 25 339 bp inverted repeat, which are separated by a 86 684 bp and a 18 482 bp single‐copy region. The genes
for 4 different rRNAs, 30 different tRNAs, 39 different proteins and 11 other predicted protein coding genes have been located. Among them, 15
genes contain introns. Blot hybridization revealed that all rRNA and tRNA genes and 27 protein genes so far analysed are transcribed in the
chloroplast and that primary transcripts of the split genes hitherto examined are spliced. Five sequences coding for proteins homologous to
components of the respiratory‐chain NADH dehydrogenase from human mitochondria have been found. The 30 tRNAs predicted from their genes
are sufficient to read all codons if the ‘two out of three’ and ‘U:N wobble’ mechanisms operate in the chloroplast. Two sequences which
autonomously replicate in yeast have also been mapped. The sequence and expression analyses indicate both prokaryotic and eukaryotic features of
the chloroplast genes.](https://image.slidesharecdn.com/chloroplastgenomeorganisation-210531151521/85/Chloroplast-genome-organisation-34-320.jpg)


![Methods for Obtaining and Analyzing
Whole Chloroplast Genome Sequences
(science direct)
Robert K.Jansen et al.(2005)
During the past decade, there has been a rapid increase in our understanding of plastid genome organization and evolution due to the
availability of many new completely sequenced genomes. There are 45 complete genomes published and ongoing projects are likely to
increase this sampling to nearly 200 genomes during the next 5 years. Several groups of researchers including ours have been developing
new techniques for gathering and analyzing entire plastid genome sequences and details of these developments are summarized in this
chapter. The most important developments that enhance our ability to generate whole chloroplast genome sequences involve the generation
of pure fractions of chloroplast genomes by whole genome amplification using rolling circle amplification, cloning
genomes into Fosmid or bacterial artificial chromosome (BAC) vectors, and the development of an organellar annotation program (Dual
Organellar GenoMe Annotator [DOGMA]). In addition to providing details of these methods, we provide an overview of methods for
analyzing complete plastid genome sequences for repeats and gene content, as well as approaches for using gene order and sequence data
for phylogeny reconstruction. This explosive increase in the number of sequenced plastid genomes and improved computational tools will
provide many insights into the evolution of these genomes and much new data for assessing relationships at deep nodes in plants and other
photosynthetic organisms.](https://image.slidesharecdn.com/chloroplastgenomeorganisation-210531151521/85/Chloroplast-genome-organisation-37-320.jpg)
