This document discusses extra nuclear genomes, specifically chloroplast DNA and mitochondrial DNA. It provides details on:
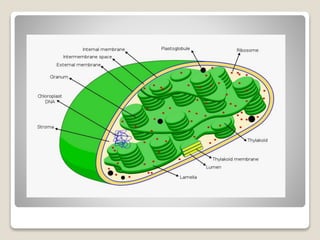
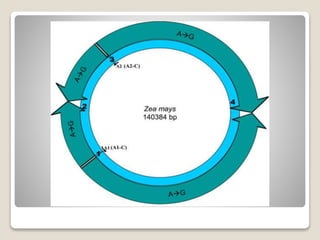
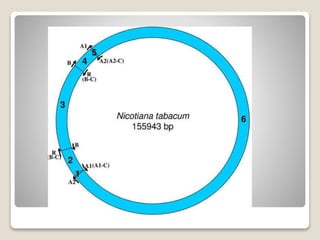
1) Chloroplast DNA is circular DNA ranging from 120-155kb that encodes around 120 genes and is present in multiple copies within chloroplasts.
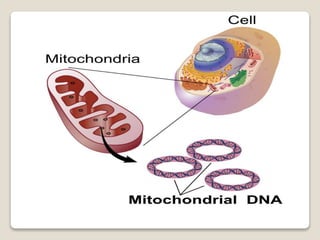
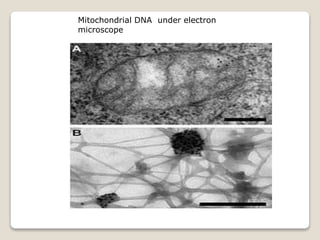
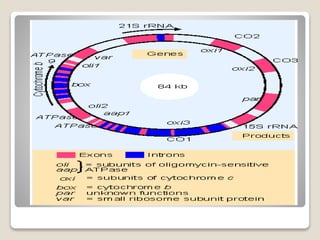
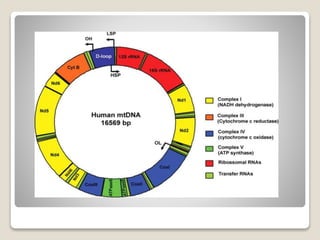
2) Mitochondrial DNA also exists as circular DNA that varies in size and encodes RNA and some proteins. In mammals it is 16.5kb while in plants it can be over 100kb.
3) Both organelle genomes are transcribed and translated within their respective organelles but rely on nuclear genes for some functions like replication machinery. They have their own genetic codes that differ slightly from nuclear codes.