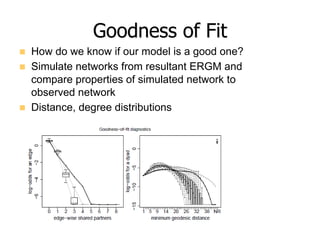
This document provides an overview of exponential random graph models (ERGMs) for statistically modeling social networks. It discusses the goals of using ERGMs, which are to understand structural features of networks, test hypotheses about network formation processes, and link macro network structures to micro behaviors. Example model terms that can be used in ERGMs are described, ranging from simple models with just edges to more complex models incorporating triangles, degree distributions, and homophily. The document outlines the challenges of estimating ERGM parameters using maximum likelihood due to the normalizing constant, and notes that simulation-based approximations are typically used instead.






![Figure 1. The largest connected component of
One mesoscopic s
consists of a grou
densely connected
nected to other de
We illustrate this
known benchmark
literature [131].
The existence o
itively clear, and th
have been studied
ogy [25, 44, 79] and
For example, Stua
to investigate poli
and George Homa
of rearranging th
matrices to reveal
1950 [60]. Robert W
formed (using org
have been the firs
nity structure in 1
espoused surprisin
ty structure and co
1960s [117]. Socia
arising in the flo
cial organizations
Coauthorship between
scholars in physics literature
Slide Credit: Porter et al. 2009](https://image.slidesharecdn.com/08ergmsnhdukepresentation-160930144534-170629181939/85/08-Exponential-Random-Graph-Models-2016-9-320.jpg)






















![MLE Estimation
n Likelihood formula:
n Want to estimate unknown parameters ,θ, given the
observed network
n Can’t get ML estimates directly (because the
normalizing constant can’t be enumerated) so
approximate by simulation
Journal of Statistical Software
aximize `(✓) directly, we will consider instead the log-ratio of likelihood v
`(✓) `(✓0) = (✓ ✓0)>
g(yobs) log
(✓, Y)
(✓0, Y)
,
arbitrarily chosen parameter vector. [Note: Previously in this article, we
the term “coe cient” in situations in which either “coe cient” or “param
ally be correct. To be precise, a coe cient in this context is a specific
namely, one that is multiplied by a statistic as in this case or in the
enerally. In this section, we may use the terms “parameter” and “coe
y.]
ation of ratios of normalizing constants such as the one in expression (
well-studied problem (Meng and Wong 1996; Gelman and Meng 1998).
xploit in the ergm function is due to Geyer and Thompson (1992) and m
llows: Starting from Equations 1 and 2, a bit of algebra reveals that
(✓, Y) n
>
o](https://image.slidesharecdn.com/08ergmsnhdukepresentation-160930144534-170629181939/85/08-Exponential-Random-Graph-Models-2016-33-320.jpg)