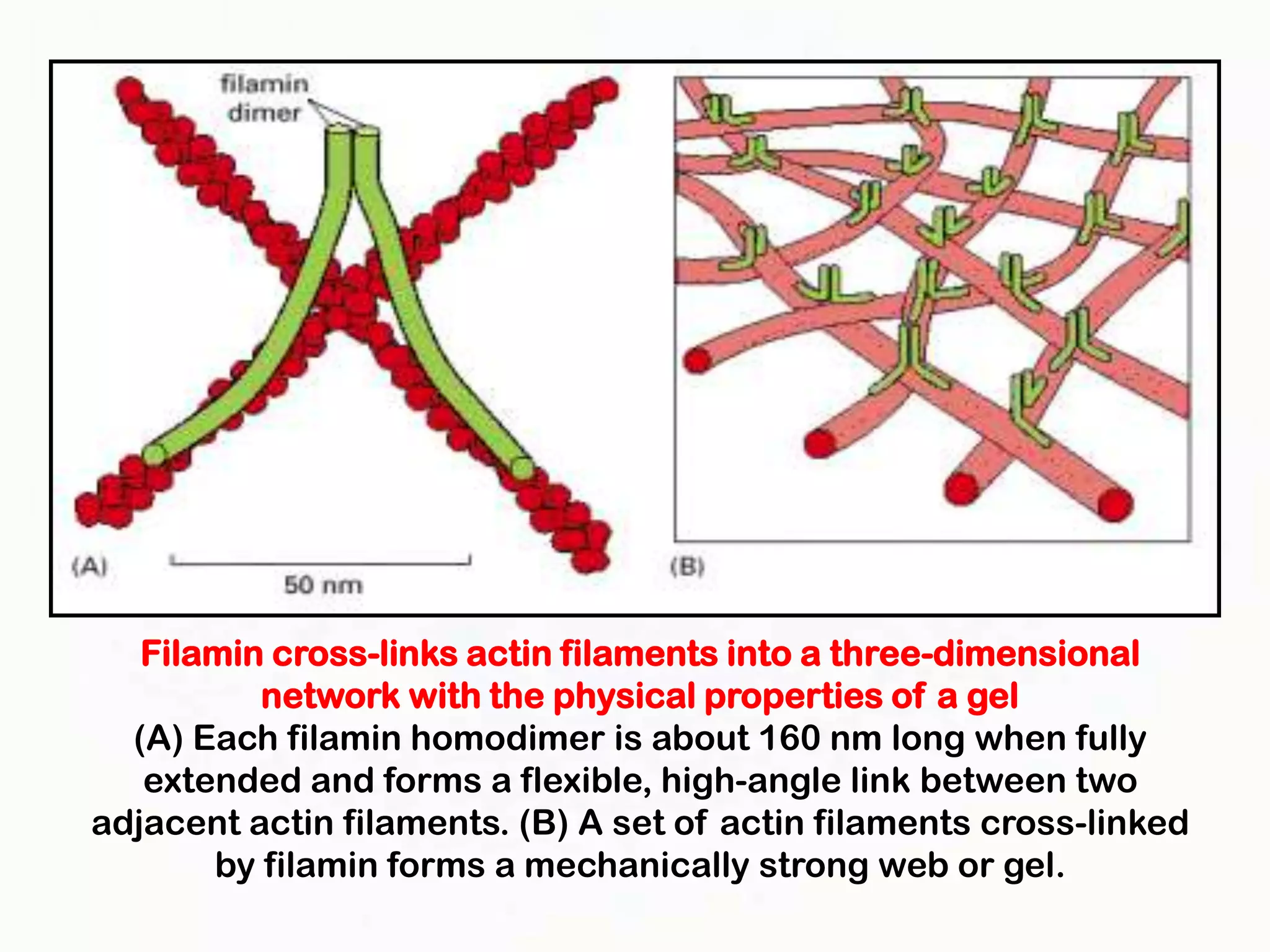
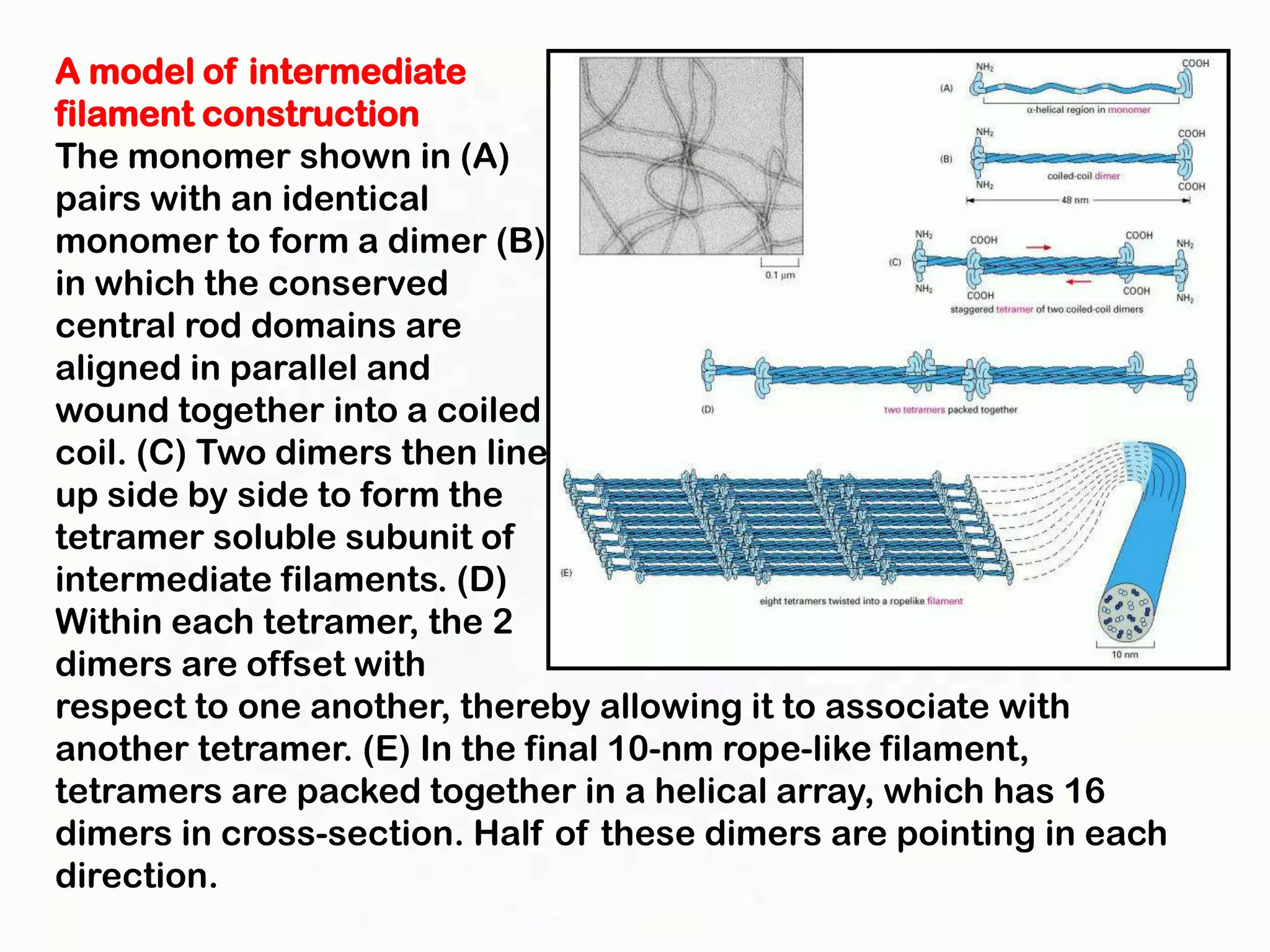
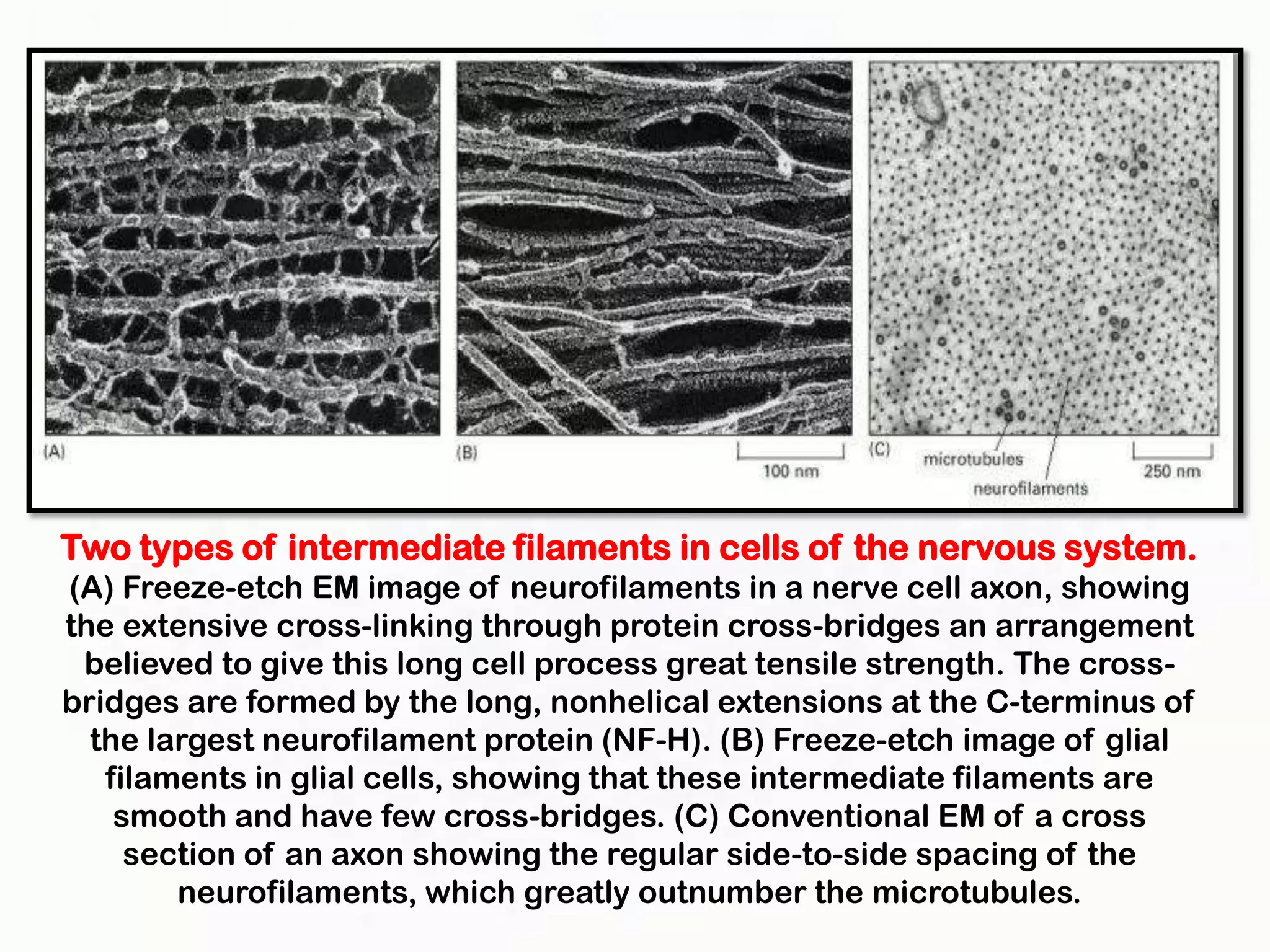
The document summarizes key aspects of the cytoskeleton, focusing on actin filaments. It describes how actin filaments: 1) maintain cell shape and generate force for cell movements through polymerization and depolymerization; 2) integrate cells through attachments to cell adhesions; and 3) produce movements within cells and at the cell membrane through interactions with myosin motor proteins. Accessory proteins regulate actin dynamics by capping, bundling, severing, and crosslinking filaments.