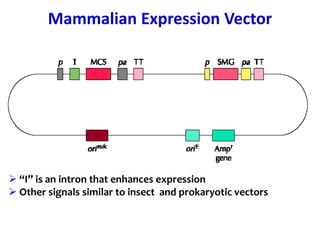
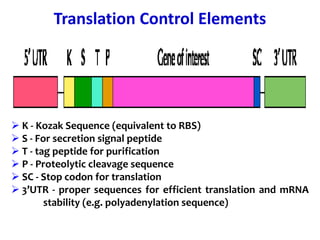
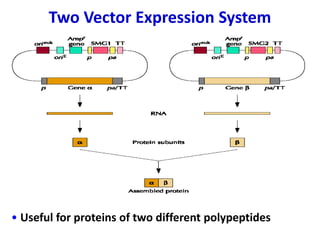
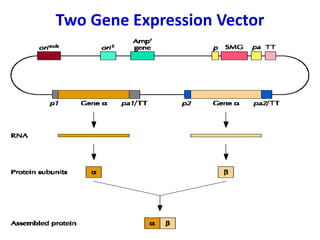
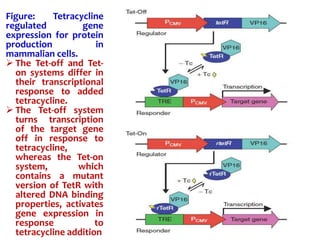
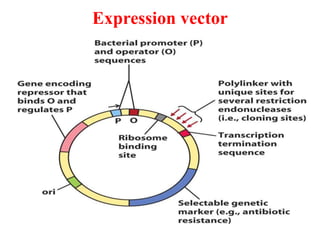
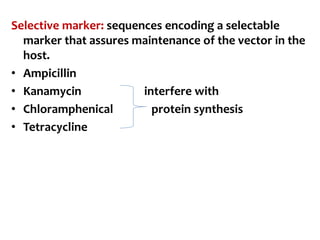
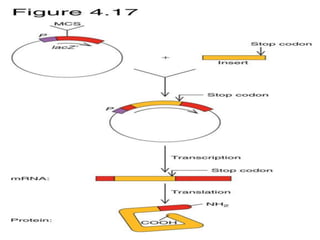
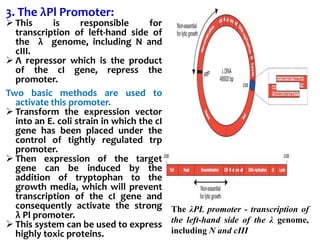
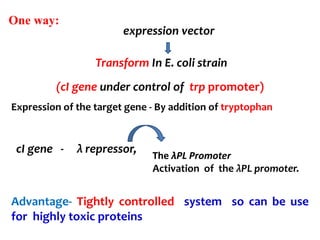
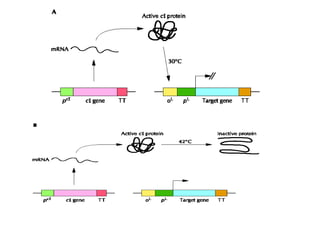
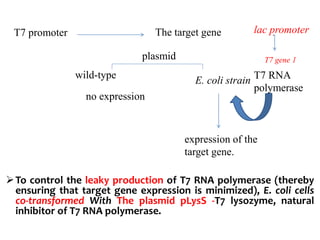
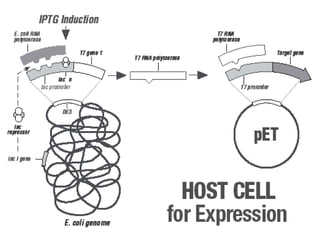
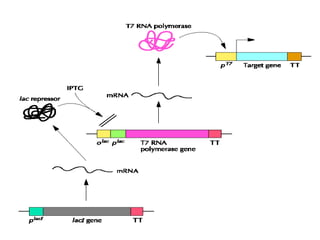
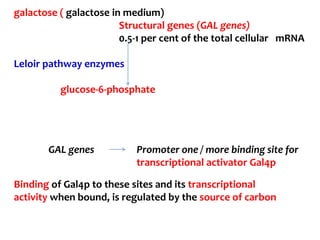
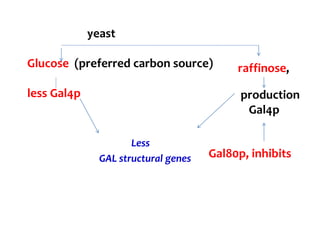
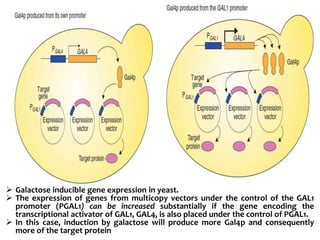
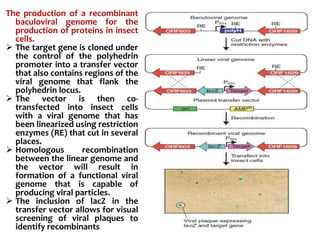
The document discusses different expression vectors and systems used for recombinant protein expression. It describes key elements required for an expression vector including an origin of replication, selective marker, promoter, multiple cloning site, and terminator. It provides details on commonly used expression systems in E. coli such as the lac, tac, lambda PL, and T7 promoters. It also summarizes protein expression in yeast using the galactose-inducible GAL promoter system.




































![Expression in Higher-Eukaryotic Cells
For the production of mammalian proteins, mammalian cells
have an obvious advantage.
Mammalian cell lines derived from humans or hamsters have
been used in synthesis of several recombinant proteins, and in
most cases these proteins have been processed correctly and
are indistinguishable from the non-recombinant versions.
Two modes of expression - transient and stable.
Three cell types are dominant in transient expression: human
embryonic kidney (HEK), COS and baby hamster kidney (BHK),
whilst CHO (Chinese hamster ovary) cells are used
predominantly for stable expression.
Note: The acronym "COS" is derived from the cells being CV-1
(simian) in Origin, and carrying the SV40 genetic material.[2] Two
forms of COS cell lines commonly used are COS-1 and COS-7.](https://image.slidesharecdn.com/expressionvectors-180609200652/85/Expression-vectors-58-320.jpg)