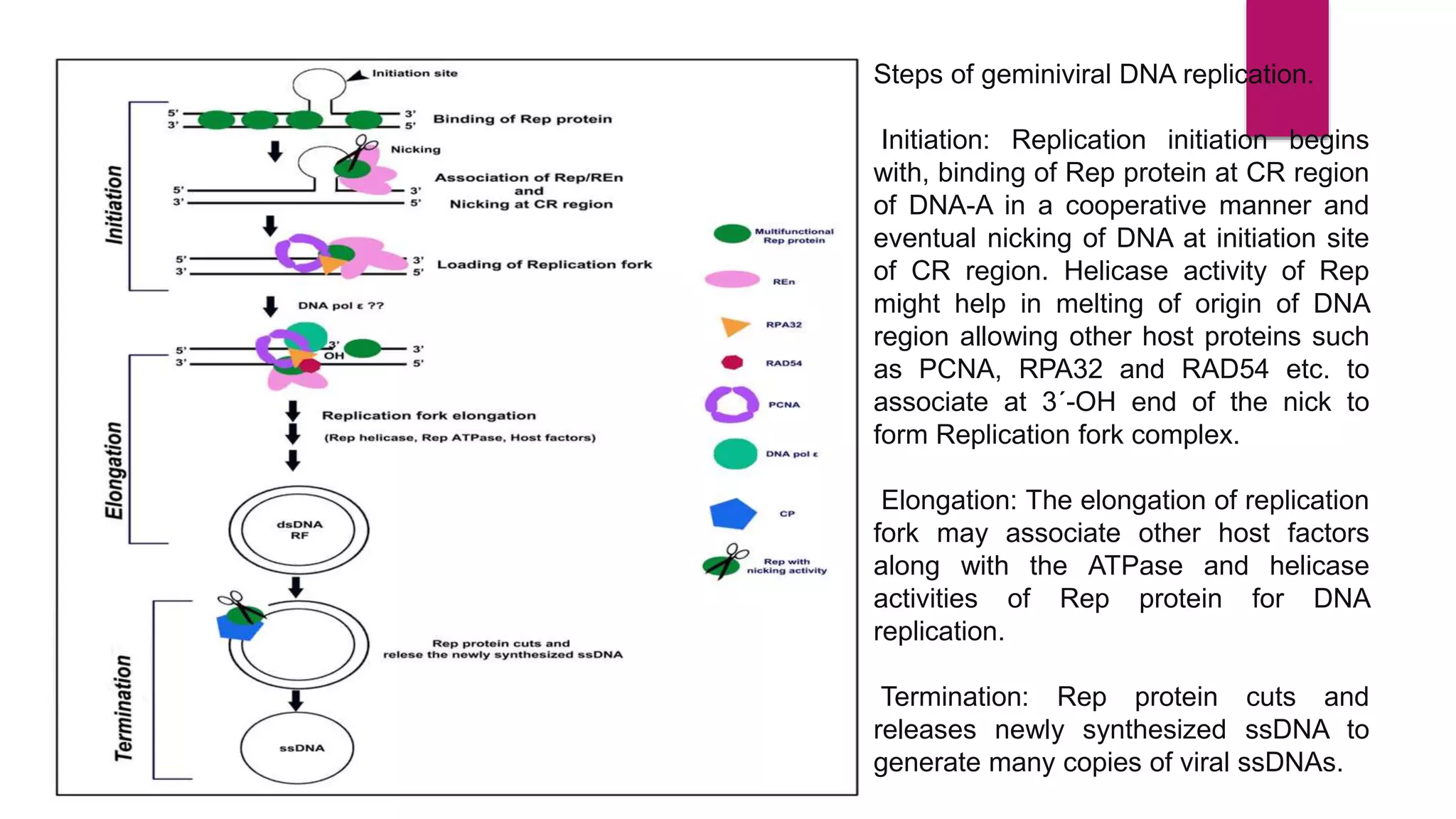
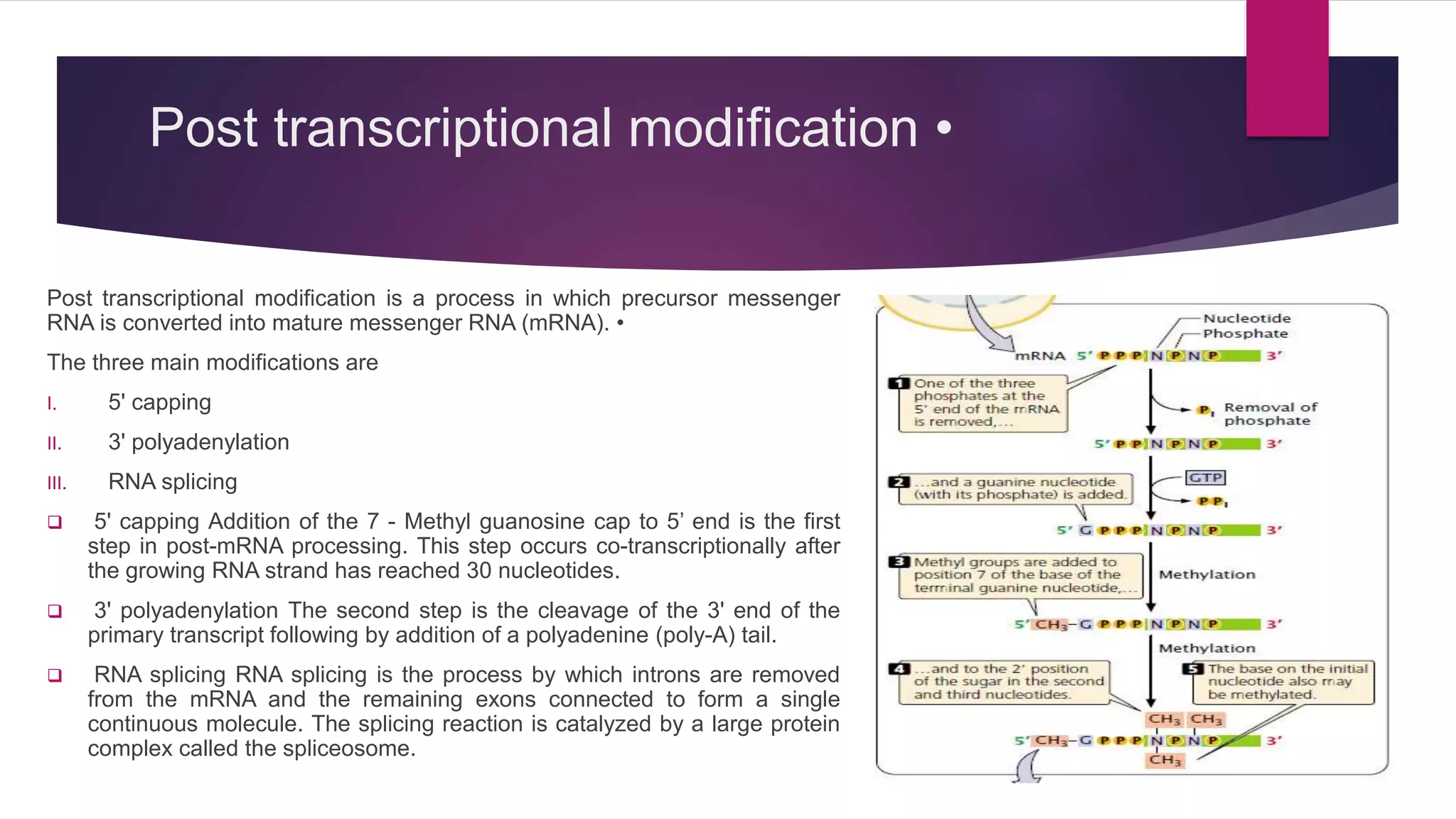
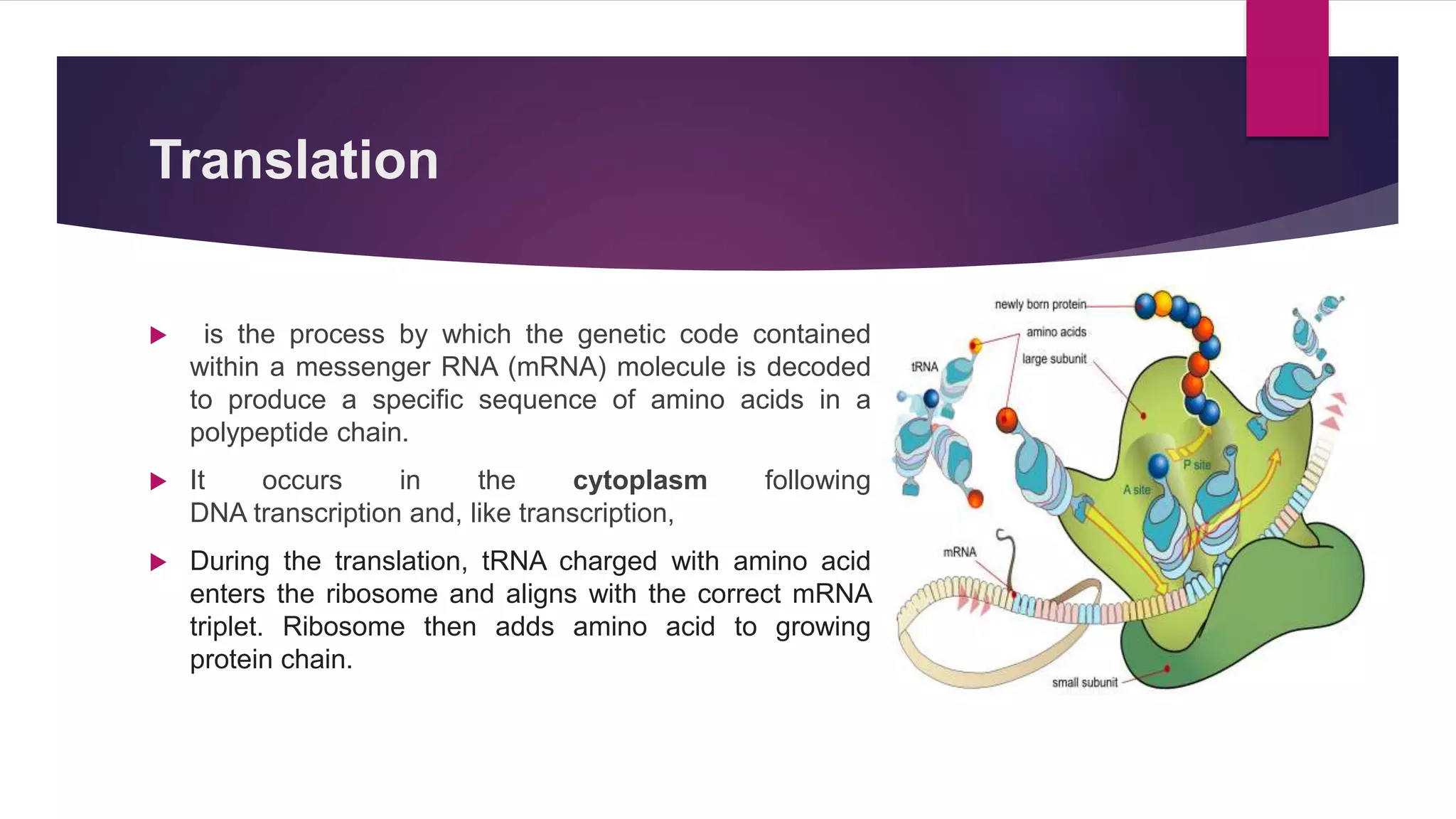
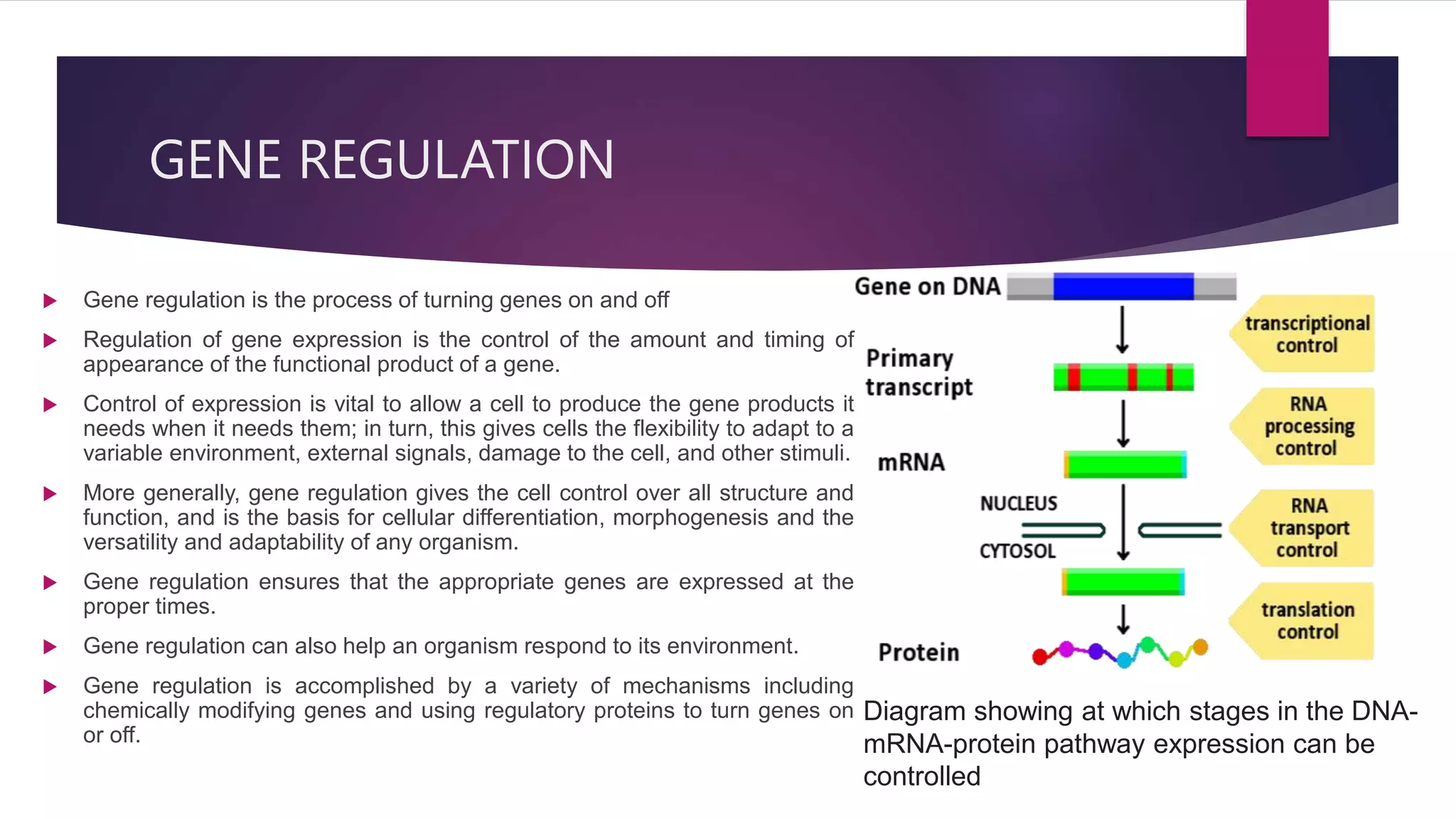
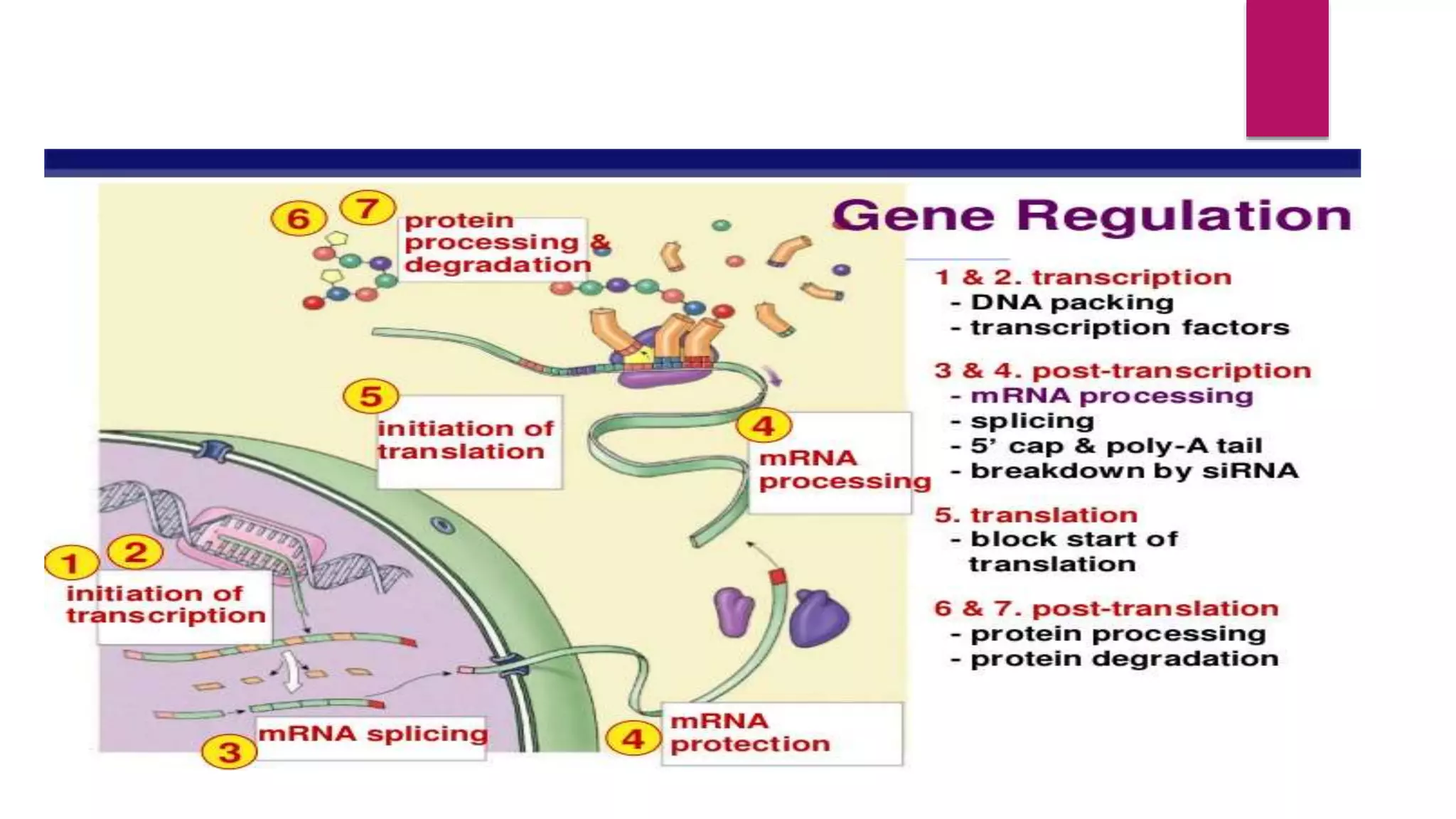
Gene expression is the process by which information from a gene is used to direct protein synthesis. It involves replication, transcription, and translation. Regulation of gene expression controls the amount and timing of gene products. Regulation occurs through transcription factors binding to regulatory elements near gene promoters to turn genes on and off. This precise control of gene expression is vital for cellular differentiation and adaptation.