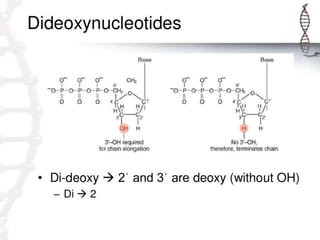
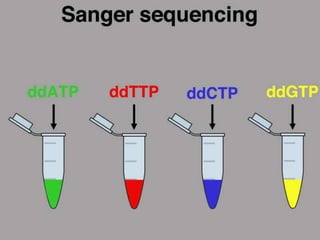
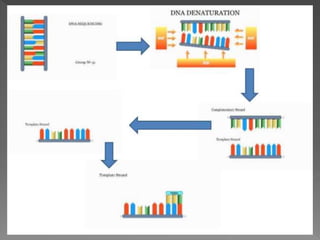
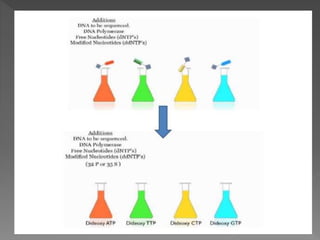
DNA sequencing is the process of determining the order of nucleotides in a DNA molecule. The Sanger method, developed in 1977, was the most widely used sequencing technique for 25 years. It utilizes chain termination with dideoxynucleotides which lack a 3' OH group, preventing formation of a phosphodiester bond and terminating strand elongation. Four reactions are run in parallel with each dideoxynucleotide labeled with a different color. Gel electrophoresis separates the terminated fragments by size, allowing the DNA sequence to be read by matching fragment sizes to nucleotide colors.