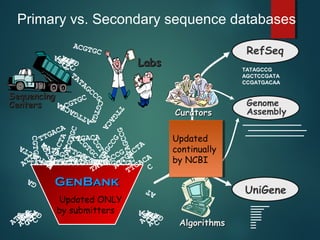
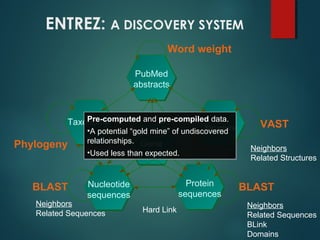
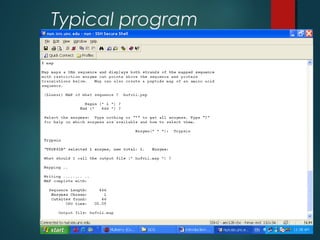
This document provides an introduction to biological databases. It discusses primary databases like GenBank which contain original sequence submissions and secondary databases derived from primary data, maintained by third parties like NCBI. Some key databases mentioned include GenBank, PDB, Swiss-Prot. The document also provides an overview of the NCBI and Entrez retrieval system, which allows integrated searches across literature and sequences.