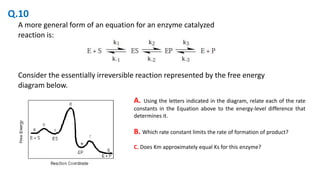
The document discusses various concepts of enzyme kinetics, including reaction orders, Michaelis constant (Km), turnover number (kcat), and steady-state approximation. It also outlines the conditions under which Km equals the rate constant Ks, as well as the effects of different factors on enzyme activity such as substrate concentration and inhibitors. Moreover, the document features problem-solving examples related to enzyme activity and kinetics, ultimately aiding in the understanding of enzyme catalysis.















![• a. True. Vmax = k2[E]t
• b. True
• c. False. The value of Km is independent of enzyme concentration for almost all enzymes
• d. False. A non-competitive inhibitor cannot be overcome by substrate concentration.
• e. True. This occurs in regulatory enzymes.
• f. True.
• g. False. The initial increasing slope of the curve shows that binding of the first substrate
molecule increases the affinity of the enzyme for subsequent substrate molecules.
A.7](https://image.slidesharecdn.com/enzymetechnologysolvedproblems-200407090146/85/Enzyme-technology-solved-problems-16-320.jpg)
![1. The _____________ of a reaction is the numerical relationship between
substrates and products
2. The rate constant ____________ of an enzyme-catalysed reaction is a measure
of the catalytic efficiency at saturating
levels of substrate.
3. _______________ inhibitors do not alter the Vmax of an enzyme-catalyzed
reaction.
4. The sigmoidal shape of the v versus [S] curve for some regulatory enzymes
results from a _______________ effect of substrate on the substrate binding sites.
5. For an enzyme whose Km can be regulated, the presence of a _____________
effector increases the level of substrate required to attain a given reaction rate.
Q.8](https://image.slidesharecdn.com/enzymetechnologysolvedproblems-200407090146/85/Enzyme-technology-solved-problems-17-320.jpg)



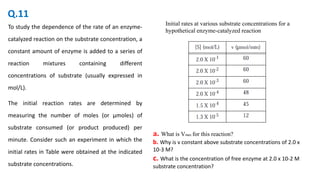

![To study the dependence of the rate of an enzyme-catalyzed
reaction on the substrate concentration, a constant amount of
enzyme is added to a series of reaction mixtures containing
different concentrations of substrate (usually expressed in mol/L).
Total reaction mixture volume is 10mL
The initial reaction rates are determined by measuring the
number of moles (or μmoles) of substrate consumed (or product
produced) per minute. Consider such an experiment in which the
initial rates in Table were obtained at the indicated substrate
concentrations.
A. What is Vmax for this concentration of enzyme?
B What is the Km of this enzyme?
C. Show that this reaction does or does not follow simple
Michaelis-Menten kinetics.
D. What are the initial rates at [S] = 1.0 x 10-6 M and at
[S] = 1.0 x 10-1 M?
E. Calculate the total amount of product made during the first
five minutes when [S] = 2.0 x 10-3 M. Could you make the
same calculation at [S] = 2.0 X 10-6 M?
F. Suppose that the enzyme concentration in each reaction
mixture were increased by a factor of 4. What would be the
value of Km? of Vmax? What would be the value of v at [S]
= 5.0 x 10-6 M?
Q.12](https://image.slidesharecdn.com/enzymetechnologysolvedproblems-200407090146/85/Enzyme-technology-solved-problems-25-320.jpg)
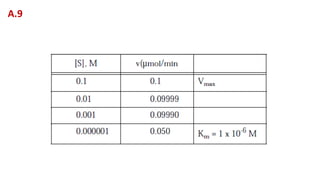
![A. Vmax = 0.25 μmol/min
B. For a reaction obeying Michaelis-Menten kinetics, Vmax and Km are
simply constants relating v to [S]. Km can be calculated by substituting
Vmax and any pair of v and [S] values at v < Vmax. For example, at [S] =
5.0 x 10-5 M and v = 0.20 μmol/min the equation becomes
A.12](https://image.slidesharecdn.com/enzymetechnologysolvedproblems-200407090146/85/Enzyme-technology-solved-problems-26-320.jpg)
![C. If the reaction follows simple Michaelis-Menten kinetics, then the
Michaelis-Menton equation should relate v to [S] over a wide range of [S].
This can be tested by determining whether the equation yields the same
value of Km at several different values of [S] and v < Vmax. Under the
conditions of this problem, the same value, Km = 1.3 x 10-5 M, is obtained at
[S] = 5.0 x 10-6 M, v = 0.071 μmol/min and at [S] = 5.0 x 10-7 M, v= 0.0096
μmol/min. Therefore, Michaelis-Menten kinetics are obeyed
A.12. Cont.](https://image.slidesharecdn.com/enzymetechnologysolvedproblems-200407090146/85/Enzyme-technology-solved-problems-27-320.jpg)

![E. At [S] = 2.0 x 10-3 M, v = Vmax = 0.25 μmol/min. Since 0.25 μmole is much less than the amount
of substrate present (2.0 x 10-3 mole/liter x 10-2 L x 106 μmol/mol = 20 μmol) the reaction can
proceed for five minutes without significantly changing the substrate concentration.
Thus, 0.25 μmol/min x 5 min = 1.25 μmol
At [S] = 2.0 x 10-6 M,
During 5 minutes at this rate, 0.033 μmol/min x 5 min = 0.17 μmol of product would be produced.
However, this value exceeds the total amount of substrate present (2.0 x 10-6 mol/L x 10-2 L x 106
μmol/mol = 0.020 μmol). Clearly, during the-5 min reaction, [S] and therefore v would decrease
significantly. Calculation of the exact amount of product made would require integration of a
differential equation; this amount obviously cannot exceed 0.020 μmole
A.12 Cont.](https://image.slidesharecdn.com/enzymetechnologysolvedproblems-200407090146/85/Enzyme-technology-solved-problems-29-320.jpg)
![F. Km is independent of enzyme concentration, since a change in [E]
does not affect the three rate constants, k1, k2, and k3.
Hence Km would remain equal to 1.25 x 10-5 M.
Since Vmax = k3[E]o, increasing the enzyme concentration by a factor
of 4 increases Vmax by a factor of 4. Therefore, Vmax = 1.0 μmole/min.
At [S] = 5.0 x 10-6 M,
A.12 Cont.](https://image.slidesharecdn.com/enzymetechnologysolvedproblems-200407090146/85/Enzyme-technology-solved-problems-30-320.jpg)
![The Km of a certain enzyme is 1.0 X 10-5 M in a reaction that is described by Michaelis-Menten kinetics. At a
substrate concentration of 0.10 M, the initial rate of the reaction is 37 μmol/min for a certain concentration of
enzyme. However, you observe that at a lower substrate concentration of 0.010 M the initial reaction rate remains 37
μmoles/min.
• a. Using numerical calculations, show why this tenfold reduction in substrate concentration does not alter the
initial reaction rate.
• b. Calculate v as a fraction of Vmax for [S] = 0.20 Km, 0.50 Km, 1.0 Km, 2.0 Km, 4.0 Km, and 10 Km.
• c. From the results in (b), sketch the curve relating v/Vmax to [S]/Km. What is the best range of [S] to use in
determining Km or investigating the dependence of v on [S]?
Q.13](https://image.slidesharecdn.com/enzymetechnologysolvedproblems-200407090146/85/Enzyme-technology-solved-problems-31-320.jpg)
![a. Since both substrate concentrations are well above Km, you
can assume that Vmax = 37 μmol/min. Then
Therefore, at [S] 1.0 x 10-2 M, v still is equal to Vmax.
A.13](https://image.slidesharecdn.com/enzymetechnologysolvedproblems-200407090146/85/Enzyme-technology-solved-problems-32-320.jpg)
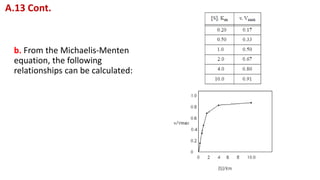
![c. When you plot these values you will be able to see that the best
range of [S] for studying the dependence of v on [S] is in the
neighborhood of Km or below it, since changes in [S] below Km cause
greater changes in v than do changes in [S] above Km. Therefore, when
using graphic methods to determine Km and Vmax, several
measurements should be made at [S] well below Km.
A.13 Cont.](https://image.slidesharecdn.com/enzymetechnologysolvedproblems-200407090146/85/Enzyme-technology-solved-problems-34-320.jpg)

![a.1 'unit' here = 10 μmole/15 min at 37ºC
= 10/15 μmole min-1
= 10/15.60 μmole s-1
When [S] >> Km,
Vo = Vmax
Here Vmax = 2800 units mg-1
= 2800 x 10/15.60
= 31.3 μmol s-1 mg-1.
NOTE. THIS IS BY DEFINITION THE SP. ACT.
A.14](https://image.slidesharecdn.com/enzymetechnologysolvedproblems-200407090146/85/Enzyme-technology-solved-problems-36-320.jpg)


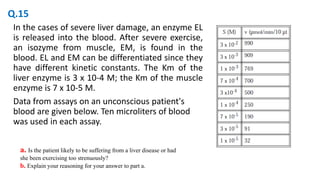
![• a. Liver
• b. The [S]Vmax/2 is close to the Km of the liver enzyme
A.15](https://image.slidesharecdn.com/enzymetechnologysolvedproblems-200407090146/85/Enzyme-technology-solved-problems-40-320.jpg)

![A. The amount of enzyme which converts 1 μmole of substrate to
product per min (at 25/37oC)
B. Under steady state conditions, S is converted into P at a constant
rate, the [S] and [P] vs time plots are linear, and [ES] is Constant
C. An enzyme with a quaternary structure, i.e., made up of more than
one subunit (protomer, monomer). The subunits may be the same
(e.g., a homodimer), or different (e.g., a heterodimer).
A.16](https://image.slidesharecdn.com/enzymetechnologysolvedproblems-200407090146/85/Enzyme-technology-solved-problems-42-320.jpg)

![• Saturating substrate ([S] >> Km)
A.17](https://image.slidesharecdn.com/enzymetechnologysolvedproblems-200407090146/85/Enzyme-technology-solved-problems-44-320.jpg)
![• Indicate the effects of substrate concentration, enzyme
concentration, temperature, inhibitors or activators on enzyme
activity by labeling correctly both axes of the graphs given shown
below
Your choice for axes are: Energy, [E], Temperature, [S], 1/[S], 1/v, v.
(You may use the same label on more than one graph.)
Q.18](https://image.slidesharecdn.com/enzymetechnologysolvedproblems-200407090146/85/Enzyme-technology-solved-problems-45-320.jpg)

![An enzyme-catalyzed reaction was assayed at several substrate
concentrations. Two data points which fell on the Lineweaver- Burk plot
are v = 41.7 μmol S/min when [S] = 5 x 10-4 M and v = 16.7 μmol S/min
when [S] = 5 x 10-6 M. Place the two points on a line on the
accompanying graph.
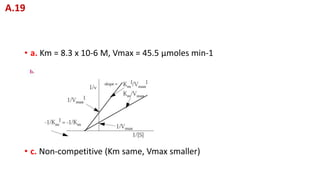
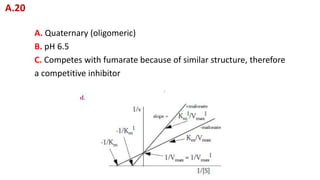
A. Determine the value of Km and Vmax in the correct units. When an
inhibitor was added, the velocities fell to 1/2 their uninhibited values.
B. Plot the inhibited line on the graph.
C. Is the inhibitor competitive or non-competitive? On what evidence
did you base your decision?
Q.19](https://image.slidesharecdn.com/enzymetechnologysolvedproblems-200407090146/85/Enzyme-technology-solved-problems-47-320.jpg)






![A. [S] >> Km
Therefore, vo = Vmax Therefore, the velocity is doubled when [E]tot is
doubled
B. None, because the enzyme is already saturated with substrate
A.23](https://image.slidesharecdn.com/enzymetechnologysolvedproblems-200407090146/85/Enzyme-technology-solved-problems-56-320.jpg)
![Answer the following with true or false; justify your answer in each
case.
A. The initial rate of an enzyme-catalyzed reaction is independent of
substrate concentration.
B. If enough substrate is added, the normal Vmax of an enzyme
catalyzed reaction can be attained even in the presence of a
noncompetitive inhibitor.
C. The rate of an enzyme-catalyzed reaction in the presence of a rate-
limiting concentration of substrate decreases with time.
D. The sigmoid shape of the v-versus-[S] curve for some regulatory
enzymes indicates that the affinity of the enzyme for substrate
decreases as [S] is increased
Q.24](https://image.slidesharecdn.com/enzymetechnologysolvedproblems-200407090146/85/Enzyme-technology-solved-problems-57-320.jpg)







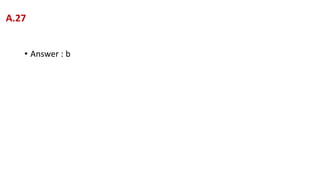




![a. The standard conditions of assay for E1 must be met (pH, ionic strength, cofactors,
temperature, etc), so the results may be compared to the range of normal values.
b. [A] must be at a sufficiently high concentration to saturate E1. [NAD+] must be at a
sufficiently high concentration to saturate E2. E2 must be present at sufficiently high levels
of activity for it not to be a limiting factor in the assay
c. The rate of the second reaction must be much greater than that of the first reaction (E2
at high activity), so that only the amount of E1 limits the rate of the reaction, i.e., Vmax2
must be >> Vmax1.
d. Increase the activity of E2 present in the assay medium, so that it is much greater than
that expected for E1.
A.31](https://image.slidesharecdn.com/enzymetechnologysolvedproblems-200407090146/85/Enzyme-technology-solved-problems-73-320.jpg)



![When the enzyme is saturated with substrate,
v0 = Vmax = kcat Et = a constant for that assay,
i.e., the rate = a constant = k, therefore it is not dependent on [S], and
zero order kinetics in [S} are observed.
A.33](https://image.slidesharecdn.com/enzymetechnologysolvedproblems-200407090146/85/Enzyme-technology-solved-problems-77-320.jpg)



![d. Vmax = k [Eo] (i.e., Vmax = kcat [E]tot)
A.35](https://image.slidesharecdn.com/enzymetechnologysolvedproblems-200407090146/85/Enzyme-technology-solved-problems-81-320.jpg)
