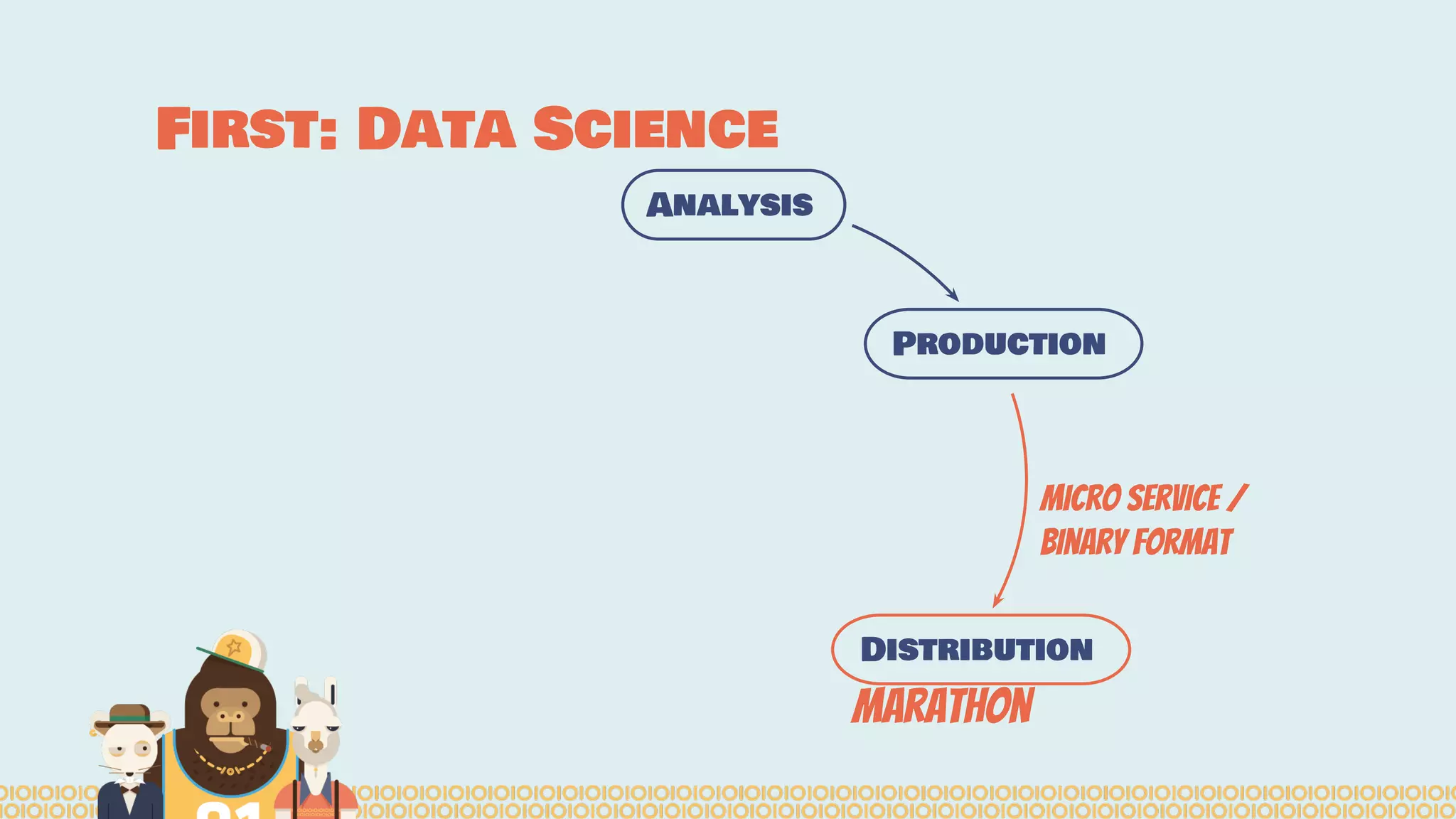
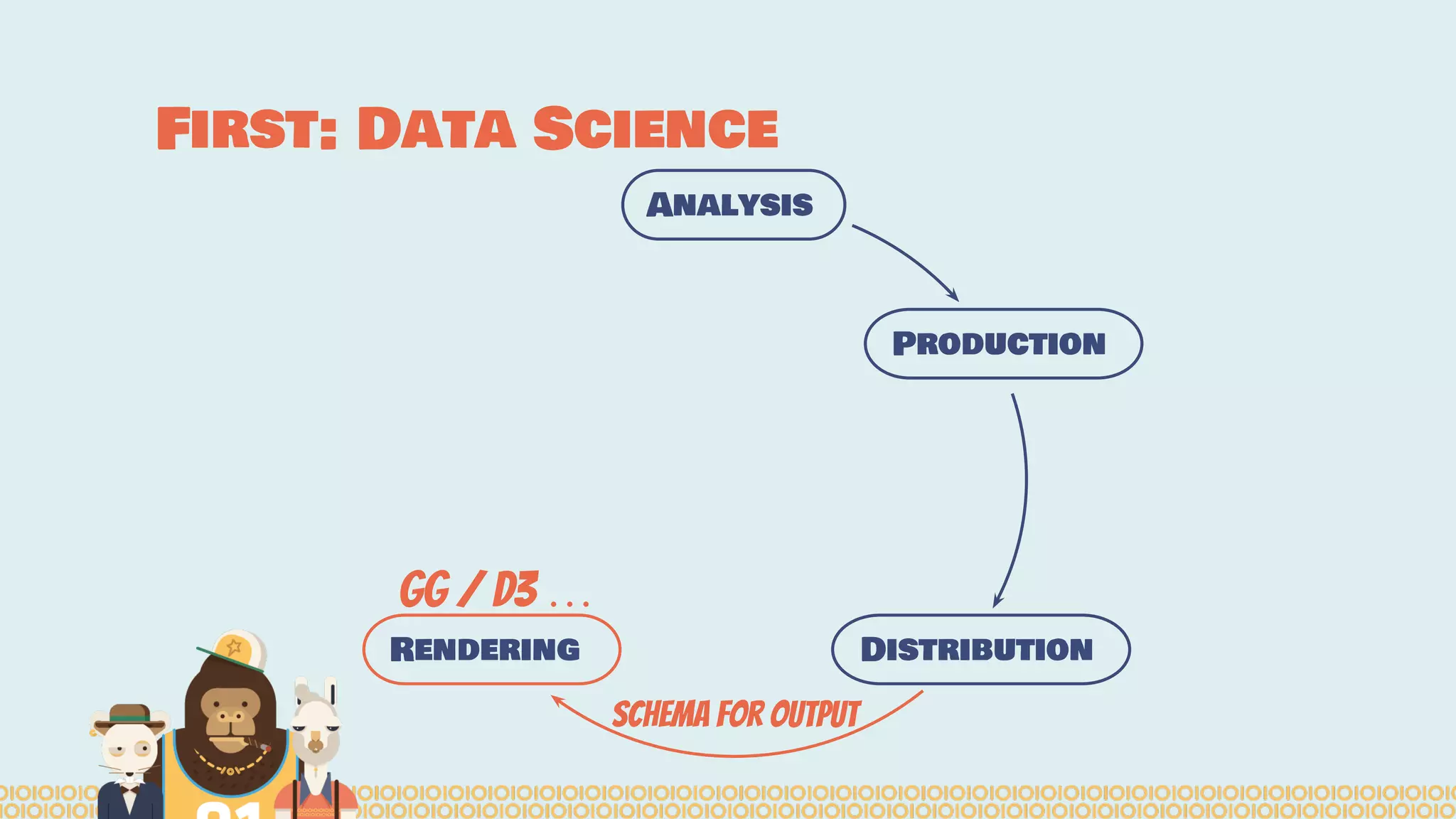
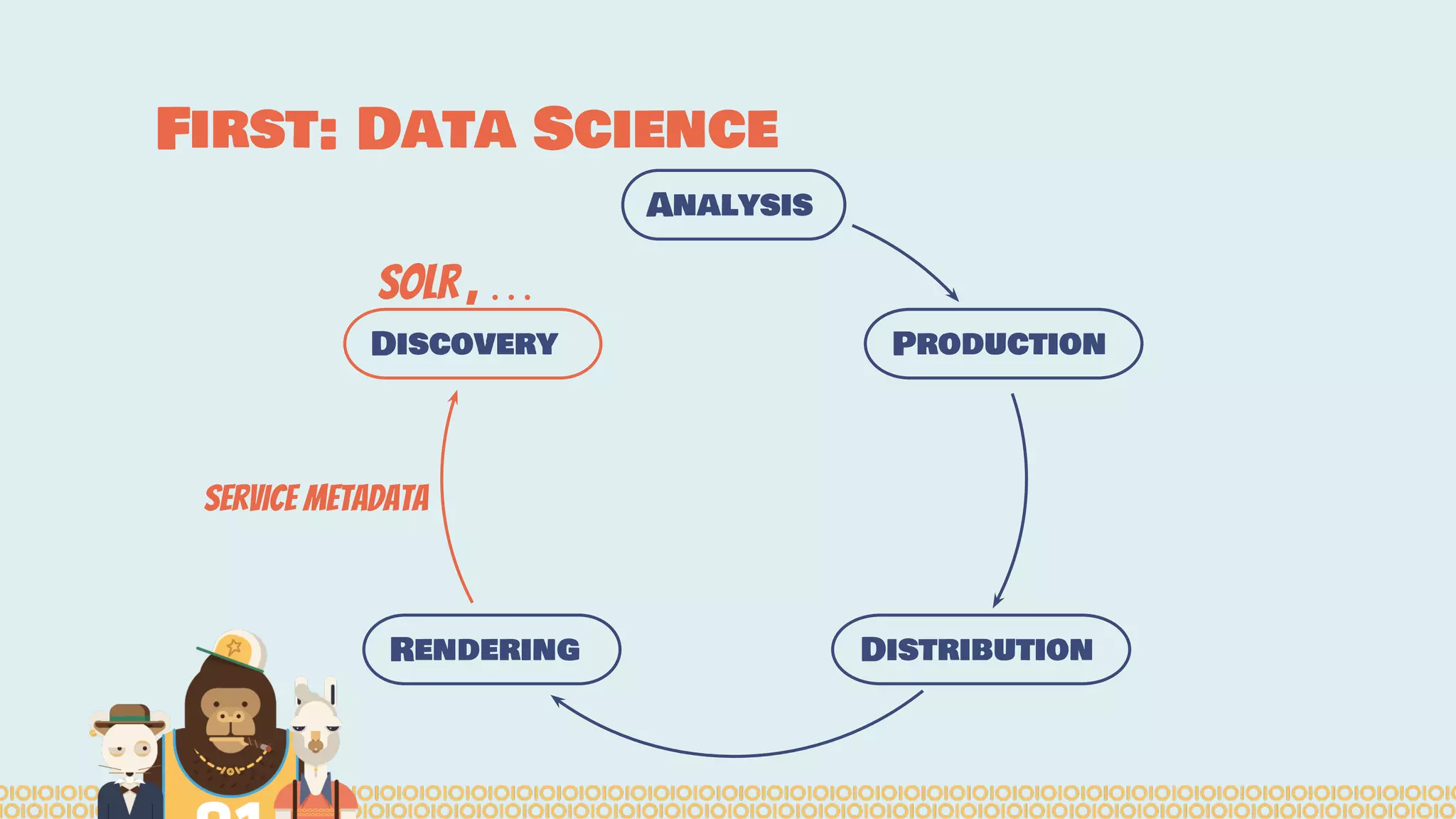
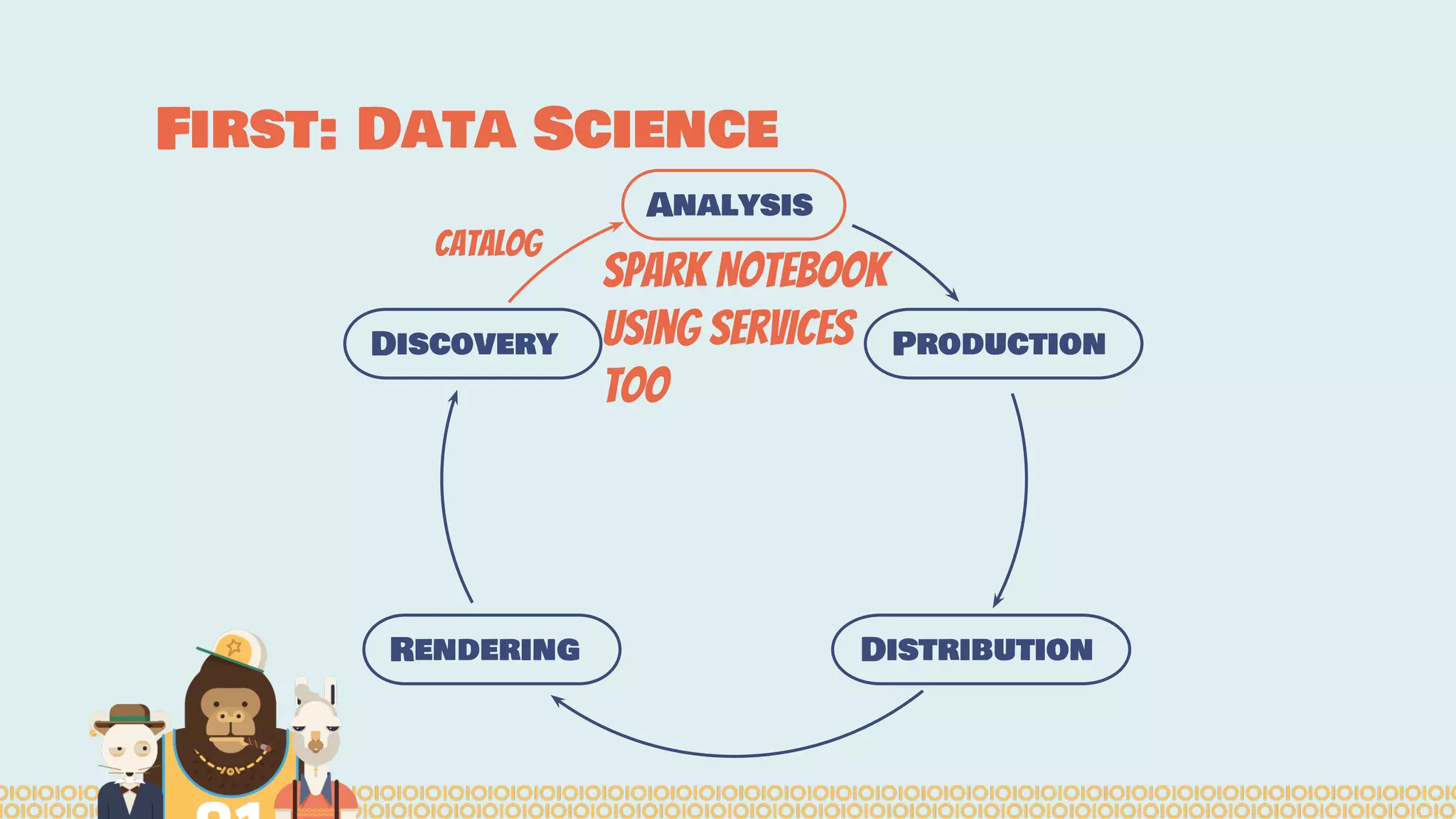
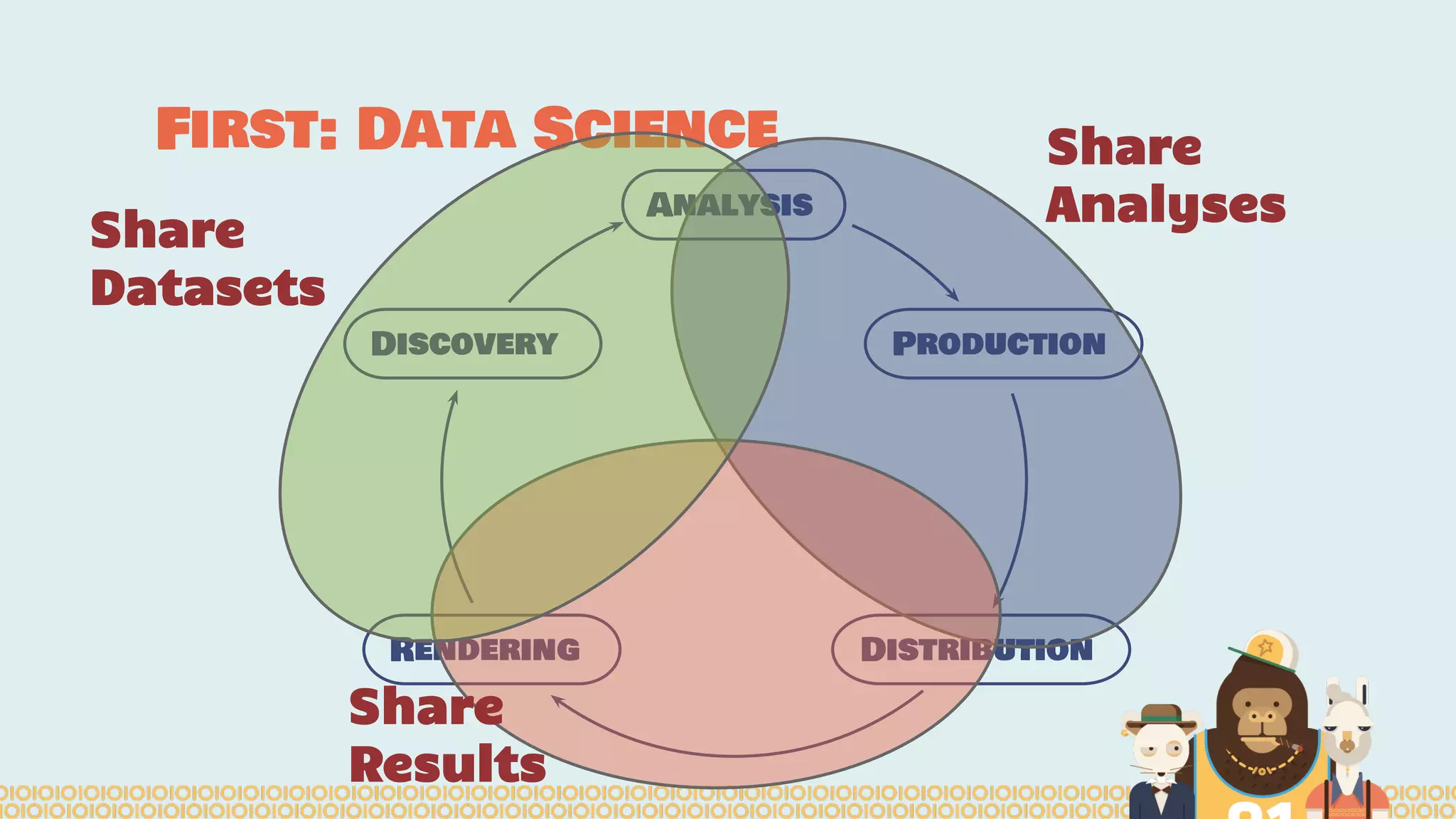
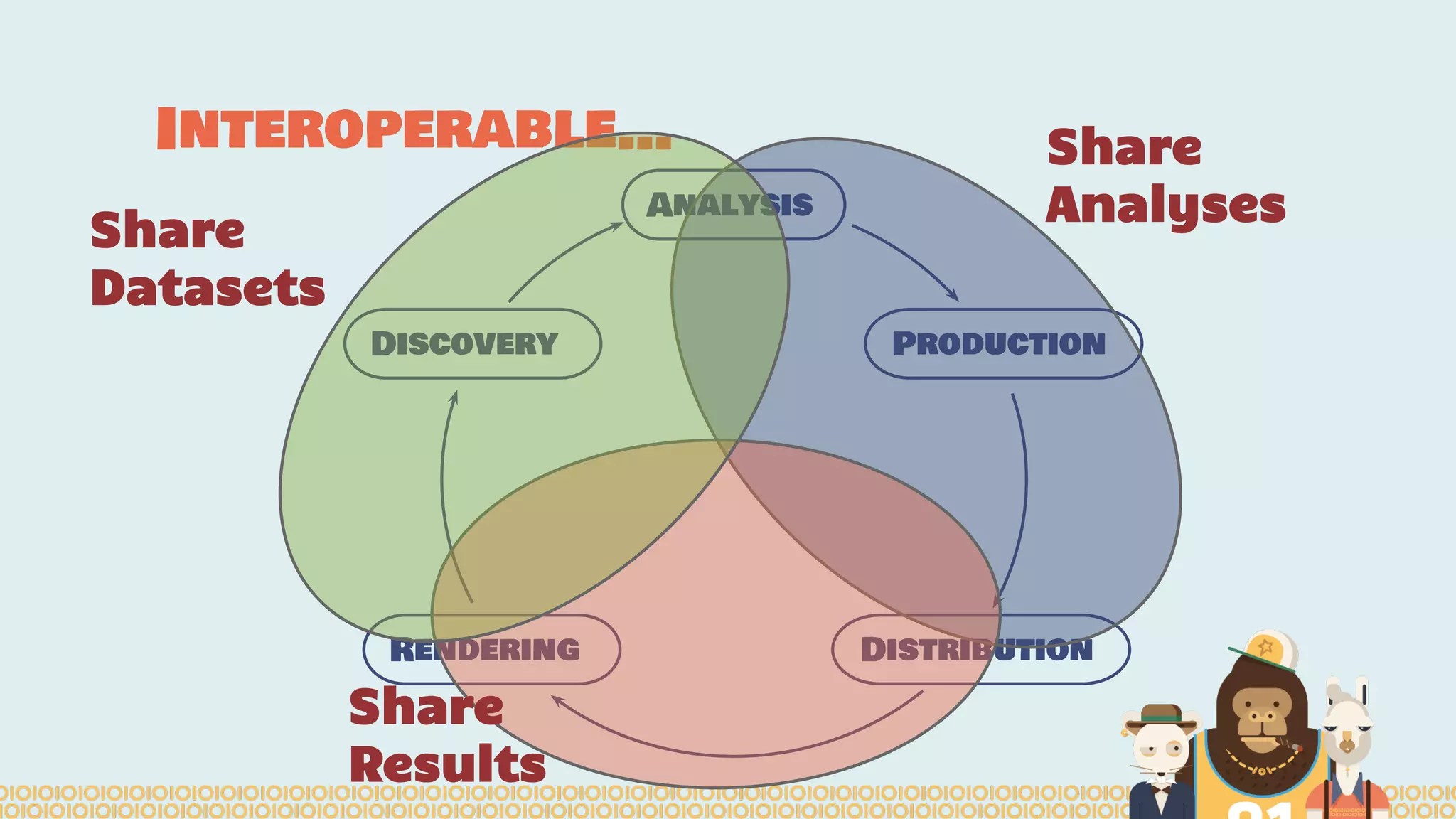
The document outlines the efforts of the Belgian startup Data Fellas in providing scalable and interoperable data services for genomics, focusing on the Big Data challenges represented by genomic data. It highlights various projects and training initiatives related to Apache Spark and distributed machine learning, emphasizing the importance of metadata and service discovery in genomic research. Future plans include expanding their services to incorporate medical data trading and geospatial modules.