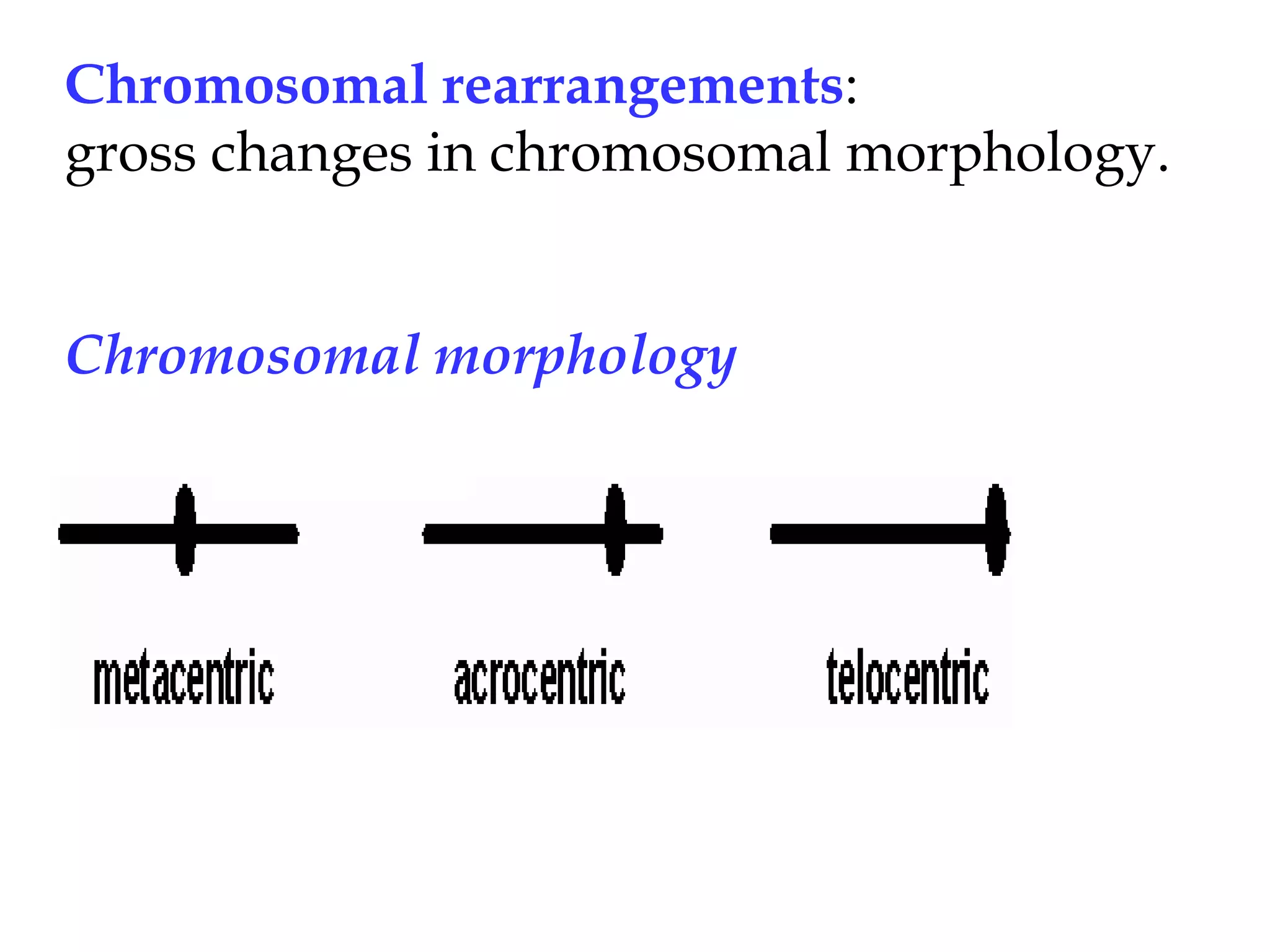
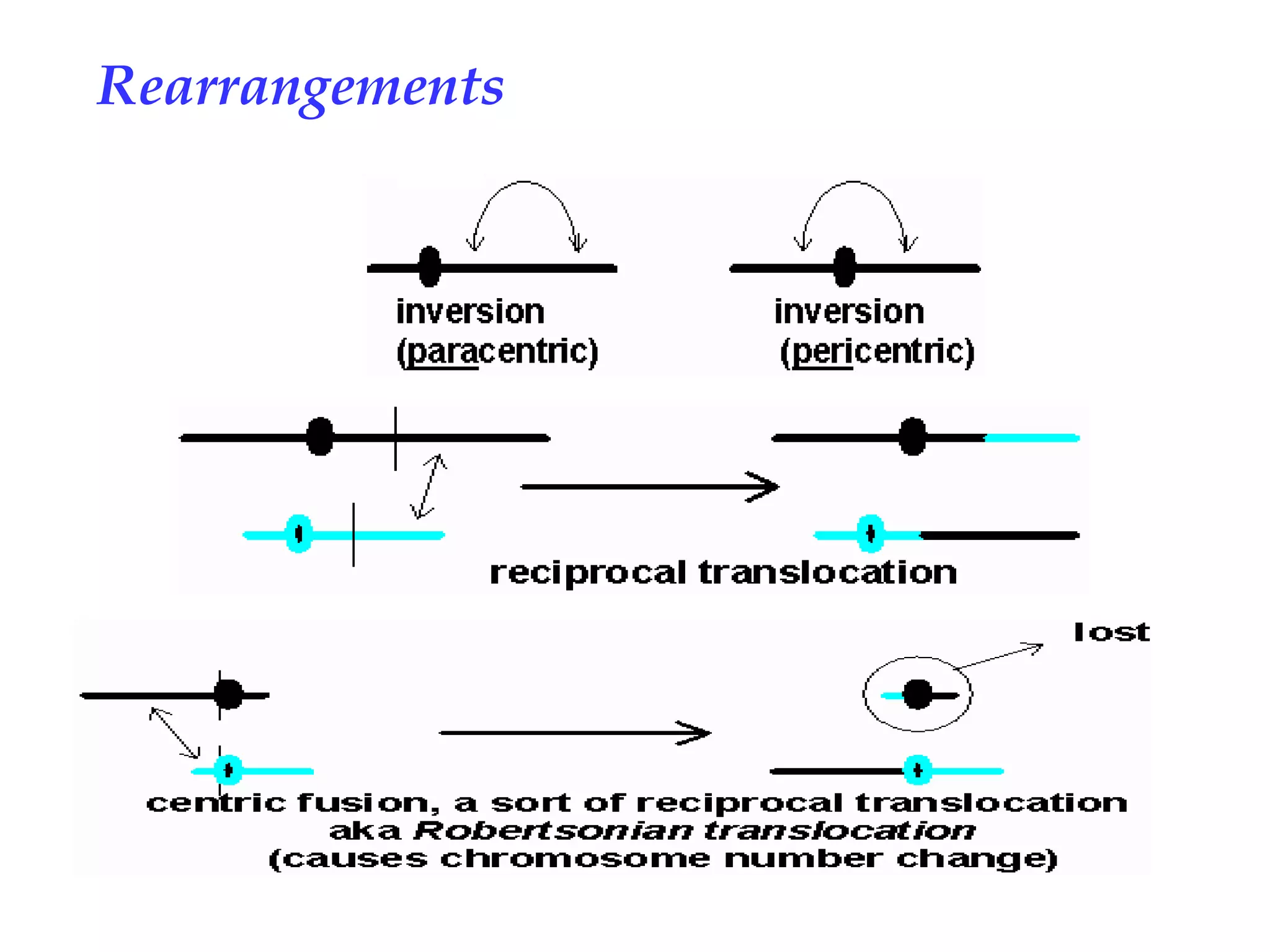
Chromosomal rearrangements are gross changes in chromosomal morphology that can occur via radiation, mutagens, or repetitive sequences in DNA involved in non-homologous recombination. Rearrangements lead to chromosome breakage and "sticky ends" that are normally capped by telomeres to prevent mutation, but uncapped ends can cause duplications or deletions during meiosis. Heterozygous rearrangements often produce unbalanced gametes, giving rearrangements a heterozygous disadvantage that leads to their fixation in populations, and can cause reproductive isolation between populations with different rearrangements. Comparison of chromosome banding patterns has shown that rearrangements have occurred during primate evolution and may have contributed to speciation by trapping groups