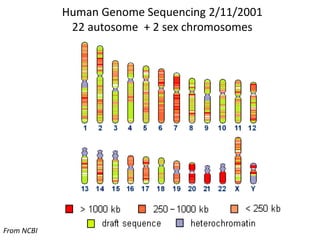
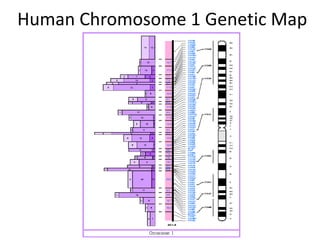
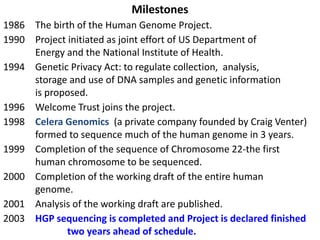
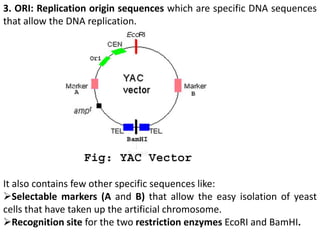
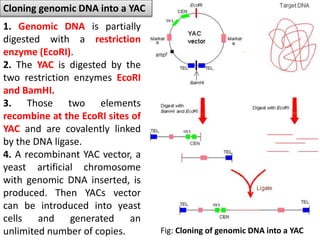
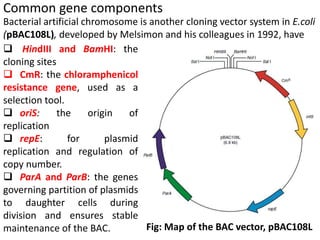
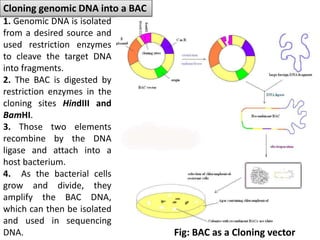
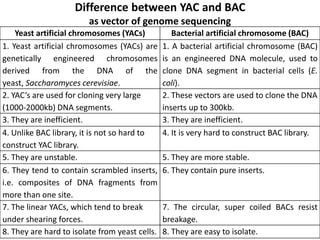
The document discusses the human genome project, which aimed to sequence the entire human genome and identify all human genes. It provides background on the human genome, describing its size, number of genes, and chromosomes. It details the goals and milestones of the human genome project from 1986 to 2003. Vectors like yeast artificial chromosomes and bacterial artificial chromosomes were used to clone large fragments of DNA for sequencing.