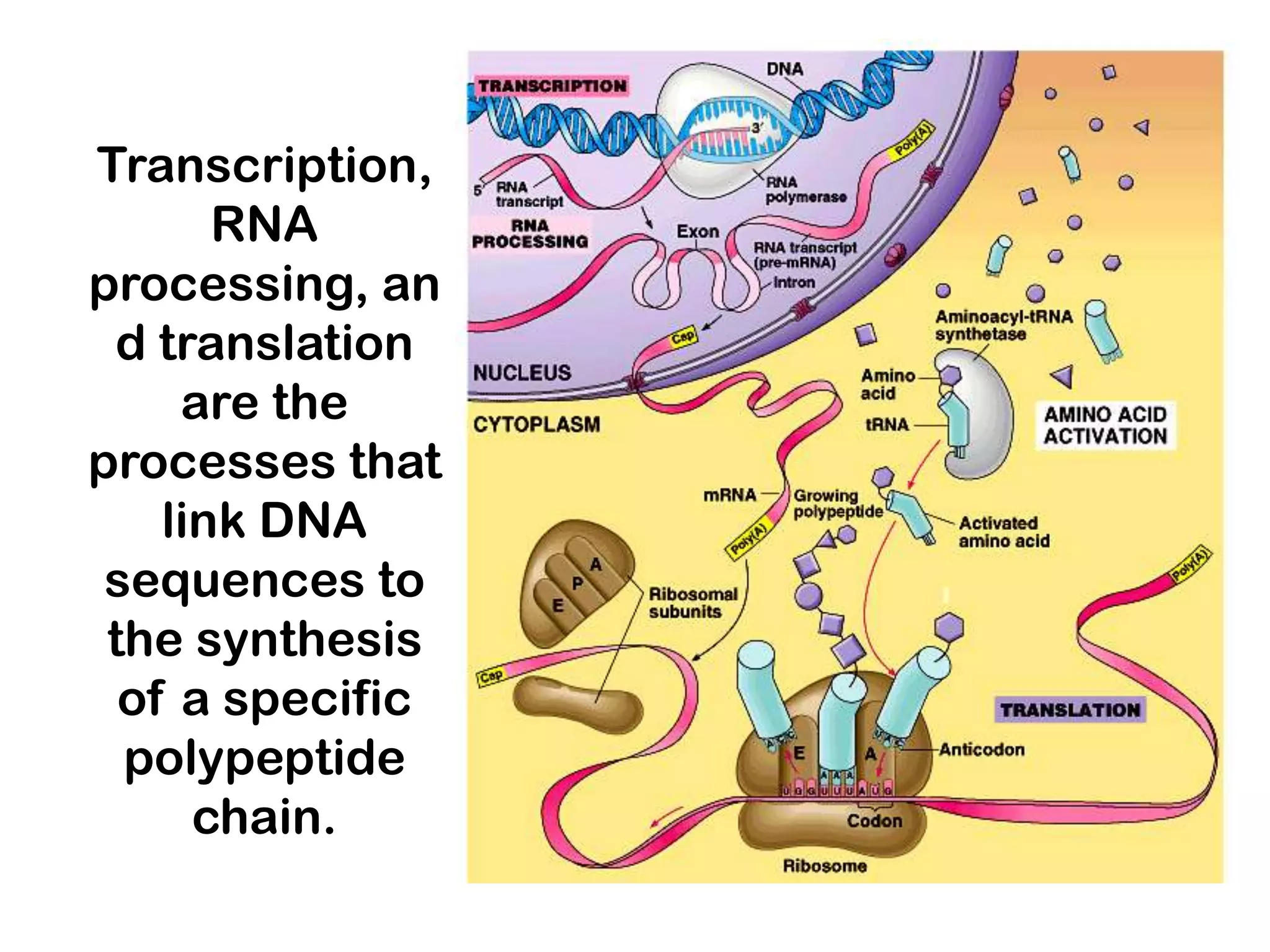
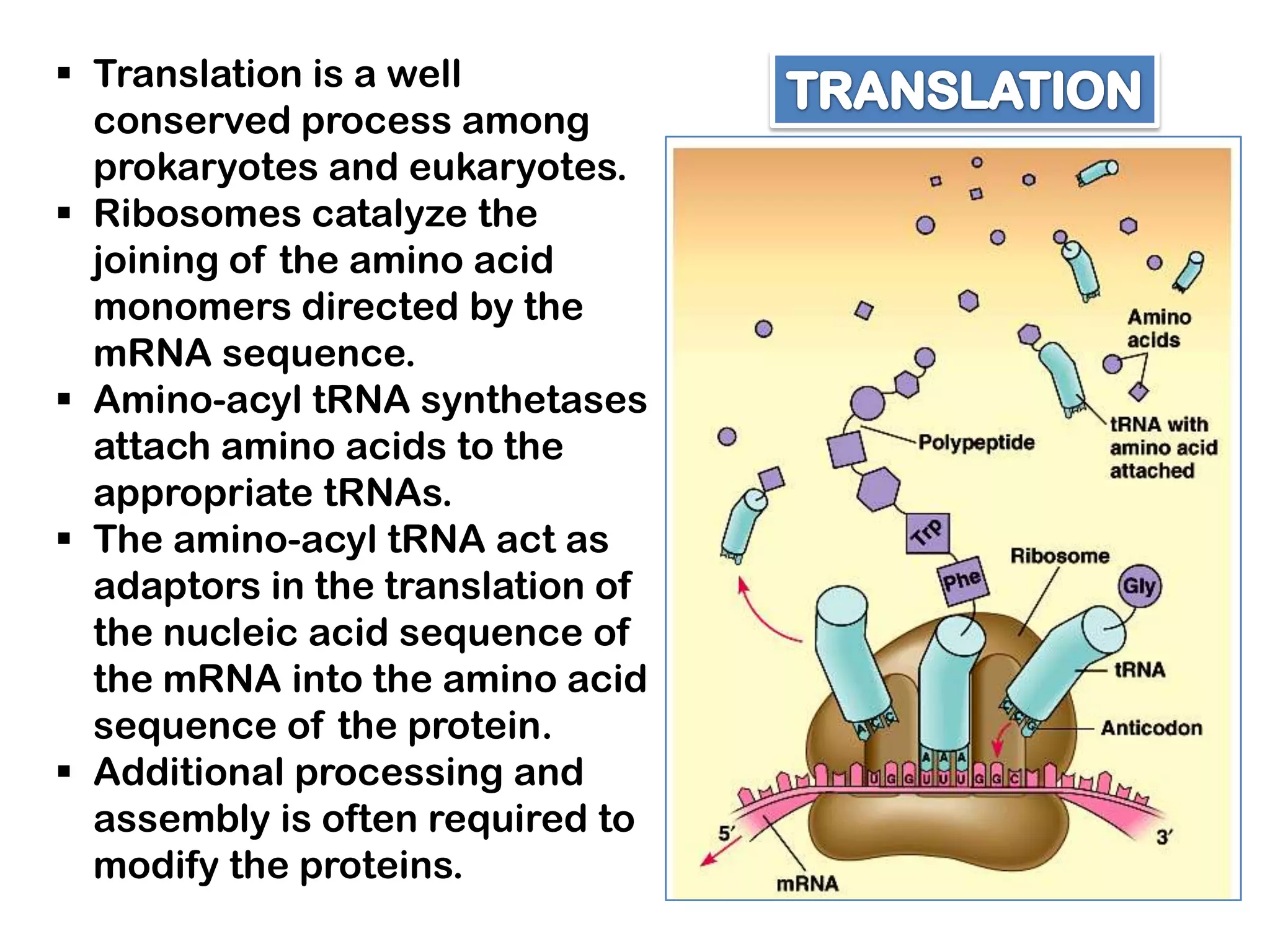
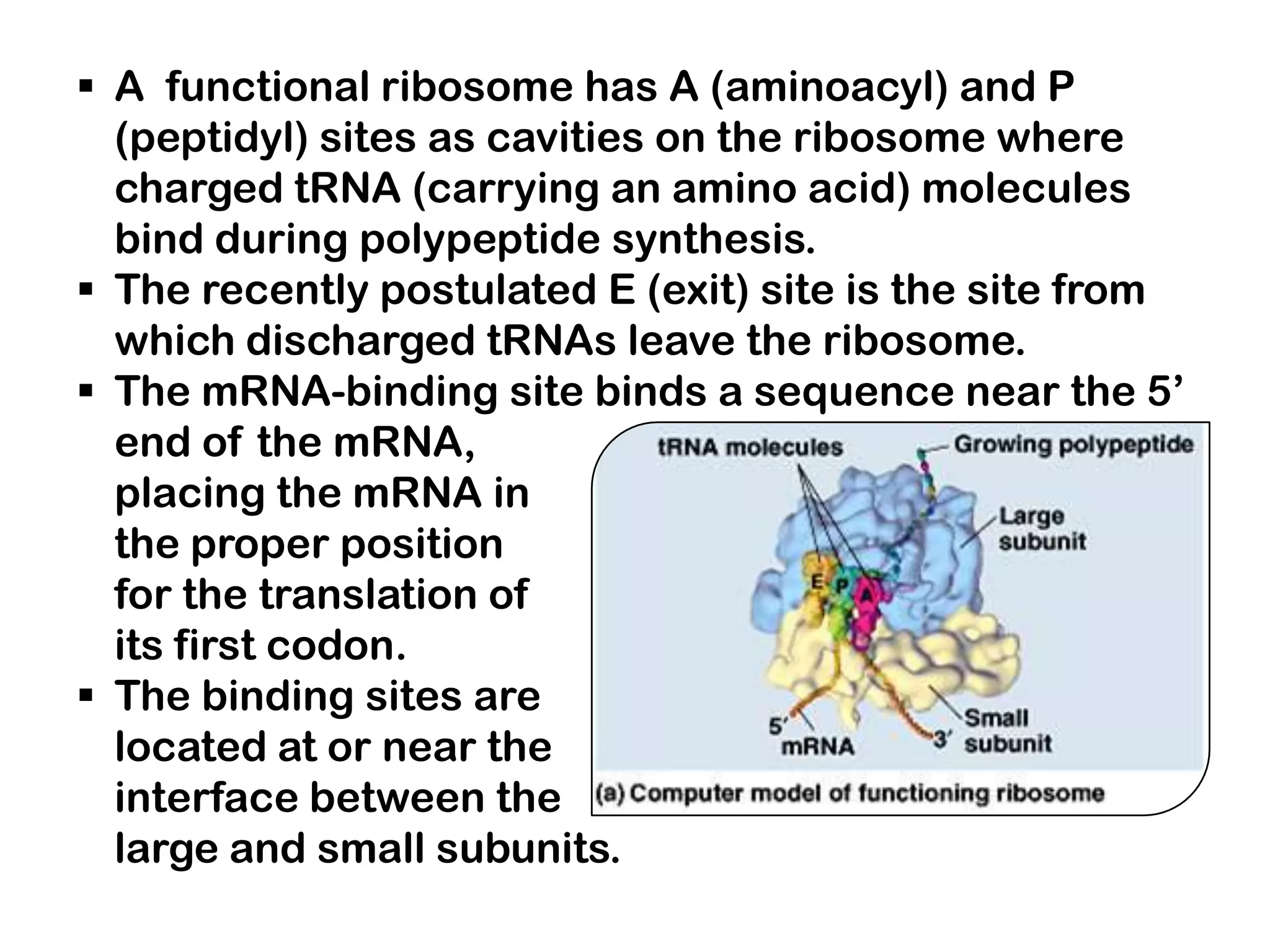
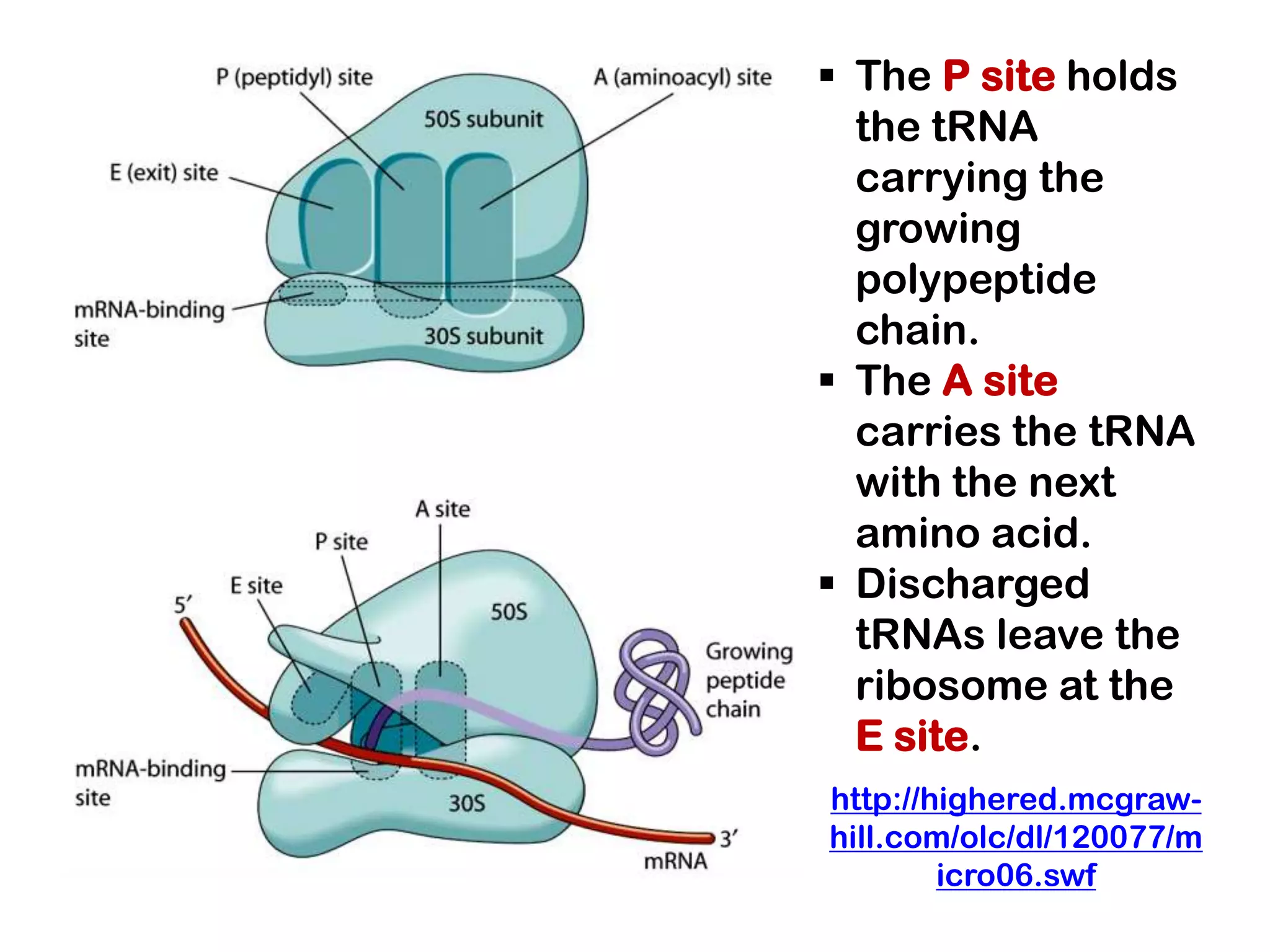
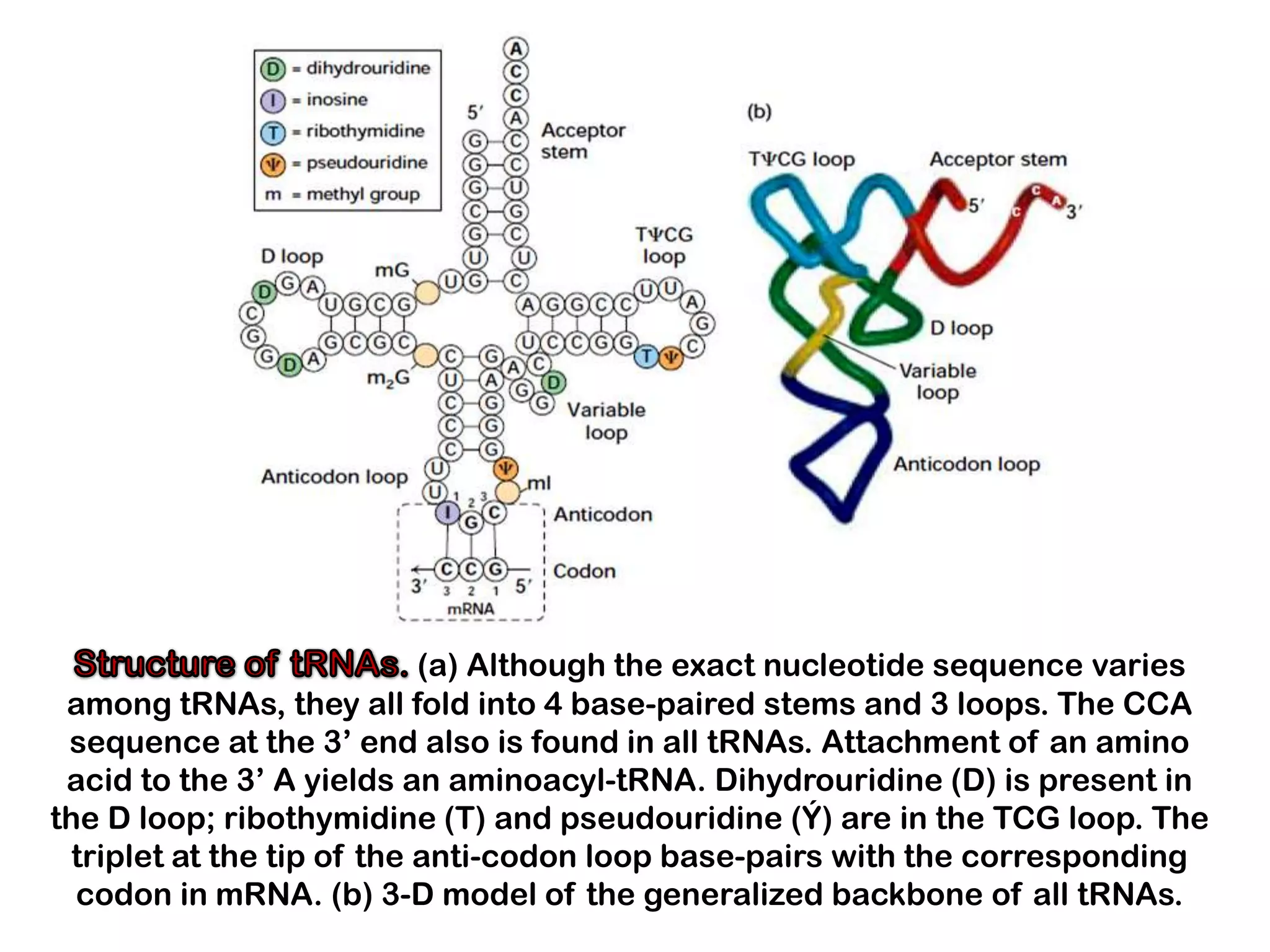
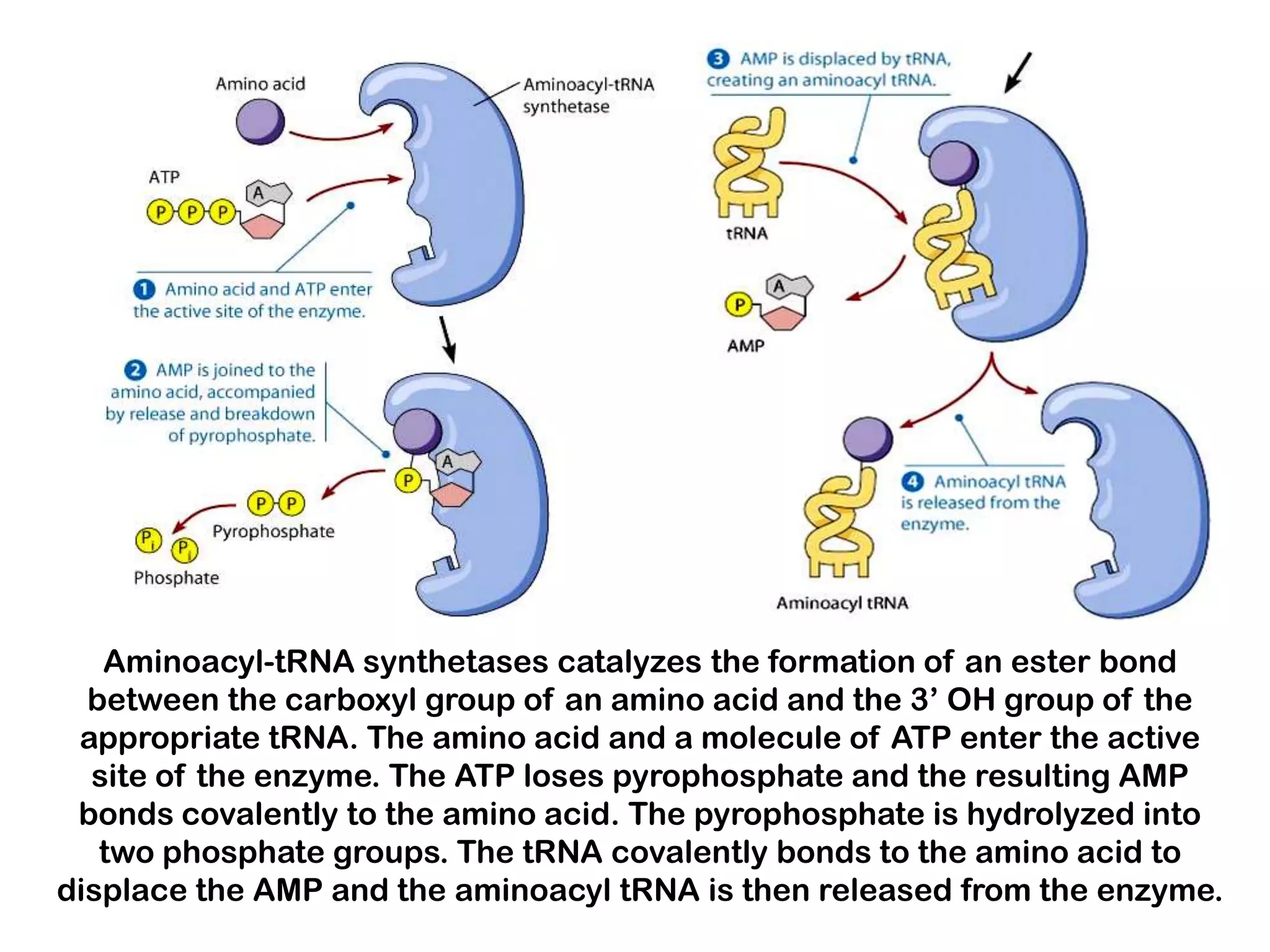
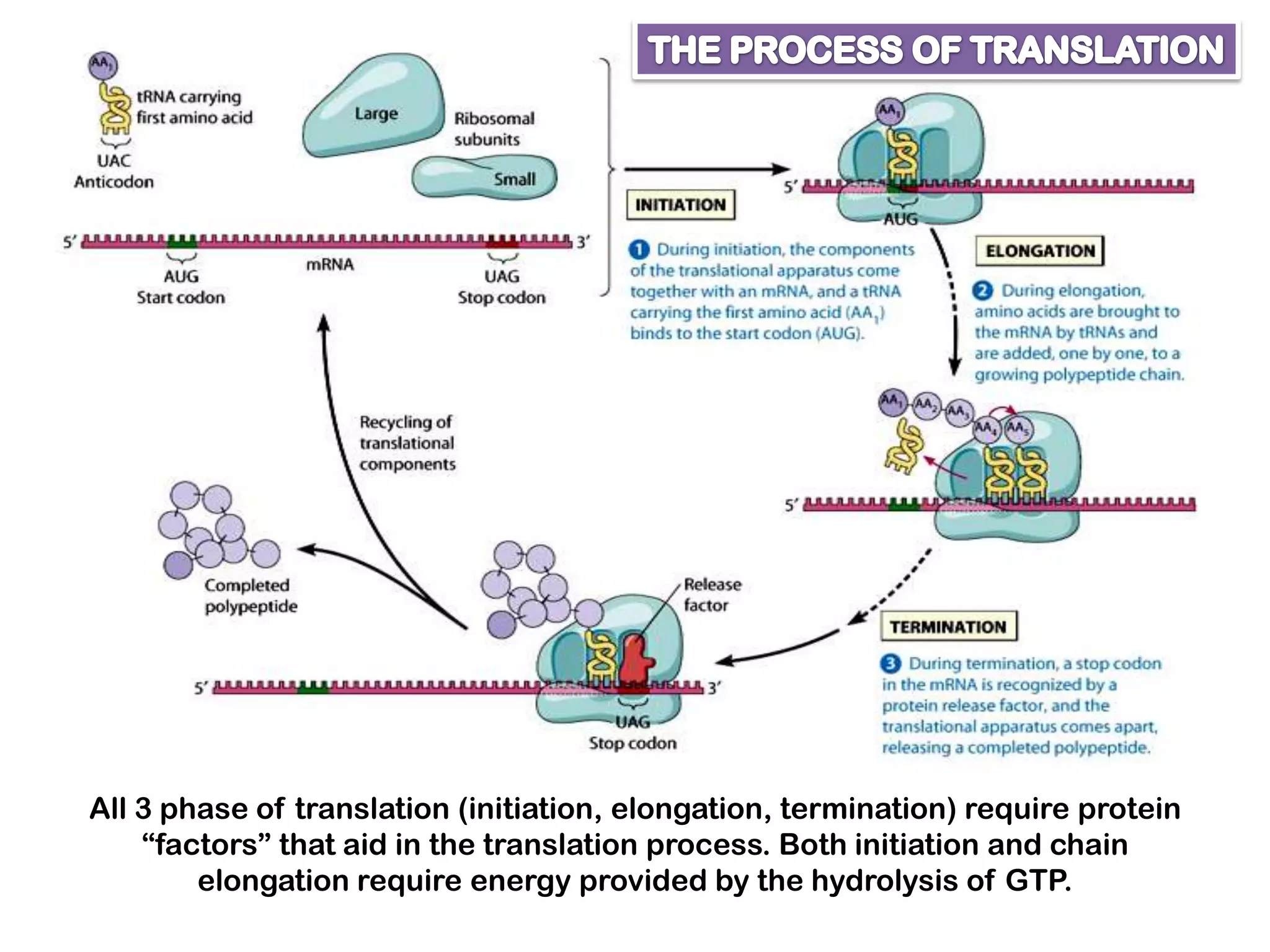
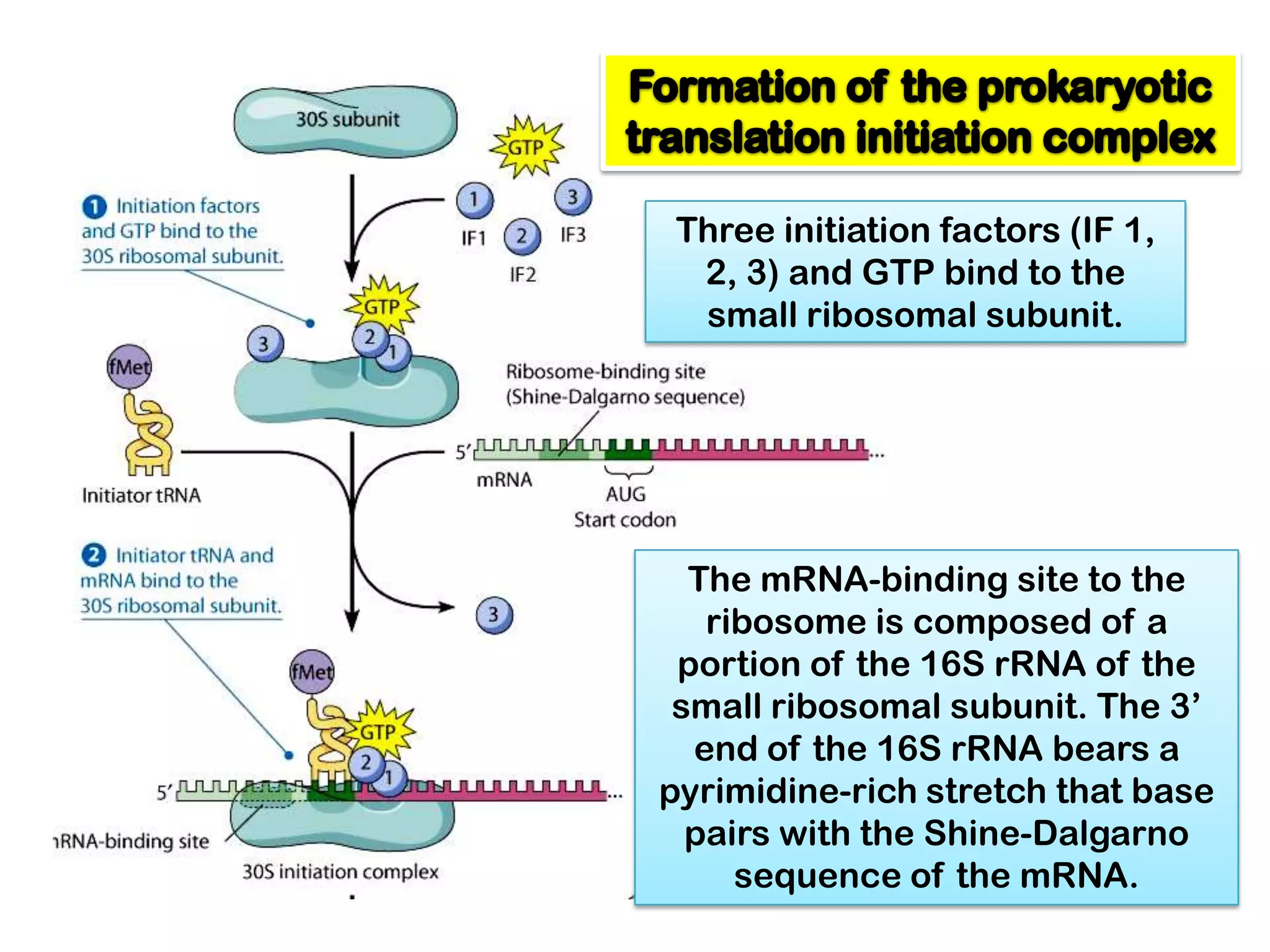
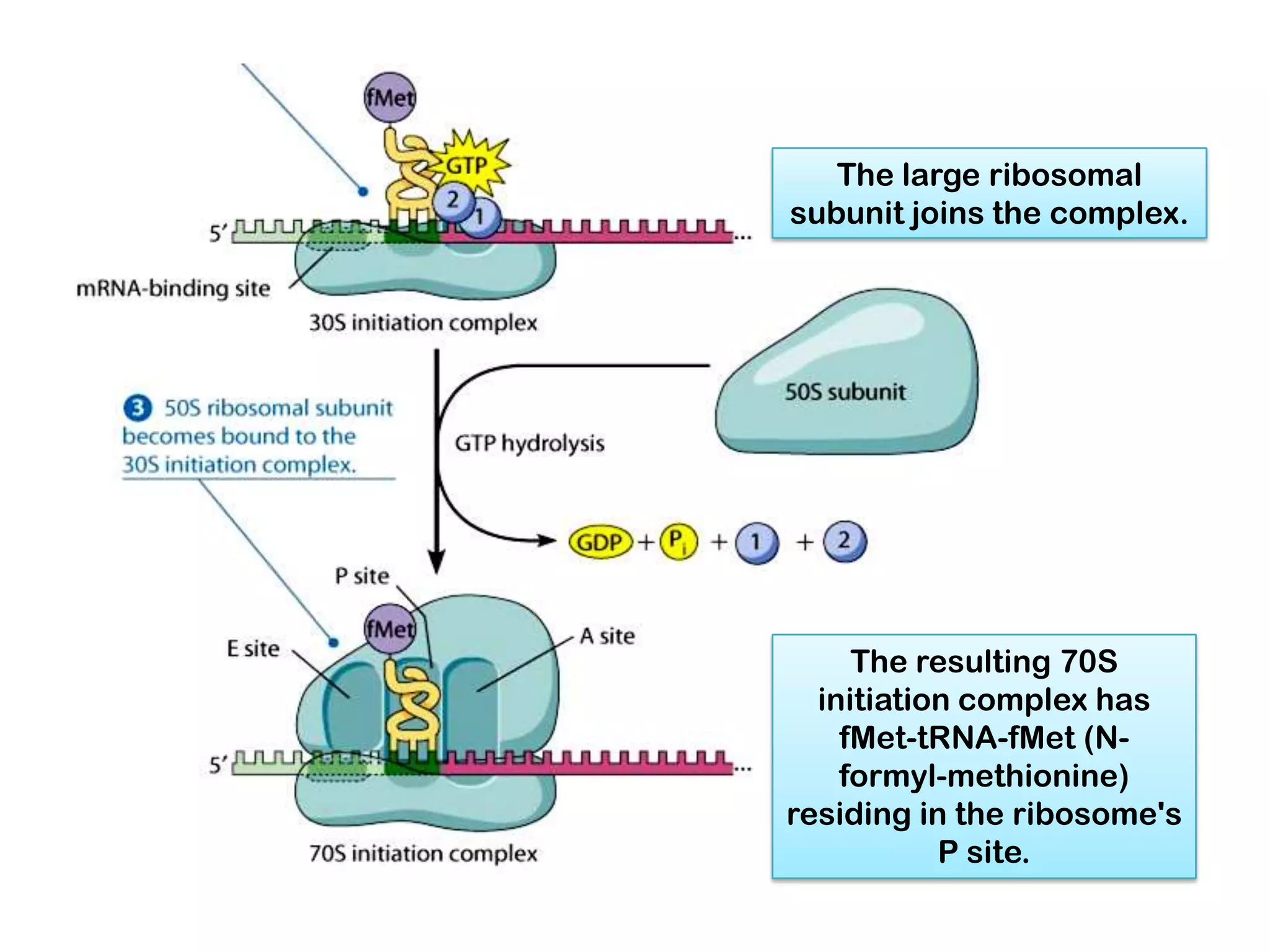
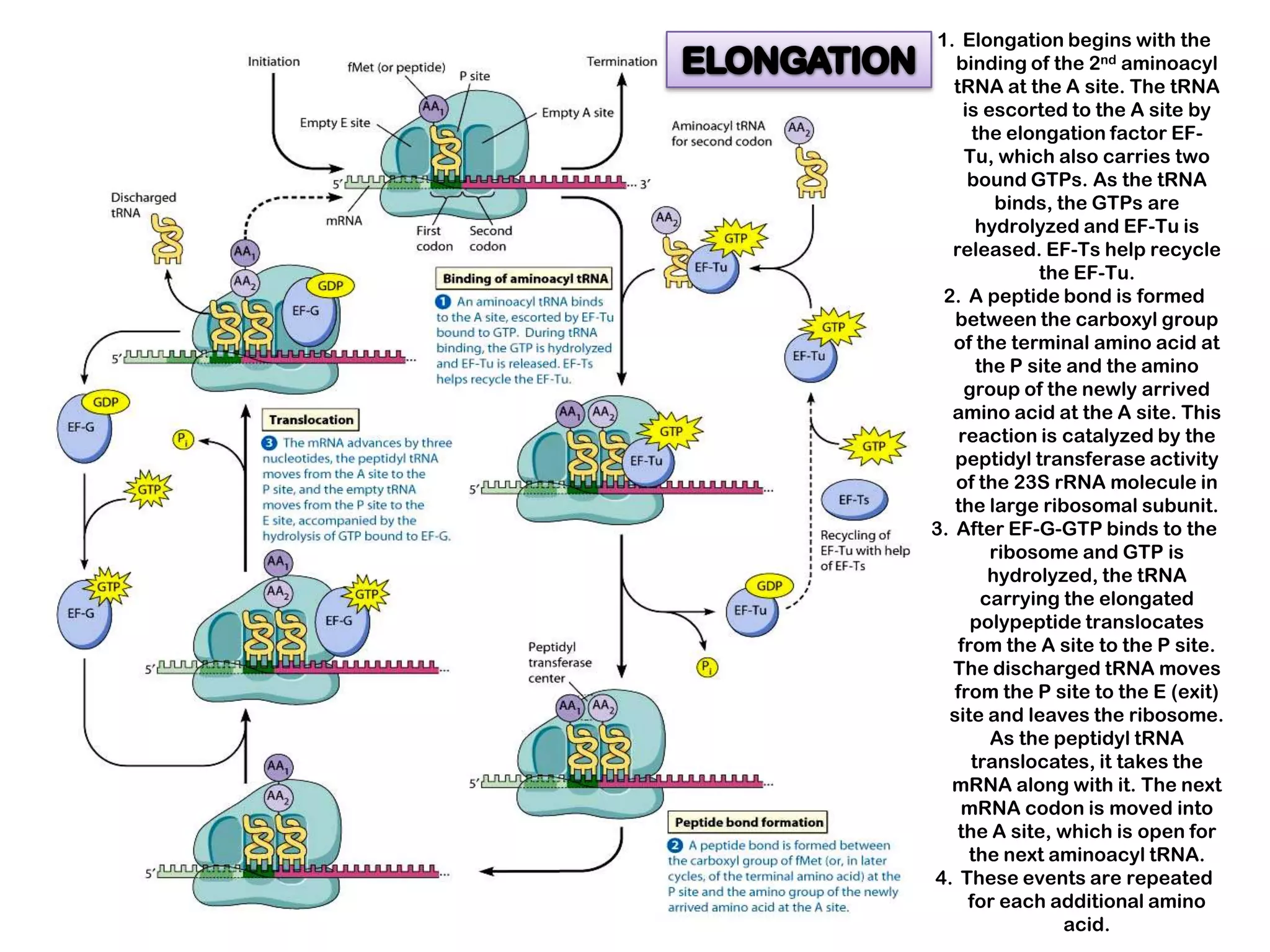
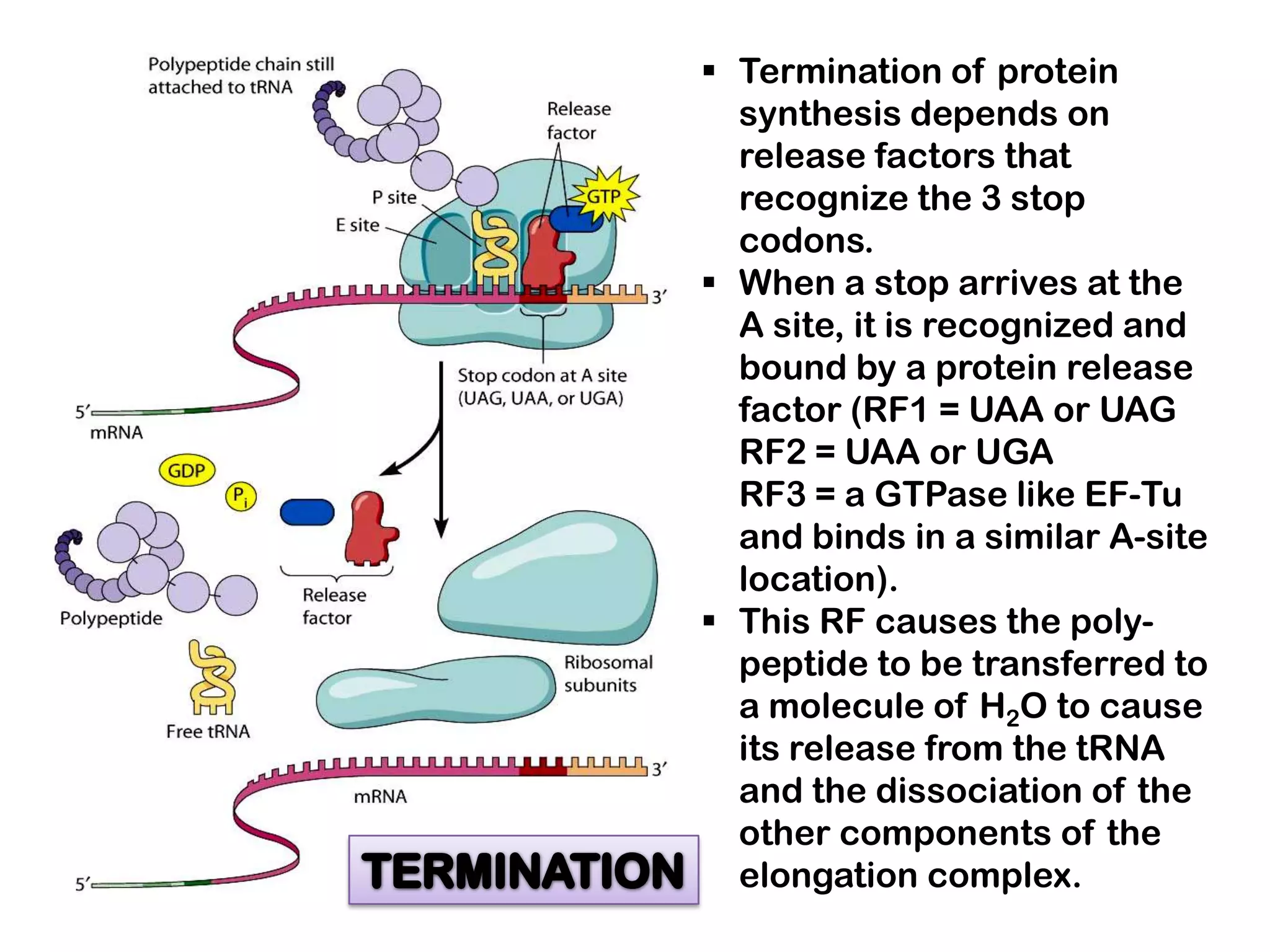
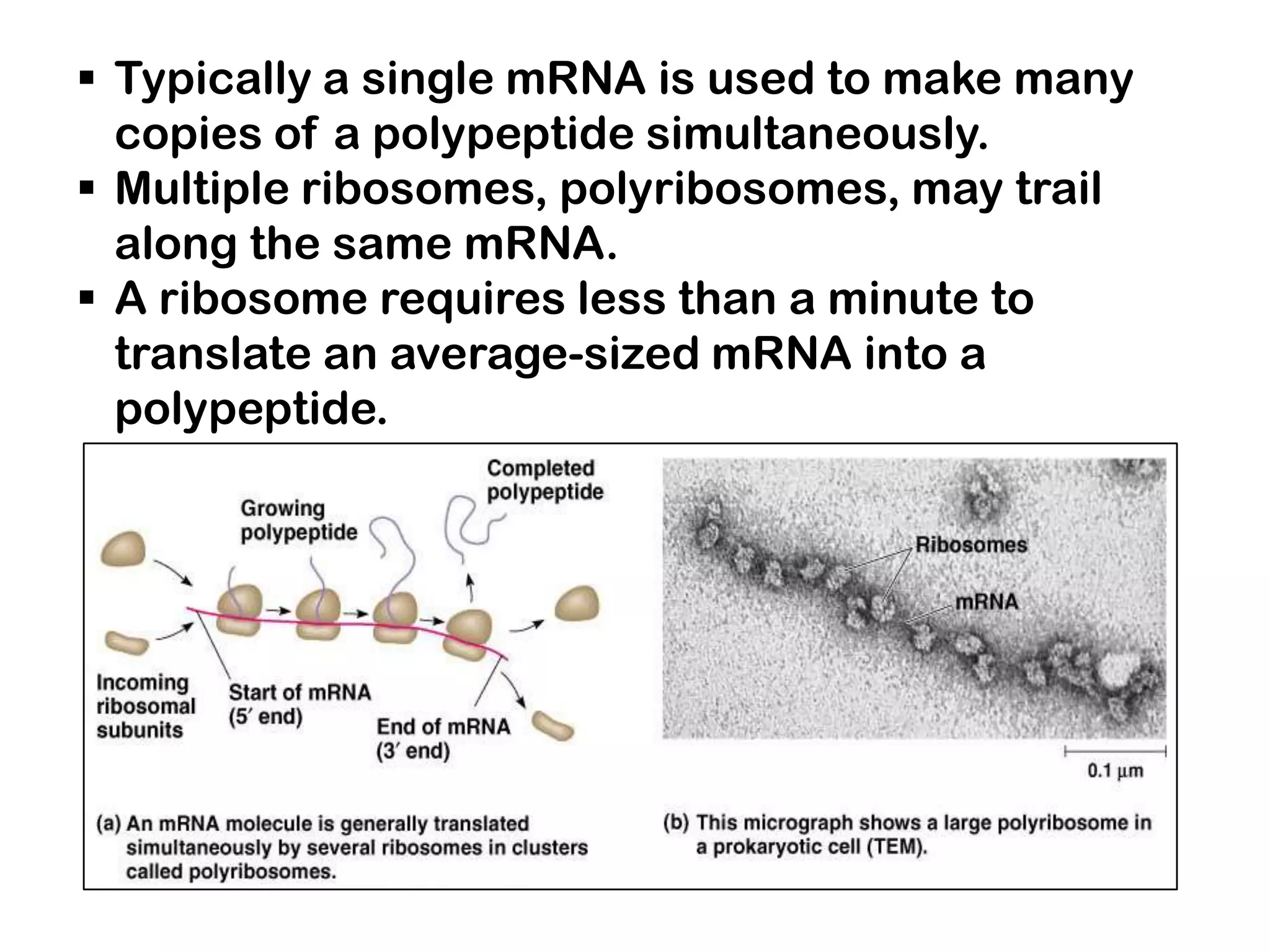
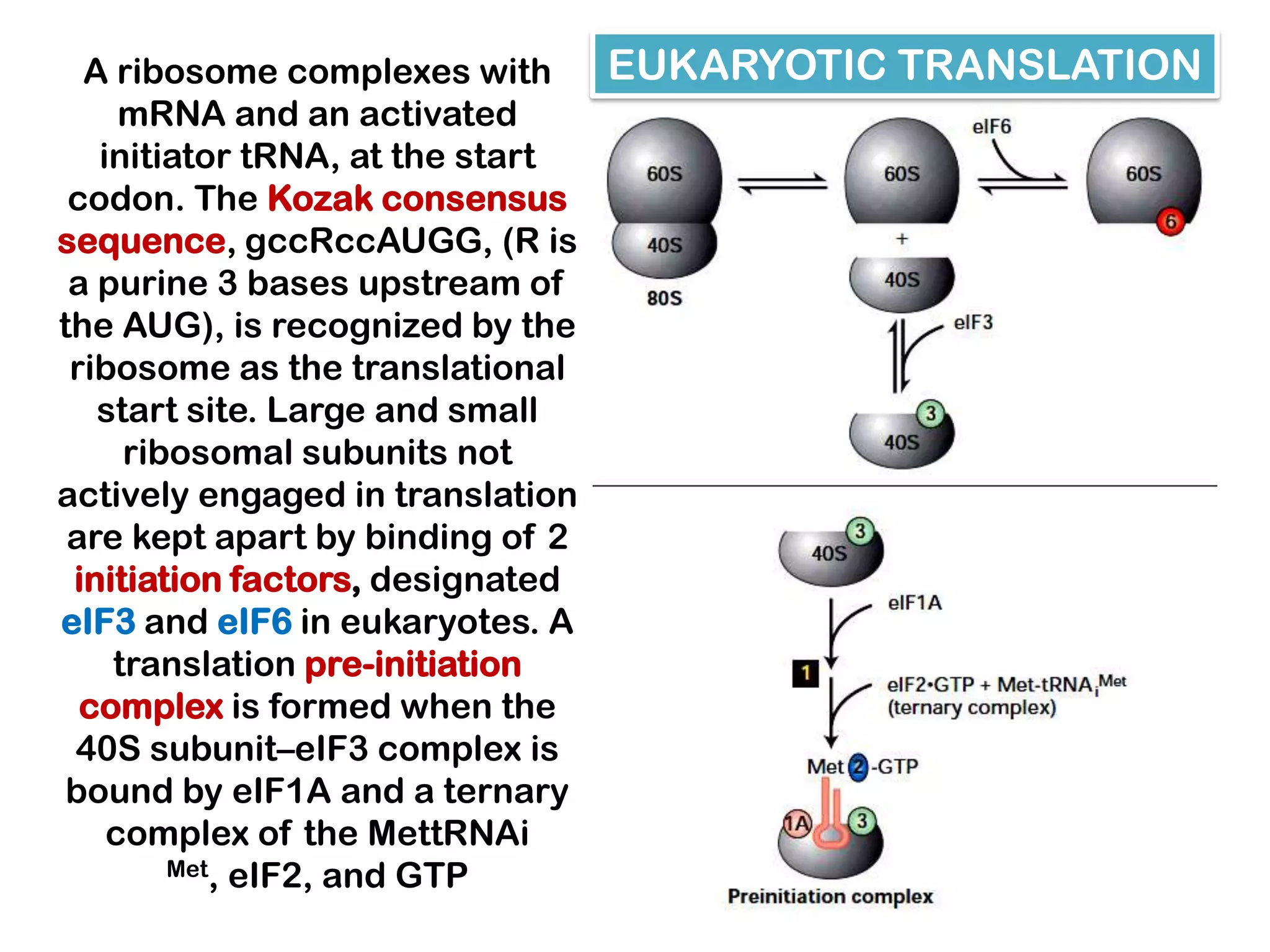
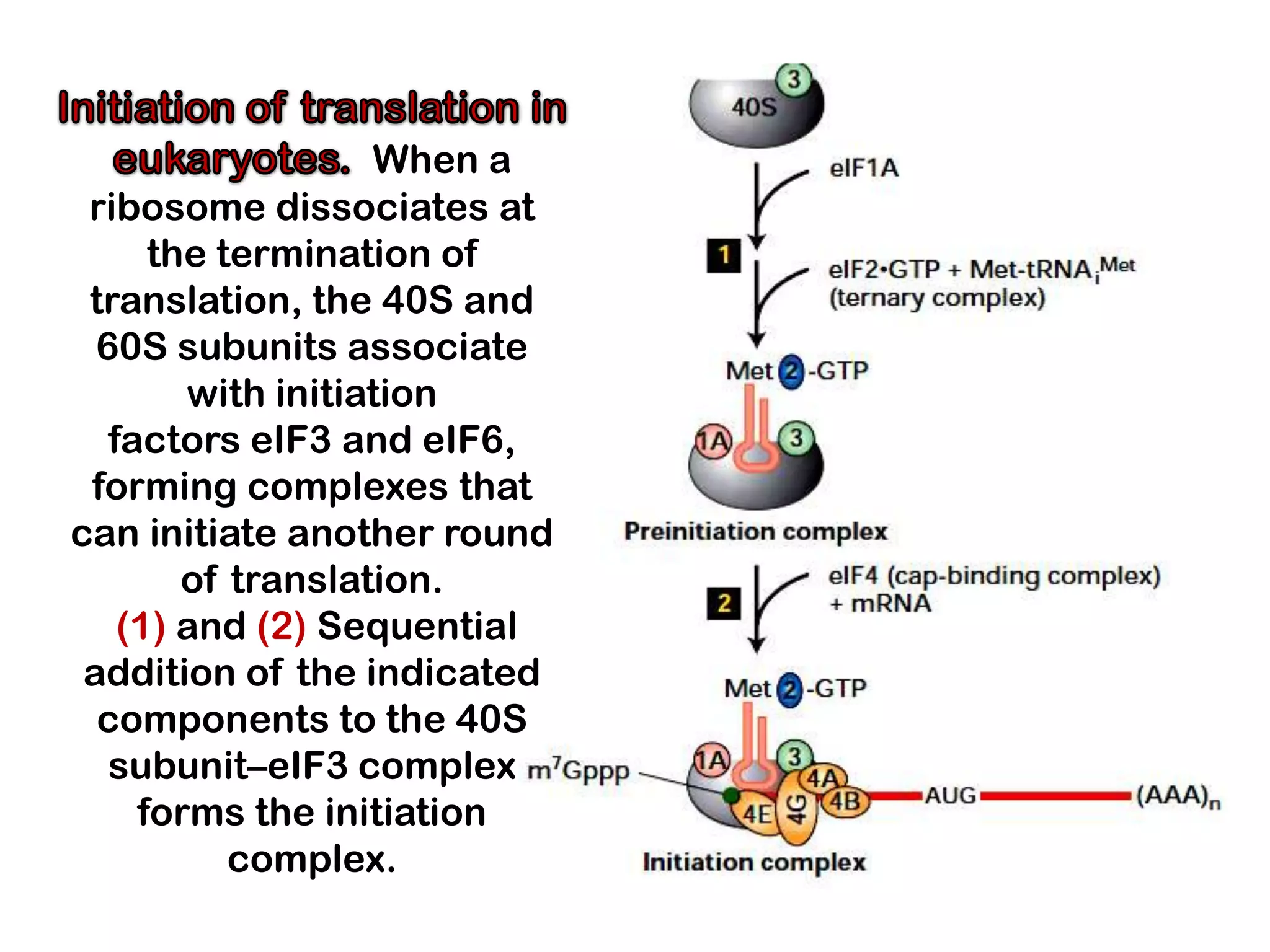
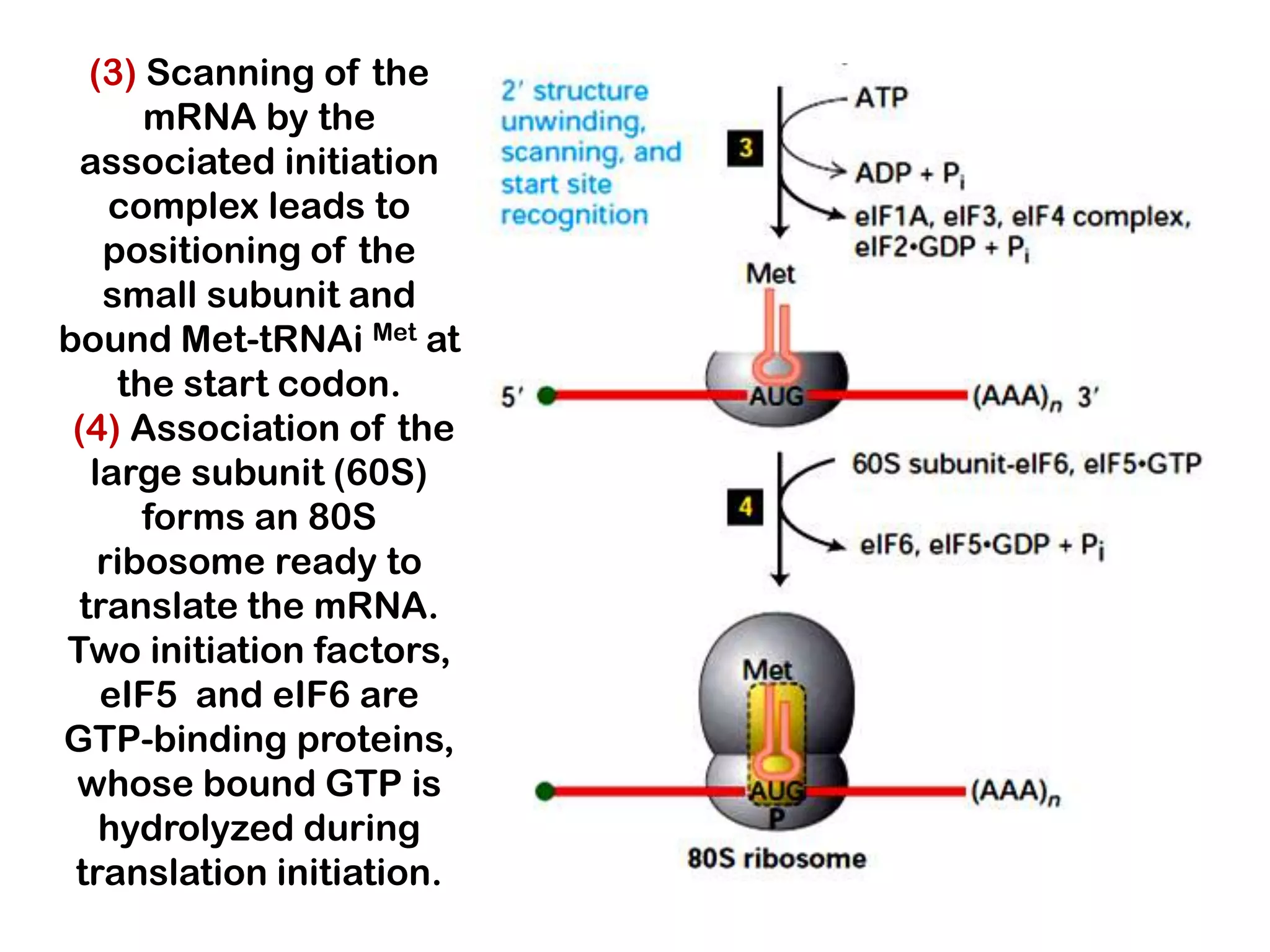
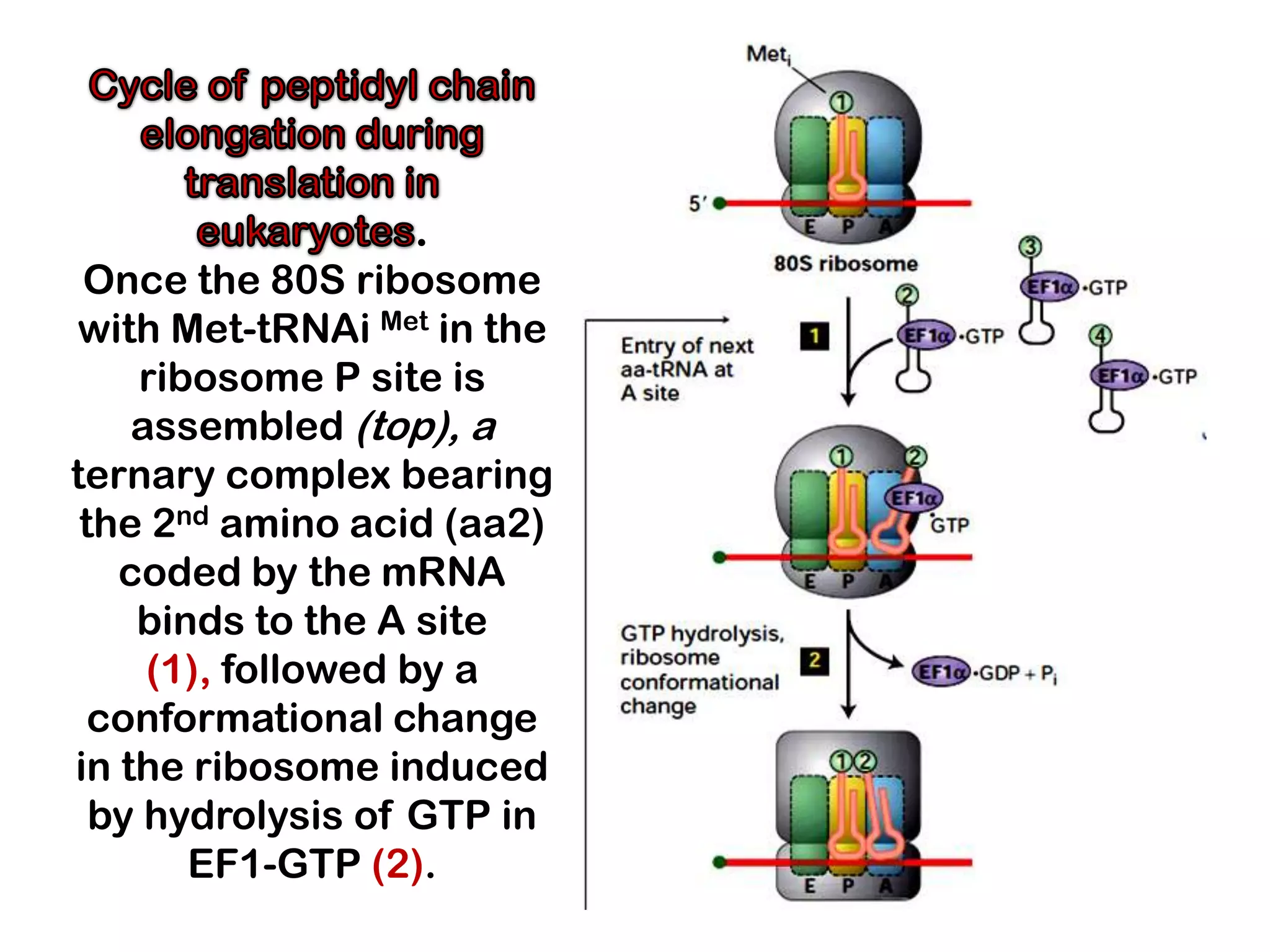
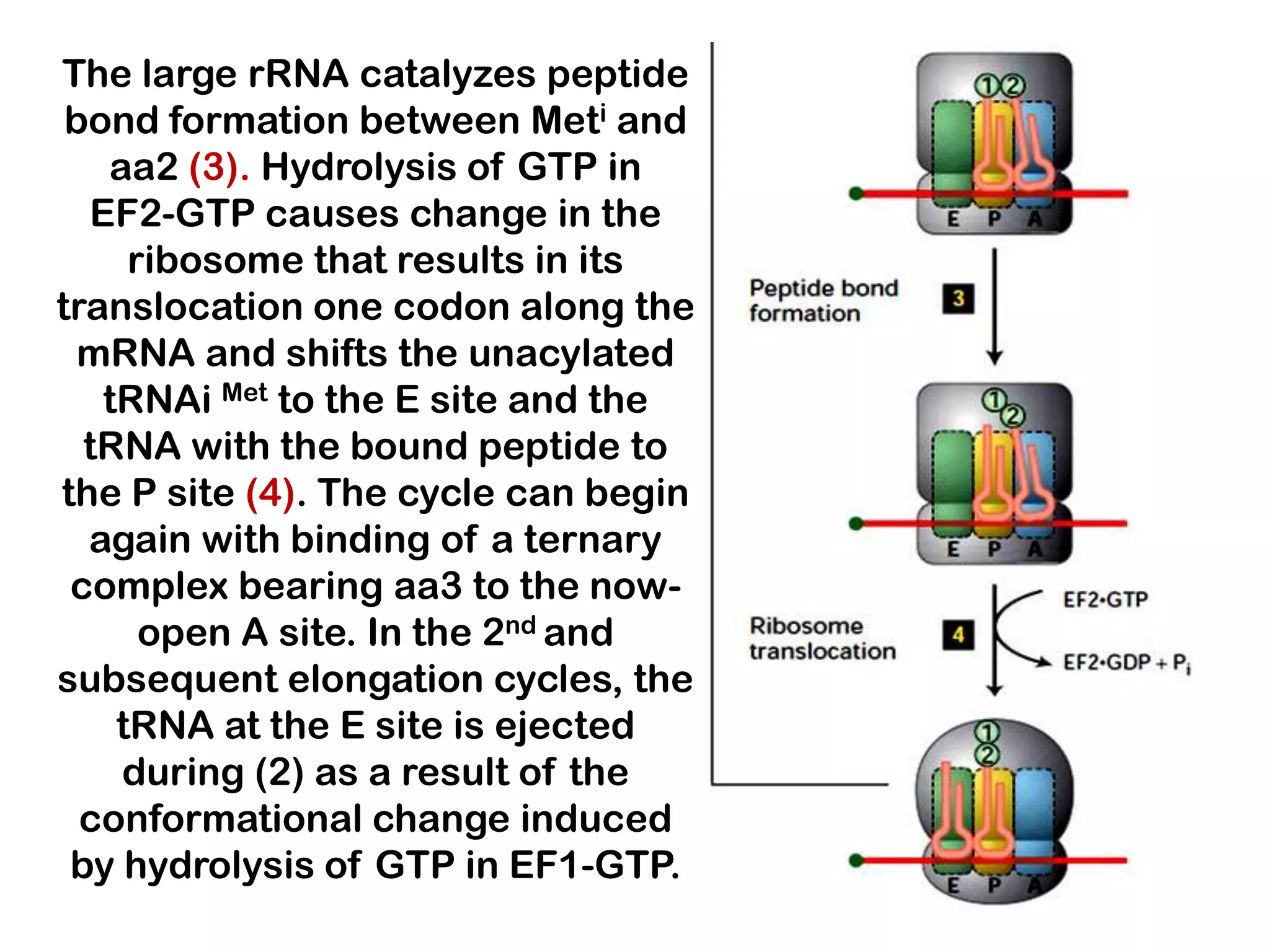
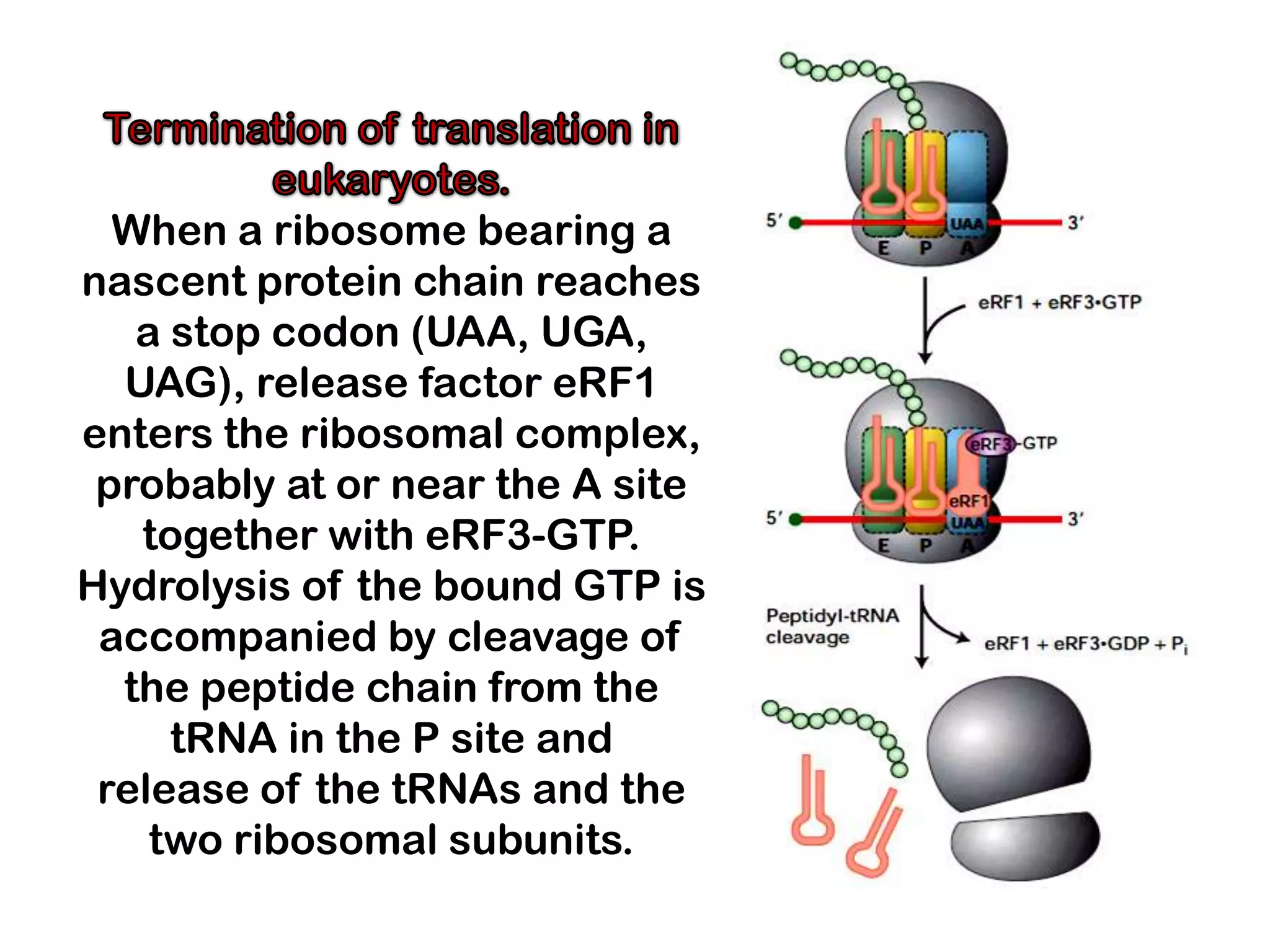
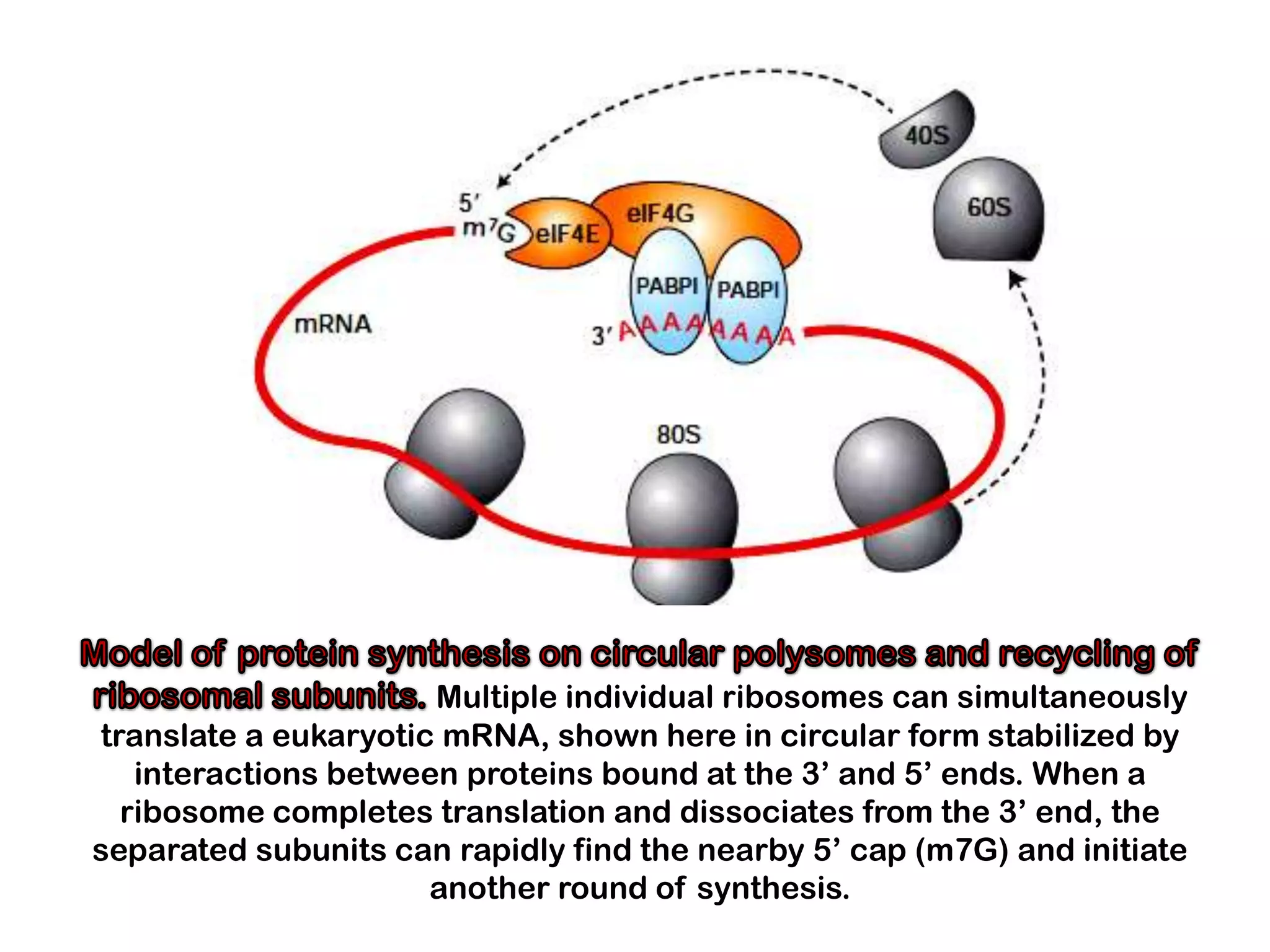
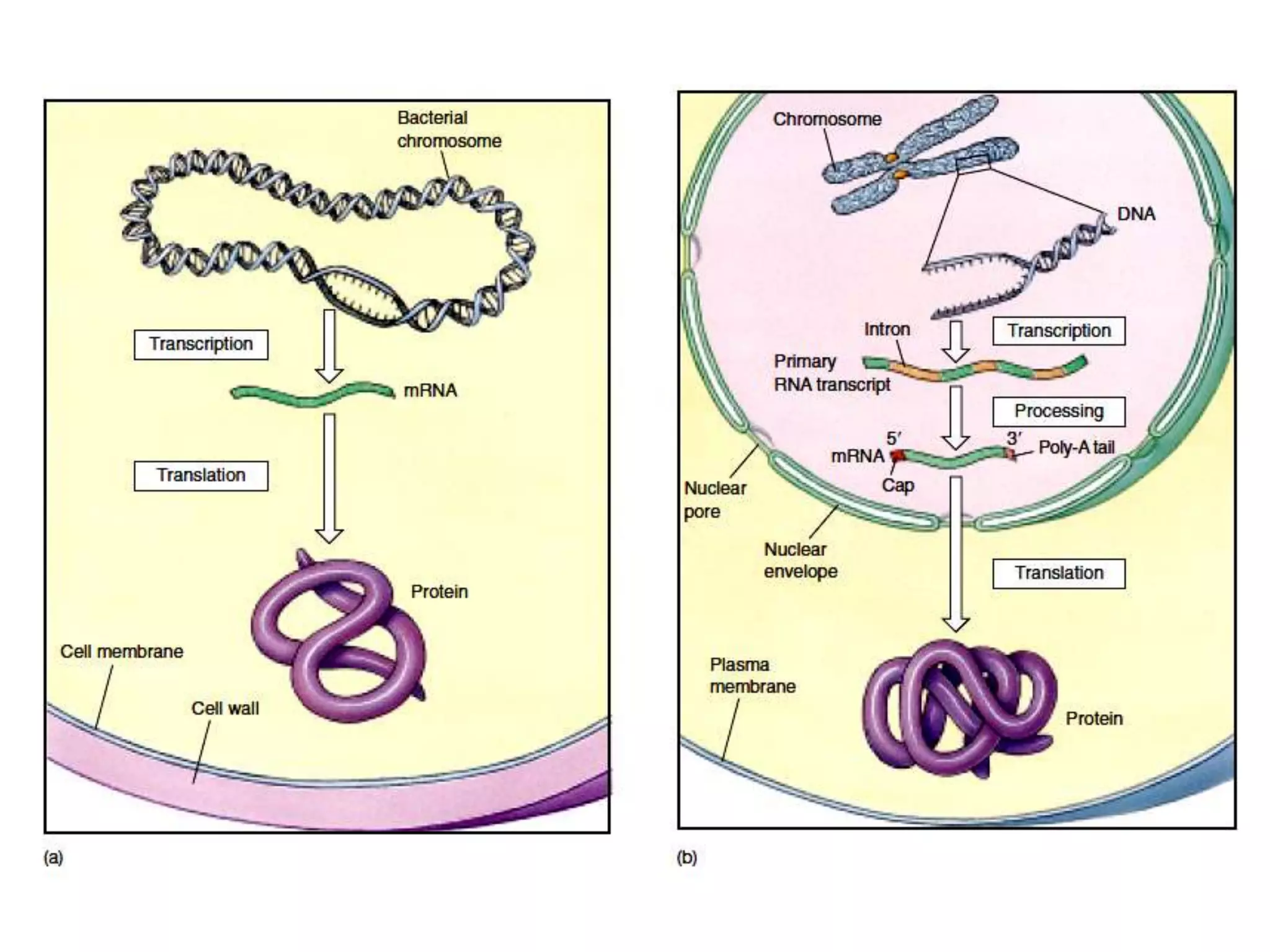
Transcription, RNA processing, and translation are the processes that link DNA sequences to protein synthesis. Translation occurs via ribosomes on the mRNA, which catalyze peptide bond formation between amino acids carried by tRNAs according to the mRNA codon sequence. Additional steps include RNA processing, modification of tRNAs and proteins, and initiation and termination factors that regulate translation.